Please be patient as the page loads
|
DEPD5_HUMAN
|
||||||
| SwissProt Accessions | O75140, Q5K3V5, Q5THY9, Q5THZ0, Q5THZ1, Q5THZ3, Q68DR1, Q6MZX3, Q6PEZ1, Q9UGV8, Q9UH13 | Gene names | DEPDC5, KIAA0645 | |||
|
Domain Architecture |

|
|||||
| Description | DEP domain-containing protein 5. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DEPD5_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O75140, Q5K3V5, Q5THY9, Q5THZ0, Q5THZ1, Q5THZ3, Q68DR1, Q6MZX3, Q6PEZ1, Q9UGV8, Q9UH13 | Gene names | DEPDC5, KIAA0645 | |||
|
Domain Architecture |

|
|||||
| Description | DEP domain-containing protein 5. | |||||
|
DEPD5_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.992797 (rank : 2) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P61460 | Gene names | Depdc5, Kiaa0645 | |||
|
Domain Architecture |

|
|||||
| Description | DEP domain-containing protein 5. | |||||
|
DVL1_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 3) | NC score | 0.140351 (rank : 5) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
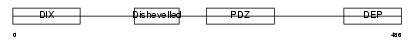
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
RPGF4_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 4) | NC score | 0.086152 (rank : 10) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9EQZ6, Q8VIP9, Q9CW52, Q9Z1P0 | Gene names | Rapgef4, Cgef2, Epac2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 4 (cAMP-regulated guanine nucleotide exchange factor II) (cAMP-GEFII) (Exchange factor directly activated by cAMP 2) (Epac 2) (cAMP-dependent Rap1 guanine-nucleotide exchange factor). | |||||
|
DVL1L_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 5) | NC score | 0.134759 (rank : 7) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
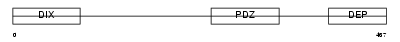
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
DVL2_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 6) | NC score | 0.137021 (rank : 6) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL3_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 7) | NC score | 0.147510 (rank : 4) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
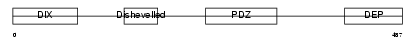
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL3_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 8) | NC score | 0.147649 (rank : 3) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
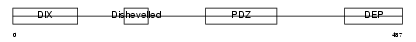
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 9) | NC score | 0.130581 (rank : 9) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL2_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 10) | NC score | 0.133602 (rank : 8) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
FYV1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.084397 (rank : 11) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z1T6 | Gene names | Pip5k3, Pikfyve | |||
|
Domain Architecture |

|
|||||
| Description | FYVE finger-containing phosphoinositide kinase (EC 2.7.1.68) (1- phosphatidylinositol-4-phosphate 5-kinase) (PIP5K) (PtdIns(4)P-5- kinase) (PIKfyve) (p235). | |||||
|
FYV1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 12) | NC score | 0.084026 (rank : 12) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y2I7, Q8NB67 | Gene names | PIP5K3, KIAA0981, PIKFYVE | |||
|
Domain Architecture |

|
|||||
| Description | FYVE finger-containing phosphoinositide kinase (EC 2.7.1.68) (1- phosphatidylinositol-4-phosphate 5-kinase) (Phosphatidylinositol-3- phosphate 5-kinase type III) (PIP5K) (PtdIns(4)P-5-kinase) (PIKfyve) (p235). | |||||
|
RPGF4_HUMAN
|
||||||
| θ value | 0.163984 (rank : 13) | NC score | 0.077563 (rank : 13) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8WZA2, O95636, Q8IXK6, Q8TAA4 | Gene names | RAPGEF4, CGEF2, EPAC2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 4 (cAMP-regulated guanine nucleotide exchange factor II) (cAMP-GEFII) (Exchange factor directly activated by cAMP 2) (Epac 2). | |||||
|
BMP10_MOUSE
|
||||||
| θ value | 0.365318 (rank : 14) | NC score | 0.027219 (rank : 18) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R229, Q9Z1V8 | Gene names | Bmp10 | |||
|
Domain Architecture |
|
|||||
| Description | Bone morphogenetic protein 10 precursor (BMP-10). | |||||
|
RPGF5_MOUSE
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.058509 (rank : 14) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C0Q9, Q8BJJ9, Q8C0R5 | Gene names | Rapgef5, Gfr, Kiaa0277, Mrgef | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 5 (Guanine nucleotide exchange factor for Rap1) (M-Ras-regulated Rap GEF) (MR-GEF). | |||||
|
DMD_MOUSE
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.014250 (rank : 32) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
NICA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 17) | NC score | 0.031140 (rank : 17) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92542, Q86VV5 | Gene names | NCSTN, KIAA0253 | |||
|
Domain Architecture |

|
|||||
| Description | Nicastrin precursor. | |||||
|
DMD_HUMAN
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.014404 (rank : 30) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
RNZ1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.055032 (rank : 16) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H777, Q9NS99 | Gene names | ELAC1, D29 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc phosphodiesterase ELAC protein 1 (EC 3.1.26.11) (Ribonuclease Z 1) (RNase Z 1) (tRNase Z 1) (tRNA 3 endonuclease 1) (ElaC homolog protein 1) (Deleted in Ma29). | |||||
|
RNZ1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.055305 (rank : 15) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VEB6, Q9CXB1, Q9ERF4 | Gene names | Elac1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc phosphodiesterase ELAC protein 1 (EC 3.1.26.11) (Ribonuclease Z 1) (RNase Z 1) (tRNase Z 1) (tRNA 3 endonuclease 1) (ElaC homolog protein 1). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.008070 (rank : 36) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
KSYK_MOUSE
|
||||||
| θ value | 3.0926 (rank : 22) | NC score | 0.003421 (rank : 39) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48025 | Gene names | Syk | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase SYK (EC 2.7.10.2) (Spleen tyrosine kinase). | |||||
|
NFAT5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 23) | NC score | 0.021726 (rank : 19) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WV30 | Gene names | Nfat5 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T cell transcription factor NFAT5) (NF-AT5) (Rel domain-containing transcription factor NFAT5). | |||||
|
GAK6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.021548 (rank : 20) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
KIF2C_MOUSE
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.009501 (rank : 35) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q922S8 | Gene names | Kif2c | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF2C (Mitotic centromere-associated kinesin) (MCAK). | |||||
|
LTBP4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.004778 (rank : 38) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K4G1, Q8K4G0 | Gene names | Ltbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 4 precursor (LTBP-4). | |||||
|
CEL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.007469 (rank : 37) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64285 | Gene names | Cel, Lip1 | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
KLDC4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.015304 (rank : 29) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TBB5, Q96F29, Q9BVN3 | Gene names | KLHDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch domain-containing protein 4. | |||||
|
KSYK_HUMAN
|
||||||
| θ value | 6.88961 (rank : 29) | NC score | 0.002541 (rank : 40) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43405 | Gene names | SYK | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase SYK (EC 2.7.10.2) (Spleen tyrosine kinase). | |||||
|
SMOO_HUMAN
|
||||||
| θ value | 6.88961 (rank : 30) | NC score | 0.014299 (rank : 31) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53814, O00569, O95769, O95937, Q9P1S8, Q9UIT1, Q9UIT2 | Gene names | SMTN, SMSMO | |||
|
Domain Architecture |

|
|||||
| Description | Smoothelin. | |||||
|
STAU2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.018356 (rank : 27) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NUL3, Q6AHY7, Q96HM0, Q96HM1, Q9NVI5, Q9UGG6 | Gene names | STAU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
STAU2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.018219 (rank : 28) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CJ67, Q8BSY8, Q8CJ66, Q8R175, Q91Z19, Q9D5N7 | Gene names | Stau2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
GAK11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.019604 (rank : 25) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.019808 (rank : 22) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.020128 (rank : 21) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62683 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.019627 (rank : 23) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.019619 (rank : 24) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.019551 (rank : 26) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
PIGG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 39) | NC score | 0.014110 (rank : 33) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5H8A4, Q2TAK5, Q6UX31, Q7L5Y4, Q8N866, Q8NCC9, Q96SY9, Q9BVT7, Q9NXG5 | Gene names | PIGG, GPI7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 2 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class G protein) (PIG-G) (GPI7 homolog) (hGPI7). | |||||
|
TF3C5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 40) | NC score | 0.014003 (rank : 34) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5Q8, Q8N2U7, Q8N857, Q96GD9, Q9H4P2 | Gene names | GTF3C5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 5 (Transcription factor IIIC-epsilon subunit) (TF3C-epsilon) (TFIIIC 63 kDa subunit) (TFIIIC63). | |||||
|
DEPD5_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O75140, Q5K3V5, Q5THY9, Q5THZ0, Q5THZ1, Q5THZ3, Q68DR1, Q6MZX3, Q6PEZ1, Q9UGV8, Q9UH13 | Gene names | DEPDC5, KIAA0645 | |||
|
Domain Architecture |

|
|||||
| Description | DEP domain-containing protein 5. | |||||
|
DEPD5_MOUSE
|
||||||
| NC score | 0.992797 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P61460 | Gene names | Depdc5, Kiaa0645 | |||
|
Domain Architecture |

|
|||||
| Description | DEP domain-containing protein 5. | |||||
|
DVL3_MOUSE
|
||||||
| NC score | 0.147649 (rank : 3) | θ value | 0.0431538 (rank : 8) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL3_HUMAN
|
||||||
| NC score | 0.147510 (rank : 4) | θ value | 0.0431538 (rank : 7) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL1_MOUSE
|
||||||
| NC score | 0.140351 (rank : 5) | θ value | 0.00869519 (rank : 3) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL2_HUMAN
|
||||||
| NC score | 0.137021 (rank : 6) | θ value | 0.0330416 (rank : 6) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL1L_HUMAN
|
||||||
| NC score | 0.134759 (rank : 7) | θ value | 0.0330416 (rank : 5) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
DVL2_MOUSE
|
||||||
| NC score | 0.133602 (rank : 8) | θ value | 0.0563607 (rank : 10) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL1_HUMAN
|
||||||
| NC score | 0.130581 (rank : 9) | θ value | 0.0563607 (rank : 9) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
RPGF4_MOUSE
|
||||||
| NC score | 0.086152 (rank : 10) | θ value | 0.0193708 (rank : 4) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9EQZ6, Q8VIP9, Q9CW52, Q9Z1P0 | Gene names | Rapgef4, Cgef2, Epac2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 4 (cAMP-regulated guanine nucleotide exchange factor II) (cAMP-GEFII) (Exchange factor directly activated by cAMP 2) (Epac 2) (cAMP-dependent Rap1 guanine-nucleotide exchange factor). | |||||
|
FYV1_MOUSE
|
||||||
| NC score | 0.084397 (rank : 11) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z1T6 | Gene names | Pip5k3, Pikfyve | |||
|
Domain Architecture |

|
|||||
| Description | FYVE finger-containing phosphoinositide kinase (EC 2.7.1.68) (1- phosphatidylinositol-4-phosphate 5-kinase) (PIP5K) (PtdIns(4)P-5- kinase) (PIKfyve) (p235). | |||||
|
FYV1_HUMAN
|
||||||
| NC score | 0.084026 (rank : 12) | θ value | 0.163984 (rank : 12) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y2I7, Q8NB67 | Gene names | PIP5K3, KIAA0981, PIKFYVE | |||
|
Domain Architecture |

|
|||||
| Description | FYVE finger-containing phosphoinositide kinase (EC 2.7.1.68) (1- phosphatidylinositol-4-phosphate 5-kinase) (Phosphatidylinositol-3- phosphate 5-kinase type III) (PIP5K) (PtdIns(4)P-5-kinase) (PIKfyve) (p235). | |||||
|
RPGF4_HUMAN
|
||||||
| NC score | 0.077563 (rank : 13) | θ value | 0.163984 (rank : 13) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8WZA2, O95636, Q8IXK6, Q8TAA4 | Gene names | RAPGEF4, CGEF2, EPAC2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 4 (cAMP-regulated guanine nucleotide exchange factor II) (cAMP-GEFII) (Exchange factor directly activated by cAMP 2) (Epac 2). | |||||
|
RPGF5_MOUSE
|
||||||
| NC score | 0.058509 (rank : 14) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C0Q9, Q8BJJ9, Q8C0R5 | Gene names | Rapgef5, Gfr, Kiaa0277, Mrgef | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 5 (Guanine nucleotide exchange factor for Rap1) (M-Ras-regulated Rap GEF) (MR-GEF). | |||||
|
RNZ1_MOUSE
|
||||||
| NC score | 0.055305 (rank : 15) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VEB6, Q9CXB1, Q9ERF4 | Gene names | Elac1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc phosphodiesterase ELAC protein 1 (EC 3.1.26.11) (Ribonuclease Z 1) (RNase Z 1) (tRNase Z 1) (tRNA 3 endonuclease 1) (ElaC homolog protein 1). | |||||
|
RNZ1_HUMAN
|
||||||
| NC score | 0.055032 (rank : 16) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H777, Q9NS99 | Gene names | ELAC1, D29 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc phosphodiesterase ELAC protein 1 (EC 3.1.26.11) (Ribonuclease Z 1) (RNase Z 1) (tRNase Z 1) (tRNA 3 endonuclease 1) (ElaC homolog protein 1) (Deleted in Ma29). | |||||
|
NICA_HUMAN
|
||||||
| NC score | 0.031140 (rank : 17) | θ value | 1.81305 (rank : 17) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92542, Q86VV5 | Gene names | NCSTN, KIAA0253 | |||
|
Domain Architecture |

|
|||||
| Description | Nicastrin precursor. | |||||
|
BMP10_MOUSE
|
||||||
| NC score | 0.027219 (rank : 18) | θ value | 0.365318 (rank : 14) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R229, Q9Z1V8 | Gene names | Bmp10 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 10 precursor (BMP-10). | |||||
|
NFAT5_MOUSE
|
||||||
| NC score | 0.021726 (rank : 19) | θ value | 3.0926 (rank : 23) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WV30 | Gene names | Nfat5 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T cell transcription factor NFAT5) (NF-AT5) (Rel domain-containing transcription factor NFAT5). | |||||
|
GAK6_HUMAN
|
||||||
| NC score | 0.021548 (rank : 20) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK1_HUMAN
|
||||||
| NC score | 0.020128 (rank : 21) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62683 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK12_HUMAN
|
||||||
| NC score | 0.019808 (rank : 22) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK2_HUMAN
|
||||||
| NC score | 0.019627 (rank : 23) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| NC score | 0.019619 (rank : 24) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK11_HUMAN
|
||||||
| NC score | 0.019604 (rank : 25) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK5_HUMAN
|
||||||
| NC score | 0.019551 (rank : 26) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
STAU2_HUMAN
|
||||||
| NC score | 0.018356 (rank : 27) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NUL3, Q6AHY7, Q96HM0, Q96HM1, Q9NVI5, Q9UGG6 | Gene names | STAU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
STAU2_MOUSE
|
||||||
| NC score | 0.018219 (rank : 28) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CJ67, Q8BSY8, Q8CJ66, Q8R175, Q91Z19, Q9D5N7 | Gene names | Stau2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
KLDC4_HUMAN
|
||||||
| NC score | 0.015304 (rank : 29) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TBB5, Q96F29, Q9BVN3 | Gene names | KLHDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch domain-containing protein 4. | |||||
|
DMD_HUMAN
|
||||||
| NC score | 0.014404 (rank : 30) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
SMOO_HUMAN
|
||||||
| NC score | 0.014299 (rank : 31) | θ value | 6.88961 (rank : 30) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53814, O00569, O95769, O95937, Q9P1S8, Q9UIT1, Q9UIT2 | Gene names | SMTN, SMSMO | |||
|
Domain Architecture |

|
|||||
| Description | Smoothelin. | |||||
|
DMD_MOUSE
|
||||||
| NC score | 0.014250 (rank : 32) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
PIGG_HUMAN
|
||||||
| NC score | 0.014110 (rank : 33) | θ value | 8.99809 (rank : 39) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5H8A4, Q2TAK5, Q6UX31, Q7L5Y4, Q8N866, Q8NCC9, Q96SY9, Q9BVT7, Q9NXG5 | Gene names | PIGG, GPI7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 2 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class G protein) (PIG-G) (GPI7 homolog) (hGPI7). | |||||
|
TF3C5_HUMAN
|
||||||
| NC score | 0.014003 (rank : 34) | θ value | 8.99809 (rank : 40) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5Q8, Q8N2U7, Q8N857, Q96GD9, Q9H4P2 | Gene names | GTF3C5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 5 (Transcription factor IIIC-epsilon subunit) (TF3C-epsilon) (TFIIIC 63 kDa subunit) (TFIIIC63). | |||||
|
KIF2C_MOUSE
|
||||||
| NC score | 0.009501 (rank : 35) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q922S8 | Gene names | Kif2c | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF2C (Mitotic centromere-associated kinesin) (MCAK). | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.008070 (rank : 36) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
CEL_MOUSE
|
||||||
| NC score | 0.007469 (rank : 37) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64285 | Gene names | Cel, Lip1 | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
LTBP4_MOUSE
|
||||||
| NC score | 0.004778 (rank : 38) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K4G1, Q8K4G0 | Gene names | Ltbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 4 precursor (LTBP-4). | |||||
|
KSYK_MOUSE
|
||||||
| NC score | 0.003421 (rank : 39) | θ value | 3.0926 (rank : 22) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48025 | Gene names | Syk | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase SYK (EC 2.7.10.2) (Spleen tyrosine kinase). | |||||
|
KSYK_HUMAN
|
||||||
| NC score | 0.002541 (rank : 40) | θ value | 6.88961 (rank : 29) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43405 | Gene names | SYK | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase SYK (EC 2.7.10.2) (Spleen tyrosine kinase). | |||||