Please be patient as the page loads
|
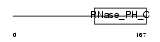
EXOS9_HUMAN
|
||||||
| SwissProt Accessions | Q06265, Q12883 | Gene names | EXOSC9, PMSCL1 | |||
|
Domain Architecture |
|
|||||
| Description | Exosome complex exonuclease RRP45 (EC 3.1.13.-) (Exosome component 9) (Polymyositis/scleroderma autoantigen 1) (Autoantigen PM/Scl 1) (Polymyositis/scleroderma autoantigen 75 kDa) (PM/Scl-75) (P75 polymyositis-scleroderma overlap syndrome-associated autoantigen). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
EXOS9_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q06265, Q12883 | Gene names | EXOSC9, PMSCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome complex exonuclease RRP45 (EC 3.1.13.-) (Exosome component 9) (Polymyositis/scleroderma autoantigen 1) (Autoantigen PM/Scl 1) (Polymyositis/scleroderma autoantigen 75 kDa) (PM/Scl-75) (P75 polymyositis-scleroderma overlap syndrome-associated autoantigen). | |||||
|
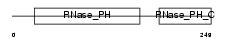
EXOS9_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.997243 (rank : 2) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JHI7, Q9CSZ2 | Gene names | Exosc9, Pmscl1 | |||
|
Domain Architecture |
|
|||||
| Description | Exosome complex exonuclease RRP45 (EC 3.1.13.-) (Exosome component 9) (Polymyositis/scleroderma autoantigen 1) (Autoantigen PM/Scl 1) (Polymyositis/scleroderma autoantigen 75 kDa) (PM/Scl-75) (P75 polymyositis-scleroderma overlap syndrome-associated autoantigen). | |||||
|
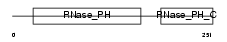
EXOS8_HUMAN
|
||||||
| θ value | 2.51237e-26 (rank : 3) | NC score | 0.789458 (rank : 3) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96B26, O43480 | Gene names | EXOSC8, OIP2, RRP43 | |||
|
Domain Architecture |
|
|||||
| Description | Exosome complex exonuclease RRP43 (EC 3.1.13.-) (Ribosomal RNA- processing protein 43) (Exosome component 8) (p9) (Opa-interacting protein 2). | |||||
|
EXOS8_MOUSE
|
||||||
| θ value | 1.37858e-24 (rank : 4) | NC score | 0.782307 (rank : 4) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D753 | Gene names | Exosc8, Rrp43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exosome complex exonuclease RRP43 (EC 3.1.13.-) (Ribosomal RNA- processing protein 43) (Exosome component 8). | |||||
|
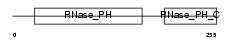
EXOS7_HUMAN
|
||||||
| θ value | 2.27762e-19 (rank : 5) | NC score | 0.732618 (rank : 5) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15024, Q96E72 | Gene names | EXOSC7, KIAA0116, RRP42 | |||
|
Domain Architecture |
|
|||||
| Description | Exosome complex exonuclease RRP42 (EC 3.1.13.-) (Ribosomal RNA- processing protein 42) (Exosome component 7) (p8). | |||||
|
EXOS7_MOUSE
|
||||||
| θ value | 2.35696e-16 (rank : 6) | NC score | 0.711781 (rank : 6) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D0M0, Q9CT77 | Gene names | Exosc7, Rrp42 | |||
|
Domain Architecture |
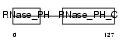
|
|||||
| Description | Exosome complex exonuclease RRP42 (EC 3.1.13.-) (Ribosomal RNA- processing protein 42) (Exosome component 7). | |||||
|
PCDGL_HUMAN
|
||||||
| θ value | 2.36792 (rank : 7) | NC score | 0.006599 (rank : 10) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5F7, Q9Y5C3 | Gene names | PCDHGC4 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin gamma C4 precursor (PCDH-gamma-C4). | |||||
|
SYS_HUMAN
|
||||||
| θ value | 5.27518 (rank : 8) | NC score | 0.018875 (rank : 8) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49591, Q9NSE3 | Gene names | SARS, SERS | |||
|
Domain Architecture |
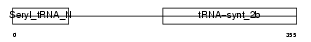
|
|||||
| Description | Seryl-tRNA synthetase (EC 6.1.1.11) (Serine--tRNA ligase) (SerRS). | |||||
|
NDUA5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 9) | NC score | 0.020080 (rank : 7) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CPP6, Q9CY90, Q9D2P2, Q9D703, Q9D739 | Gene names | Ndufa5 | |||
|
Domain Architecture |
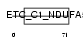
|
|||||
| Description | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 5 (EC 1.6.5.3) (EC 1.6.99.3) (NADH-ubiquinone oxidoreductase 13 kDa-B subunit) (Complex I-13kD-B) (CI-13kD-B) (Complex I subunit B13). | |||||
|
THOC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 10) | NC score | 0.012967 (rank : 9) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
EXOS9_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q06265, Q12883 | Gene names | EXOSC9, PMSCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome complex exonuclease RRP45 (EC 3.1.13.-) (Exosome component 9) (Polymyositis/scleroderma autoantigen 1) (Autoantigen PM/Scl 1) (Polymyositis/scleroderma autoantigen 75 kDa) (PM/Scl-75) (P75 polymyositis-scleroderma overlap syndrome-associated autoantigen). | |||||
|
EXOS9_MOUSE
|
||||||
| NC score | 0.997243 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JHI7, Q9CSZ2 | Gene names | Exosc9, Pmscl1 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome complex exonuclease RRP45 (EC 3.1.13.-) (Exosome component 9) (Polymyositis/scleroderma autoantigen 1) (Autoantigen PM/Scl 1) (Polymyositis/scleroderma autoantigen 75 kDa) (PM/Scl-75) (P75 polymyositis-scleroderma overlap syndrome-associated autoantigen). | |||||
|
EXOS8_HUMAN
|
||||||
| NC score | 0.789458 (rank : 3) | θ value | 2.51237e-26 (rank : 3) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96B26, O43480 | Gene names | EXOSC8, OIP2, RRP43 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome complex exonuclease RRP43 (EC 3.1.13.-) (Ribosomal RNA- processing protein 43) (Exosome component 8) (p9) (Opa-interacting protein 2). | |||||
|
EXOS8_MOUSE
|
||||||
| NC score | 0.782307 (rank : 4) | θ value | 1.37858e-24 (rank : 4) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D753 | Gene names | Exosc8, Rrp43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exosome complex exonuclease RRP43 (EC 3.1.13.-) (Ribosomal RNA- processing protein 43) (Exosome component 8). | |||||
|
EXOS7_HUMAN
|
||||||
| NC score | 0.732618 (rank : 5) | θ value | 2.27762e-19 (rank : 5) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15024, Q96E72 | Gene names | EXOSC7, KIAA0116, RRP42 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome complex exonuclease RRP42 (EC 3.1.13.-) (Ribosomal RNA- processing protein 42) (Exosome component 7) (p8). | |||||
|
EXOS7_MOUSE
|
||||||
| NC score | 0.711781 (rank : 6) | θ value | 2.35696e-16 (rank : 6) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D0M0, Q9CT77 | Gene names | Exosc7, Rrp42 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome complex exonuclease RRP42 (EC 3.1.13.-) (Ribosomal RNA- processing protein 42) (Exosome component 7). | |||||
|
NDUA5_MOUSE
|
||||||
| NC score | 0.020080 (rank : 7) | θ value | 8.99809 (rank : 9) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CPP6, Q9CY90, Q9D2P2, Q9D703, Q9D739 | Gene names | Ndufa5 | |||
|
Domain Architecture |

|
|||||
| Description | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 5 (EC 1.6.5.3) (EC 1.6.99.3) (NADH-ubiquinone oxidoreductase 13 kDa-B subunit) (Complex I-13kD-B) (CI-13kD-B) (Complex I subunit B13). | |||||
|
SYS_HUMAN
|
||||||
| NC score | 0.018875 (rank : 8) | θ value | 5.27518 (rank : 8) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49591, Q9NSE3 | Gene names | SARS, SERS | |||
|
Domain Architecture |

|
|||||
| Description | Seryl-tRNA synthetase (EC 6.1.1.11) (Serine--tRNA ligase) (SerRS). | |||||
|
THOC2_HUMAN
|
||||||
| NC score | 0.012967 (rank : 9) | θ value | 8.99809 (rank : 10) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
PCDGL_HUMAN
|
||||||
| NC score | 0.006599 (rank : 10) | θ value | 2.36792 (rank : 7) | |||
| Query Neighborhood Hits | 10 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5F7, Q9Y5C3 | Gene names | PCDHGC4 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin gamma C4 precursor (PCDH-gamma-C4). | |||||