Please be patient as the page loads
|
CPXM1_HUMAN
|
||||||
| SwissProt Accessions | Q96SM3, Q6P4G8, Q6UW65, Q9NUB5 | Gene names | CPXM1, CPX1, CPXM | |||
|
Domain Architecture |

|
|||||
| Description | Probable carboxypeptidase X1 precursor (EC 3.4.17.-) (Metallocarboxypeptidase CPX-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CPXM1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q96SM3, Q6P4G8, Q6UW65, Q9NUB5 | Gene names | CPXM1, CPX1, CPXM | |||
|
Domain Architecture |

|
|||||
| Description | Probable carboxypeptidase X1 precursor (EC 3.4.17.-) (Metallocarboxypeptidase CPX-1). | |||||
|
CPXM1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.997373 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9Z100, Q99LA3 | Gene names | Cpxm1, Cpx1, Cpxm | |||
|
Domain Architecture |

|
|||||
| Description | Probable carboxypeptidase X1 precursor (EC 3.4.17.-) (Metallocarboxypeptidase CPX-1). | |||||
|
CPXM2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.990065 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8N436 | Gene names | CPXM2 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase-like protein X2 precursor. | |||||
|
CPXM2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.990666 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D2L5, O54860, Q8VDQ4 | Gene names | Cpxm2, Cpx2 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase-like protein X2 precursor. | |||||
|
CBPE_MOUSE
|
||||||
| θ value | 1.39067e-109 (rank : 5) | NC score | 0.878648 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q00493, Q64439 | Gene names | Cpe | |||
|
Domain Architecture |
|
|||||
| Description | Carboxypeptidase E precursor (EC 3.4.17.10) (CPE) (Carboxypeptidase H) (CPH) (Enkephalin convertase) (Prohormone-processing carboxypeptidase). | |||||
|
CBPE_HUMAN
|
||||||
| θ value | 3.0981e-109 (rank : 6) | NC score | 0.878590 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P16870, Q9UIU9 | Gene names | CPE | |||
|
Domain Architecture |
|
|||||
| Description | Carboxypeptidase E precursor (EC 3.4.17.10) (CPE) (Carboxypeptidase H) (CPH) (Enkephalin convertase) (Prohormone-processing carboxypeptidase). | |||||
|
CBPZ_HUMAN
|
||||||
| θ value | 1.30163e-107 (rank : 7) | NC score | 0.790153 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q66K79, O00520, Q96MX2 | Gene names | CPZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase Z precursor (EC 3.4.17.-) (CPZ). | |||||
|
CBPZ_MOUSE
|
||||||
| θ value | 6.45994e-107 (rank : 8) | NC score | 0.806005 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R4V4 | Gene names | Cpz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase Z precursor (EC 3.4.17.-) (CPZ). | |||||
|
CBPN_HUMAN
|
||||||
| θ value | 2.07809e-105 (rank : 9) | NC score | 0.879841 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P15169 | Gene names | CPN1, ACBP | |||
|
Domain Architecture |
|
|||||
| Description | Carboxypeptidase N catalytic chain precursor (EC 3.4.17.3) (CPN) (Carboxypeptidase N polypeptide 1) (Carboxypeptidase N small subunit) (Lysine carboxypeptidase) (Arginine carboxypeptidase) (Kininase-1) (Serum carboxypeptidase N) (SCPN) (Anaphylatoxin inactivator) (Plasma carboxypeptidase B). | |||||
|
CBPN_MOUSE
|
||||||
| θ value | 1.75921e-104 (rank : 10) | NC score | 0.879445 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9JJN5, Q91WM9 | Gene names | Cpn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase N catalytic chain precursor (EC 3.4.17.3) (CPN) (Carboxypeptidase N polypeptide 1) (Carboxypeptidase N small subunit). | |||||
|
CBPD_MOUSE
|
||||||
| θ value | 2.81514e-94 (rank : 11) | NC score | 0.844853 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O89001 | Gene names | Cpd | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase D precursor (EC 3.4.17.22) (Metallocarboxypeptidase D) (gp180). | |||||
|
CBPD_HUMAN
|
||||||
| θ value | 2.38316e-93 (rank : 12) | NC score | 0.844918 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75976, O15377, Q86SH9, Q86XE6 | Gene names | CPD | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase D precursor (EC 3.4.17.22) (Metallocarboxypeptidase D) (gp180). | |||||
|
CBPM_MOUSE
|
||||||
| θ value | 1.5555e-68 (rank : 13) | NC score | 0.857771 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80V42, Q497S5, Q9CYH8 | Gene names | Cpm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase M precursor (EC 3.4.17.12) (CPM). | |||||
|
CBPM_HUMAN
|
||||||
| θ value | 1.6097e-65 (rank : 14) | NC score | 0.857662 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P14384, Q9H2K9 | Gene names | CPM | |||
|
Domain Architecture |
|
|||||
| Description | Carboxypeptidase M precursor (EC 3.4.17.12) (CPM). | |||||
|
MFGM_HUMAN
|
||||||
| θ value | 7.82807e-20 (rank : 15) | NC score | 0.518597 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08431 | Gene names | MFGE8 | |||
|
Domain Architecture |
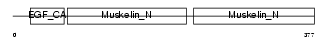
|
|||||
| Description | Lactadherin precursor (Milk fat globule-EGF factor 8) (MFG-E8) (HMFG) (Breast epithelial antigen BA46) (MFGM) [Contains: Lactadherin short form; Medin]. | |||||
|
EDIL3_MOUSE
|
||||||
| θ value | 1.02238e-19 (rank : 16) | NC score | 0.362083 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35474, O35475 | Gene names | Edil3, Del1 | |||
|
Domain Architecture |
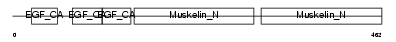
|
|||||
| Description | EGF-like repeat and discoidin I-like domain-containing protein 3 precursor (EGF-like repeats and discoidin I-like domains protein 3) (Developmentally-regulated endothelial cell locus 1 protein) (Integrin-binding protein DEL1). | |||||
|
MFGM_MOUSE
|
||||||
| θ value | 1.74391e-19 (rank : 17) | NC score | 0.449669 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P21956, P97800, Q3TBN5, Q9R1X9, Q9WTS3 | Gene names | Mfge8 | |||
|
Domain Architecture |
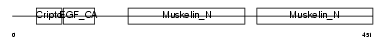
|
|||||
| Description | Lactadherin precursor (Milk fat globule-EGF factor 8) (MFG-E8) (MFGM) (Sperm surface protein SP47) (MP47). | |||||
|
EDIL3_HUMAN
|
||||||
| θ value | 6.62687e-19 (rank : 18) | NC score | 0.367580 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43854, O43855 | Gene names | EDIL3, DEL1 | |||
|
Domain Architecture |
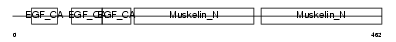
|
|||||
| Description | EGF-like repeat and discoidin I-like domain-containing protein 3 precursor (EGF-like repeats and discoidin I-like domains protein 3) (Developmentally-regulated endothelial cell locus 1 protein) (Integrin-binding protein DEL1). | |||||
|
XLRS1_MOUSE
|
||||||
| θ value | 9.56915e-18 (rank : 19) | NC score | 0.577953 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Z1L4 | Gene names | Rs1, Rs1h, Xlrs1 | |||
|
Domain Architecture |
|
|||||
| Description | Retinoschisin precursor (X-linked juvenile retinoschisis protein homolog). | |||||
|
XLRS1_HUMAN
|
||||||
| θ value | 3.63628e-17 (rank : 20) | NC score | 0.576018 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15537 | Gene names | RS1, XLRS1 | |||
|
Domain Architecture |
|
|||||
| Description | Retinoschisin precursor (X-linked juvenile retinoschisis protein). | |||||
|
FA5_HUMAN
|
||||||
| θ value | 3.07829e-16 (rank : 21) | NC score | 0.445552 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P12259, Q14285, Q6UPU6 | Gene names | F5 | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor V precursor (Activated protein C cofactor) [Contains: Coagulation factor V heavy chain; Coagulation factor V light chain]. | |||||
|
FA8_HUMAN
|
||||||
| θ value | 1.16975e-15 (rank : 22) | NC score | 0.442074 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P00451 | Gene names | F8, F8C | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component) (Antihemophilic factor) (AHF) [Contains: Factor VIIIa heavy chain, 200 kDa isoform; Factor VIIIa heavy chain, 92 kDa isoform; Factor VIII B chain; Factor VIIIa light chain]. | |||||
|
FA8_MOUSE
|
||||||
| θ value | 1.52774e-15 (rank : 23) | NC score | 0.442625 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q06194 | Gene names | F8, Cf8, F8c | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component). | |||||
|
CNTP2_MOUSE
|
||||||
| θ value | 9.90251e-15 (rank : 24) | NC score | 0.368365 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9CPW0, Q6P2K4, Q6ZQ31 | Gene names | Cntnap2, Kiaa0868 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Contactin-associated protein-like 2 precursor (Cell recognition molecule Caspr2). | |||||
|
CNTP2_HUMAN
|
||||||
| θ value | 1.68911e-14 (rank : 25) | NC score | 0.374746 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UHC6, Q52LV1, Q5H9Q7, Q9UQ12 | Gene names | CNTNAP2, CASPR2, KIAA0868 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 2 precursor (Cell recognition molecule Caspr2). | |||||
|
NRP2_HUMAN
|
||||||
| θ value | 1.68911e-14 (rank : 26) | NC score | 0.364294 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O60462, O14820, O14821 | Gene names | NRP2, VEGF165R2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-2 precursor (Vascular endothelial cell growth factor 165 receptor 2). | |||||
|
NRP2_MOUSE
|
||||||
| θ value | 4.91457e-14 (rank : 27) | NC score | 0.364740 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35375, O35373, O35374, O35376, O35377, O35378 | Gene names | Nrp2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-2 precursor (Vascular endothelial cell growth factor 165 receptor 2). | |||||
|
DCBD2_HUMAN
|
||||||
| θ value | 4.16044e-13 (rank : 28) | NC score | 0.410855 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96PD2, Q8N6M4, Q8TDX2 | Gene names | DCBLD2, CLCP1, ESDN | |||
|
Domain Architecture |
|
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 2 precursor (Endothelial and smooth muscle cell-derived neuropilin-like protein) (CUB, LCCL and coagulation factor V/VIII-homology domains protein 1). | |||||
|
DCBD2_MOUSE
|
||||||
| θ value | 4.16044e-13 (rank : 29) | NC score | 0.424364 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q91ZV3, Q8BKI4 | Gene names | Dcbld2, Esdn | |||
|
Domain Architecture |
|
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 2 precursor (Endothelial and smooth muscle cell-derived neuropilin-like protein). | |||||
|
CNTP3_HUMAN
|
||||||
| θ value | 2.0648e-12 (rank : 30) | NC score | 0.363582 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9BZ76, Q9C0E9 | Gene names | CNTNAP3, CASPR3, KIAA1714 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 3 precursor (Cell recognition molecule Caspr3). | |||||
|
NRP1_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 31) | NC score | 0.351479 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O14786, O60461, Q96IH5 | Gene names | NRP1, NRP, VEGF165R | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-1 precursor (Vascular endothelial cell growth factor 165 receptor) (CD304 antigen). | |||||
|
DDR2_HUMAN
|
||||||
| θ value | 3.89403e-11 (rank : 32) | NC score | 0.130787 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 818 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q16832 | Gene names | DDR2, NTRKR3, TKT, TYRO10 | |||
|
Domain Architecture |

|
|||||
| Description | Discoidin domain-containing receptor 2 precursor (EC 2.7.10.1) (Discoidin domain receptor 2) (Receptor protein-tyrosine kinase TKT) (Tyrosine-protein kinase TYRO 10) (Neurotrophic tyrosine kinase, receptor-related 3) (CD167b antigen). | |||||
|
DDR2_MOUSE
|
||||||
| θ value | 3.89403e-11 (rank : 33) | NC score | 0.131890 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q62371 | Gene names | Ddr2, Ntrkr3, Tkt, Tyro10 | |||
|
Domain Architecture |

|
|||||
| Description | Discoidin domain-containing receptor 2 precursor (EC 2.7.10.1) (Discoidin domain receptor 2) (Receptor protein-tyrosine kinase TKT) (Tyrosine-protein kinase TYRO 10) (Neurotrophic tyrosine kinase, receptor-related 3) (CD167b antigen). | |||||
|
DDR1_HUMAN
|
||||||
| θ value | 6.64225e-11 (rank : 34) | NC score | 0.121572 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 819 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08345, Q14196, Q16562, Q5ST12, Q9UD37 | Gene names | DDR1, CAK, EDDR1, RTK6, TRKE | |||
|
Domain Architecture |

|
|||||
| Description | Epithelial discoidin domain-containing receptor 1 precursor (EC 2.7.10.1) (Epithelial discoidin domain receptor 1) (Tyrosine kinase DDR) (Discoidin receptor tyrosine kinase) (Tyrosine-protein kinase CAK) (Cell adhesion kinase) (TRK E) (Protein-tyrosine kinase RTK 6) (HGK2) (CD167a antigen). | |||||
|
DCBD1_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 35) | NC score | 0.413157 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8N8Z6, Q5H992, Q8IYK5, Q8N7L9, Q96NH2 | Gene names | DCBLD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 1 precursor. | |||||
|
NRP1_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 36) | NC score | 0.349112 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P97333 | Gene names | Nrp1, Nrp | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-1 precursor (A5 protein). | |||||
|
CNTP4_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 37) | NC score | 0.349995 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99P47 | Gene names | Cntnap4, Caspr4 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 4 precursor (Cell recognition molecule Caspr4). | |||||
|
CNTP4_HUMAN
|
||||||
| θ value | 8.11959e-09 (rank : 38) | NC score | 0.346843 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9C0A0 | Gene names | CNTNAP4, CASPR4, KIAA1763 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 4 precursor (Cell recognition molecule Caspr4). | |||||
|
DDR1_MOUSE
|
||||||
| θ value | 1.06045e-08 (rank : 39) | NC score | 0.115395 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 818 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q03146 | Gene names | Ddr1, Cak, Eddr1, Mpk6 | |||
|
Domain Architecture |

|
|||||
| Description | Epithelial discoidin domain-containing receptor 1 precursor (EC 2.7.10.1) (Epithelial discoidin domain receptor 1) (Tyrosine kinase DDR) (Discoidin receptor tyrosine kinase) (Tyrosine-protein kinase CAK) (Cell adhesion kinase) (Protein-tyrosine kinase MPK-6) (CD167a antigen). | |||||
|
CNTP1_MOUSE
|
||||||
| θ value | 1.80886e-08 (rank : 40) | NC score | 0.340278 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O54991 | Gene names | Cntnap1, Nrxn4 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein 1 precursor (Caspr) (Caspr1) (Neurexin 4) (Neurexin IV) (Paranodin) (NCP1) (MHDNIV). | |||||
|
CBPA4_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 41) | NC score | 0.321561 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UI42, Q86UY9 | Gene names | CPA4, CPA3 | |||
|
Domain Architecture |
|
|||||
| Description | Carboxypeptidase A4 precursor (EC 3.4.17.-) (Carboxypeptidase A3). | |||||
|
CBPA6_MOUSE
|
||||||
| θ value | 2.36244e-08 (rank : 42) | NC score | 0.283374 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5U901, Q8BVD0 | Gene names | Cpa6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase A6 precursor (EC 3.4.17.1). | |||||
|
CNTP1_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 43) | NC score | 0.339766 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P78357 | Gene names | CNTNAP1, CASPR, NRXN4 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein 1 precursor (Caspr) (Caspr1) (Neurexin 4) (Neurexin IV) (p190). | |||||
|
CBPB2_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 44) | NC score | 0.292798 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9JHH6, Q5EBI3, Q9QZF0 | Gene names | Cpb2, Tafi | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase B2 precursor (EC 3.4.17.20) (Carboxypeptidase U) (CPU) (Thrombin-activable fibrinolysis inhibitor) (TAFI) (Carboxypeptidase R) (CPR). | |||||
|
CBPA6_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 45) | NC score | 0.269262 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N4T0, Q8NEX8, Q8TDE8, Q9NRI9 | Gene names | CPA6, CPAH | |||
|
Domain Architecture |
|
|||||
| Description | Carboxypeptidase A6 precursor (EC 3.4.17.1) (Carboxypeptidase B). | |||||
|
CBPA4_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 46) | NC score | 0.280554 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P8K8, Q8BMK6, Q9CTE6 | Gene names | Cpa4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase A4 precursor (EC 3.4.17.-). | |||||
|
CBPB2_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 47) | NC score | 0.249810 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96IY4, Q15114, Q5T9K1, Q5T9K2, Q9P2Y6 | Gene names | CPB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase B2 precursor (EC 3.4.17.20) (Carboxypeptidase U) (CPU) (Thrombin-activable fibrinolysis inhibitor) (TAFI) (Plasma carboxypeptidase B) (pCPB). | |||||
|
CBPA3_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 48) | NC score | 0.221833 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P15089 | Gene names | Cpa3 | |||
|
Domain Architecture |
|
|||||
| Description | Mast cell carboxypeptidase A precursor (EC 3.4.17.1) (MC-CPA) (Carboxypeptidase A3). | |||||
|
CBPA3_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 49) | NC score | 0.214515 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P15088, Q96E94 | Gene names | CPA3 | |||
|
Domain Architecture |
|
|||||
| Description | Mast cell carboxypeptidase A precursor (EC 3.4.17.1) (MC-CPA) (Carboxypeptidase A3). | |||||
|
CBPA2_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 50) | NC score | 0.256427 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P48052, Q96A12, Q96QN3 | Gene names | CPA2 | |||
|
Domain Architecture |
|
|||||
| Description | Carboxypeptidase A2 precursor (EC 3.4.17.15). | |||||
|
CBPB1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 51) | NC score | 0.188548 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15086, O60834, Q96BQ8 | Gene names | CPB1, CPB, PCPB | |||
|
Domain Architecture |
|
|||||
| Description | Carboxypeptidase B precursor (EC 3.4.17.2) (Pancreas-specific protein) (PASP). | |||||
|
CBPO_HUMAN
|
||||||
| θ value | 0.62314 (rank : 52) | NC score | 0.192764 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IVL8, Q2M277, Q7RTW7 | Gene names | CPO | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase O precursor (EC 3.4.17.-) (CPO). | |||||
|
ENTK_MOUSE
|
||||||
| θ value | 0.62314 (rank : 53) | NC score | 0.019539 (rank : 84) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97435 | Gene names | Prss7, Entk | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
TRIM8_MOUSE
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.007179 (rank : 90) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PJ2, Q8C508, Q8C700, Q8CGI2, Q99PQ4 | Gene names | Trim8, Gerp, Rnf27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
CSPG5_MOUSE
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.020761 (rank : 83) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
CBPA1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.162257 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P15085, Q9BS67 | Gene names | CPA1, CPA | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase A1 precursor (EC 3.4.17.1). | |||||
|
FOXC2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.002276 (rank : 93) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99958 | Gene names | FOXC2, FKHL14, MFH1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
PCD12_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.000290 (rank : 95) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NPG4, Q6UXB6, Q96KB8, Q9H7Y6, Q9H8E0 | Gene names | PCDH12 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin-12 precursor (Vascular cadherin-2) (Vascular endothelial cadherin-2) (VE-cadherin-2) (VE-cad-2). | |||||
|
CBPA1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.163008 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7TPZ8 | Gene names | Cpa1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase A1 precursor (EC 3.4.17.1). | |||||
|
MATN3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.001008 (rank : 94) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15232 | Gene names | MATN3 | |||
|
Domain Architecture |
|
|||||
| Description | Matrilin-3 precursor. | |||||
|
PRGC2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.008364 (rank : 88) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
PTC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.006859 (rank : 91) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6C5, O95341, O95856, Q5QP87, Q6UX14 | Gene names | PTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein patched homolog 2 (PTC2). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | -0.000537 (rank : 96) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TRIM8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.005538 (rank : 92) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BZR9, Q9C028 | Gene names | TRIM8, GERP, RNF27 | |||
|
Domain Architecture |
|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
TROAP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.010927 (rank : 87) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
ZN318_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.008191 (rank : 89) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VUA4, O94796, Q4G0E4, Q8NEM6, Q9UNU8, Q9Y2W9 | Gene names | ZNF318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Endocrine regulatory protein). | |||||
|
SSF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.016690 (rank : 86) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91YU8 | Gene names | Ppan | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of SWI4 1 homolog (Ssf-1) (Peter Pan homolog). | |||||
|
NKX12_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | -0.000979 (rank : 97) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42580, Q9JHF2, Q9JKW5, Q9JKW6, Q9JKW9, Q9JKX0, Q9JKX1 | Gene names | Nkx1-2, Nkx1.1, Sax1 | |||
|
Domain Architecture |

|
|||||
| Description | NK1 transcription factor-related protein 2 (Homeobox protein SAX-1) (NKX-1.1). | |||||
|
SSF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.016876 (rank : 85) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQ55, Q9BW97, Q9H170 | Gene names | PPAN, SSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of SWI4 1 homolog (Ssf-1) (Peter Pan homolog). | |||||
|
CBPA5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.186849 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WXQ8, Q86SE2, Q86XM3, Q8NA08 | Gene names | CPA5 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase A5 precursor (EC 3.4.17.1). | |||||
|
CBPA5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.219339 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R4H4 | Gene names | Cpa5 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase A5 precursor (EC 3.4.17.1). | |||||
|
CERU_HUMAN
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.078425 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P00450, Q14063, Q2PP18, Q9UKS4 | Gene names | CP | |||
|
Domain Architecture |
|
|||||
| Description | Ceruloplasmin precursor (EC 1.16.3.1) (Ferroxidase). | |||||
|
CERU_MOUSE
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.077628 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61147 | Gene names | Cp | |||
|
Domain Architecture |

|
|||||
| Description | Ceruloplasmin precursor (EC 1.16.3.1) (Ferroxidase). | |||||
|
CUBN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.050747 (rank : 79) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60494, Q5VTA6, Q96RU9 | Gene names | CUBN, IFCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor) (Intrinsic factor-vitamin B12 receptor) (460 kDa receptor) (Intestinal intrinsic factor receptor). | |||||
|
CUBN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.050484 (rank : 81) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JLB4 | Gene names | Cubn, Ifcr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor). | |||||
|
CUZD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.050408 (rank : 82) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86UP6, Q7Z660, Q7Z661, Q86SG1, Q86UP5, Q9HAR7 | Gene names | CUZD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Transmembrane protein UO-44). | |||||
|
DCBD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.098672 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D4J3, Q8R327, Q9D696 | Gene names | Dcbld1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 1 precursor. | |||||
|
GLPC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.088511 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P04921, Q92642 | Gene names | GYPC, GPC | |||
|
Domain Architecture |
|
|||||
| Description | Glycophorin C (PAS-2') (Glycoprotein beta) (GLPC) (Glycoconnectin) (Sialoglycoprotein D) (Glycophorin D) (GPD) (CD236 antigen). | |||||
|
HEPH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.077949 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BQS7, O75180, Q6UW45, Q9C058 | Gene names | HEPH, KIAA0698 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hephaestin precursor (EC 1.-.-.-). | |||||
|
HEPH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.079007 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z0Z4, Q6ZQ65, Q80Y80, Q8C4S2 | Gene names | Heph, Kiaa0698 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hephaestin precursor (EC 1.-.-.-). | |||||
|
MFRP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.068152 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BY79, Q335M3, Q96DQ9 | Gene names | MFRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane frizzled-related protein (Membrane-type frizzled-related protein). | |||||
|
MFRP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.069322 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K480, Q8BPP4 | Gene names | Mfrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane frizzled-related protein (Membrane-type frizzled-related protein). | |||||
|
NETO1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.053536 (rank : 78) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TDF5, Q86W85, Q8ND78, Q8TDF4 | Gene names | NETO1, BTCL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 1 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 1). | |||||
|
NETO1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.055084 (rank : 77) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4I7, Q80X39, Q8C4S3, Q8CCM2 | Gene names | Neto1, Btcl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 1 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 1). | |||||
|
NETO2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.057666 (rank : 73) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NC67, Q7Z381, Q8ND51, Q96SP4, Q9NVY8 | Gene names | NETO2, BTCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 2 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 2). | |||||
|
NETO2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.056608 (rank : 74) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BNJ6, Q5VM49, Q8C4Q8 | Gene names | Neto2, Btcl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 2 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 2). | |||||
|
NRX1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.073288 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9ULB1, O60323, Q53TJ9, Q53TQ1, Q9C079, Q9C080, Q9C081, Q9H3M2, Q9UDM6 | Gene names | NRXN1, KIAA0578 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
NRX1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.072363 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CS84, O88722, Q80Y87, Q8CHE6 | Gene names | Nrxn1, Kiaa0578 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
NRX2A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.061890 (rank : 72) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P2S2, Q9Y2D6 | Gene names | NRXN2, KIAA0921 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-2-alpha precursor (Neurexin II-alpha). | |||||
|
NRX3A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.071421 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y4C0, O95378, Q9NS47, Q9P1V3, Q9P1V6, Q9UIE2, Q9UIE3, Q9ULA5, Q9Y486 | Gene names | NRXN3, KIAA0743 | |||
|
Domain Architecture |
|
|||||
| Description | Neurexin-3-alpha precursor (Neurexin III-alpha). | |||||
|
PCOC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.063654 (rank : 71) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15113, O14550 | Gene names | PCOLCE, PCPE1 | |||
|
Domain Architecture |
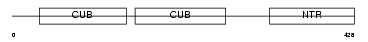
|
|||||
| Description | Procollagen C-endopeptidase enhancer 1 precursor (Procollagen COOH- terminal proteinase enhancer 1) (Procollagen C-proteinase enhancer 1) (PCPE-1) (Type I procollagen COOH-terminal proteinase enhancer) (Type 1 procollagen C-proteinase enhancer protein). | |||||
|
PCOC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.064853 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61398, O35113 | Gene names | Pcolce, Pcpe1 | |||
|
Domain Architecture |
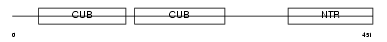
|
|||||
| Description | Procollagen C-endopeptidase enhancer 1 precursor (Procollagen COOH- terminal proteinase enhancer 1) (Procollagen C-proteinase enhancer 1) (PCPE-1) (Type I procollagen COOH-terminal proteinase enhancer) (Type 1 procollagen C-proteinase enhancer protein) (P14). | |||||
|
PCOC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.063716 (rank : 70) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UKZ9, Q9BRH3 | Gene names | PCOLCE2, PCPE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Procollagen C-endopeptidase enhancer 2 precursor (Procollagen COOH- terminal proteinase enhancer 2) (Procollagen C-proteinase enhancer 2) (PCPE-2). | |||||
|
PCOC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.065521 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4W6, Q3V1K6, Q9CX06 | Gene names | Pcolce2, Pcpe2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Procollagen C-endopeptidase enhancer 2 precursor (Procollagen COOH- terminal proteinase enhancer 2) (Procollagen C-proteinase enhancer 2) (PCPE-2). | |||||
|
PDGFD_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.050518 (rank : 80) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9GZP0, Q9BWV5 | Gene names | PDGFD, IEGF, SCDGFB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor D precursor (PDGF D) (Iris-expressed growth factor) (Spinal cord-derived growth factor B) (SCDGF-B). | |||||
|
TSG6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.056320 (rank : 76) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P98066, Q8WWI9 | Gene names | TNFAIP6, TSG6 | |||
|
Domain Architecture |
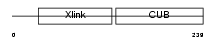
|
|||||
| Description | Tumor necrosis factor-inducible protein TSG-6 precursor (TNF- stimulated gene 6 protein) (Hyaluronate-binding protein). | |||||
|
TSG6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.056576 (rank : 75) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O08859 | Gene names | Tnfaip6, Tnfip6, Tsg6 | |||
|
Domain Architecture |
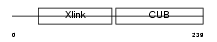
|
|||||
| Description | Tumor necrosis factor-inducible protein TSG-6 precursor (TNF- stimulated gene 6 protein). | |||||
|
CPXM1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q96SM3, Q6P4G8, Q6UW65, Q9NUB5 | Gene names | CPXM1, CPX1, CPXM | |||
|
Domain Architecture |

|
|||||
| Description | Probable carboxypeptidase X1 precursor (EC 3.4.17.-) (Metallocarboxypeptidase CPX-1). | |||||
|
CPXM1_MOUSE
|
||||||
| NC score | 0.997373 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9Z100, Q99LA3 | Gene names | Cpxm1, Cpx1, Cpxm | |||
|
Domain Architecture |

|
|||||
| Description | Probable carboxypeptidase X1 precursor (EC 3.4.17.-) (Metallocarboxypeptidase CPX-1). | |||||
|
CPXM2_MOUSE
|
||||||
| NC score | 0.990666 (rank : 3) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D2L5, O54860, Q8VDQ4 | Gene names | Cpxm2, Cpx2 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase-like protein X2 precursor. | |||||
|
CPXM2_HUMAN
|
||||||
| NC score | 0.990065 (rank : 4) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8N436 | Gene names | CPXM2 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase-like protein X2 precursor. | |||||
|
CBPN_HUMAN
|
||||||
| NC score | 0.879841 (rank : 5) | θ value | 2.07809e-105 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P15169 | Gene names | CPN1, ACBP | |||
|
Domain Architecture |
|
|||||
| Description | Carboxypeptidase N catalytic chain precursor (EC 3.4.17.3) (CPN) (Carboxypeptidase N polypeptide 1) (Carboxypeptidase N small subunit) (Lysine carboxypeptidase) (Arginine carboxypeptidase) (Kininase-1) (Serum carboxypeptidase N) (SCPN) (Anaphylatoxin inactivator) (Plasma carboxypeptidase B). | |||||
|
CBPN_MOUSE
|
||||||
| NC score | 0.879445 (rank : 6) | θ value | 1.75921e-104 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9JJN5, Q91WM9 | Gene names | Cpn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase N catalytic chain precursor (EC 3.4.17.3) (CPN) (Carboxypeptidase N polypeptide 1) (Carboxypeptidase N small subunit). | |||||
|
CBPE_MOUSE
|
||||||
| NC score | 0.878648 (rank : 7) | θ value | 1.39067e-109 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q00493, Q64439 | Gene names | Cpe | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase E precursor (EC 3.4.17.10) (CPE) (Carboxypeptidase H) (CPH) (Enkephalin convertase) (Prohormone-processing carboxypeptidase). | |||||
|
CBPE_HUMAN
|
||||||
| NC score | 0.878590 (rank : 8) | θ value | 3.0981e-109 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P16870, Q9UIU9 | Gene names | CPE | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase E precursor (EC 3.4.17.10) (CPE) (Carboxypeptidase H) (CPH) (Enkephalin convertase) (Prohormone-processing carboxypeptidase). | |||||
|
CBPM_MOUSE
|
||||||
| NC score | 0.857771 (rank : 9) | θ value | 1.5555e-68 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80V42, Q497S5, Q9CYH8 | Gene names | Cpm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase M precursor (EC 3.4.17.12) (CPM). | |||||
|
CBPM_HUMAN
|
||||||
| NC score | 0.857662 (rank : 10) | θ value | 1.6097e-65 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P14384, Q9H2K9 | Gene names | CPM | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase M precursor (EC 3.4.17.12) (CPM). | |||||
|
CBPD_HUMAN
|
||||||
| NC score | 0.844918 (rank : 11) | θ value | 2.38316e-93 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75976, O15377, Q86SH9, Q86XE6 | Gene names | CPD | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase D precursor (EC 3.4.17.22) (Metallocarboxypeptidase D) (gp180). | |||||
|
CBPD_MOUSE
|
||||||
| NC score | 0.844853 (rank : 12) | θ value | 2.81514e-94 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O89001 | Gene names | Cpd | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase D precursor (EC 3.4.17.22) (Metallocarboxypeptidase D) (gp180). | |||||
|
CBPZ_MOUSE
|
||||||
| NC score | 0.806005 (rank : 13) | θ value | 6.45994e-107 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R4V4 | Gene names | Cpz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase Z precursor (EC 3.4.17.-) (CPZ). | |||||
|
CBPZ_HUMAN
|
||||||
| NC score | 0.790153 (rank : 14) | θ value | 1.30163e-107 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q66K79, O00520, Q96MX2 | Gene names | CPZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase Z precursor (EC 3.4.17.-) (CPZ). | |||||
|
XLRS1_MOUSE
|
||||||
| NC score | 0.577953 (rank : 15) | θ value | 9.56915e-18 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Z1L4 | Gene names | Rs1, Rs1h, Xlrs1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoschisin precursor (X-linked juvenile retinoschisis protein homolog). | |||||
|
XLRS1_HUMAN
|
||||||
| NC score | 0.576018 (rank : 16) | θ value | 3.63628e-17 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15537 | Gene names | RS1, XLRS1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoschisin precursor (X-linked juvenile retinoschisis protein). | |||||
|
MFGM_HUMAN
|
||||||
| NC score | 0.518597 (rank : 17) | θ value | 7.82807e-20 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08431 | Gene names | MFGE8 | |||
|
Domain Architecture |

|
|||||
| Description | Lactadherin precursor (Milk fat globule-EGF factor 8) (MFG-E8) (HMFG) (Breast epithelial antigen BA46) (MFGM) [Contains: Lactadherin short form; Medin]. | |||||
|
MFGM_MOUSE
|
||||||
| NC score | 0.449669 (rank : 18) | θ value | 1.74391e-19 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P21956, P97800, Q3TBN5, Q9R1X9, Q9WTS3 | Gene names | Mfge8 | |||
|
Domain Architecture |

|
|||||
| Description | Lactadherin precursor (Milk fat globule-EGF factor 8) (MFG-E8) (MFGM) (Sperm surface protein SP47) (MP47). | |||||
|
FA5_HUMAN
|
||||||
| NC score | 0.445552 (rank : 19) | θ value | 3.07829e-16 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P12259, Q14285, Q6UPU6 | Gene names | F5 | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor V precursor (Activated protein C cofactor) [Contains: Coagulation factor V heavy chain; Coagulation factor V light chain]. | |||||
|
FA8_MOUSE
|
||||||
| NC score | 0.442625 (rank : 20) | θ value | 1.52774e-15 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q06194 | Gene names | F8, Cf8, F8c | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component). | |||||
|
FA8_HUMAN
|
||||||
| NC score | 0.442074 (rank : 21) | θ value | 1.16975e-15 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P00451 | Gene names | F8, F8C | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component) (Antihemophilic factor) (AHF) [Contains: Factor VIIIa heavy chain, 200 kDa isoform; Factor VIIIa heavy chain, 92 kDa isoform; Factor VIII B chain; Factor VIIIa light chain]. | |||||
|
DCBD2_MOUSE
|
||||||
| NC score | 0.424364 (rank : 22) | θ value | 4.16044e-13 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q91ZV3, Q8BKI4 | Gene names | Dcbld2, Esdn | |||
|
Domain Architecture |

|
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 2 precursor (Endothelial and smooth muscle cell-derived neuropilin-like protein). | |||||
|
DCBD1_HUMAN
|
||||||
| NC score | 0.413157 (rank : 23) | θ value | 1.133e-10 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8N8Z6, Q5H992, Q8IYK5, Q8N7L9, Q96NH2 | Gene names | DCBLD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 1 precursor. | |||||
|
DCBD2_HUMAN
|
||||||
| NC score | 0.410855 (rank : 24) | θ value | 4.16044e-13 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96PD2, Q8N6M4, Q8TDX2 | Gene names | DCBLD2, CLCP1, ESDN | |||
|
Domain Architecture |

|
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 2 precursor (Endothelial and smooth muscle cell-derived neuropilin-like protein) (CUB, LCCL and coagulation factor V/VIII-homology domains protein 1). | |||||
|
CNTP2_HUMAN
|
||||||
| NC score | 0.374746 (rank : 25) | θ value | 1.68911e-14 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UHC6, Q52LV1, Q5H9Q7, Q9UQ12 | Gene names | CNTNAP2, CASPR2, KIAA0868 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 2 precursor (Cell recognition molecule Caspr2). | |||||
|
CNTP2_MOUSE
|
||||||
| NC score | 0.368365 (rank : 26) | θ value | 9.90251e-15 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9CPW0, Q6P2K4, Q6ZQ31 | Gene names | Cntnap2, Kiaa0868 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Contactin-associated protein-like 2 precursor (Cell recognition molecule Caspr2). | |||||
|
EDIL3_HUMAN
|
||||||
| NC score | 0.367580 (rank : 27) | θ value | 6.62687e-19 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43854, O43855 | Gene names | EDIL3, DEL1 | |||
|
Domain Architecture |
|
|||||
| Description | EGF-like repeat and discoidin I-like domain-containing protein 3 precursor (EGF-like repeats and discoidin I-like domains protein 3) (Developmentally-regulated endothelial cell locus 1 protein) (Integrin-binding protein DEL1). | |||||
|
NRP2_MOUSE
|
||||||
| NC score | 0.364740 (rank : 28) | θ value | 4.91457e-14 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35375, O35373, O35374, O35376, O35377, O35378 | Gene names | Nrp2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-2 precursor (Vascular endothelial cell growth factor 165 receptor 2). | |||||
|
NRP2_HUMAN
|
||||||
| NC score | 0.364294 (rank : 29) | θ value | 1.68911e-14 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O60462, O14820, O14821 | Gene names | NRP2, VEGF165R2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-2 precursor (Vascular endothelial cell growth factor 165 receptor 2). | |||||
|
CNTP3_HUMAN
|
||||||
| NC score | 0.363582 (rank : 30) | θ value | 2.0648e-12 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9BZ76, Q9C0E9 | Gene names | CNTNAP3, CASPR3, KIAA1714 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 3 precursor (Cell recognition molecule Caspr3). | |||||
|
EDIL3_MOUSE
|
||||||
| NC score | 0.362083 (rank : 31) | θ value | 1.02238e-19 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35474, O35475 | Gene names | Edil3, Del1 | |||
|
Domain Architecture |
|
|||||
| Description | EGF-like repeat and discoidin I-like domain-containing protein 3 precursor (EGF-like repeats and discoidin I-like domains protein 3) (Developmentally-regulated endothelial cell locus 1 protein) (Integrin-binding protein DEL1). | |||||
|
NRP1_HUMAN
|
||||||
| NC score | 0.351479 (rank : 32) | θ value | 1.74796e-11 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O14786, O60461, Q96IH5 | Gene names | NRP1, NRP, VEGF165R | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-1 precursor (Vascular endothelial cell growth factor 165 receptor) (CD304 antigen). | |||||
|
CNTP4_MOUSE
|
||||||
| NC score | 0.349995 (rank : 33) | θ value | 9.59137e-10 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99P47 | Gene names | Cntnap4, Caspr4 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 4 precursor (Cell recognition molecule Caspr4). | |||||
|
NRP1_MOUSE
|
||||||
| NC score | 0.349112 (rank : 34) | θ value | 1.133e-10 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P97333 | Gene names | Nrp1, Nrp | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-1 precursor (A5 protein). | |||||
|
CNTP4_HUMAN
|
||||||
| NC score | 0.346843 (rank : 35) | θ value | 8.11959e-09 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9C0A0 | Gene names | CNTNAP4, CASPR4, KIAA1763 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 4 precursor (Cell recognition molecule Caspr4). | |||||
|
CNTP1_MOUSE
|
||||||
| NC score | 0.340278 (rank : 36) | θ value | 1.80886e-08 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O54991 | Gene names | Cntnap1, Nrxn4 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein 1 precursor (Caspr) (Caspr1) (Neurexin 4) (Neurexin IV) (Paranodin) (NCP1) (MHDNIV). | |||||
|
CNTP1_HUMAN
|
||||||
| NC score | 0.339766 (rank : 37) | θ value | 2.36244e-08 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P78357 | Gene names | CNTNAP1, CASPR, NRXN4 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein 1 precursor (Caspr) (Caspr1) (Neurexin 4) (Neurexin IV) (p190). | |||||
|
CBPA4_HUMAN
|
||||||
| NC score | 0.321561 (rank : 38) | θ value | 2.36244e-08 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UI42, Q86UY9 | Gene names | CPA4, CPA3 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase A4 precursor (EC 3.4.17.-) (Carboxypeptidase A3). | |||||
|
CBPB2_MOUSE
|
||||||
| NC score | 0.292798 (rank : 39) | θ value | 2.21117e-06 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9JHH6, Q5EBI3, Q9QZF0 | Gene names | Cpb2, Tafi | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase B2 precursor (EC 3.4.17.20) (Carboxypeptidase U) (CPU) (Thrombin-activable fibrinolysis inhibitor) (TAFI) (Carboxypeptidase R) (CPR). | |||||
|
CBPA6_MOUSE
|
||||||
| NC score | 0.283374 (rank : 40) | θ value | 2.36244e-08 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5U901, Q8BVD0 | Gene names | Cpa6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase A6 precursor (EC 3.4.17.1). | |||||
|
CBPA4_MOUSE
|
||||||
| NC score | 0.280554 (rank : 41) | θ value | 2.44474e-05 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P8K8, Q8BMK6, Q9CTE6 | Gene names | Cpa4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase A4 precursor (EC 3.4.17.-). | |||||
|
CBPA6_HUMAN
|
||||||
| NC score | 0.269262 (rank : 42) | θ value | 6.43352e-06 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N4T0, Q8NEX8, Q8TDE8, Q9NRI9 | Gene names | CPA6, CPAH | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase A6 precursor (EC 3.4.17.1) (Carboxypeptidase B). | |||||
|
CBPA2_HUMAN
|
||||||
| NC score | 0.256427 (rank : 43) | θ value | 0.00390308 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P48052, Q96A12, Q96QN3 | Gene names | CPA2 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase A2 precursor (EC 3.4.17.15). | |||||
|
CBPB2_HUMAN
|
||||||
| NC score | 0.249810 (rank : 44) | θ value | 5.44631e-05 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96IY4, Q15114, Q5T9K1, Q5T9K2, Q9P2Y6 | Gene names | CPB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase B2 precursor (EC 3.4.17.20) (Carboxypeptidase U) (CPU) (Thrombin-activable fibrinolysis inhibitor) (TAFI) (Plasma carboxypeptidase B) (pCPB). | |||||
|
CBPA3_MOUSE
|
||||||
| NC score | 0.221833 (rank : 45) | θ value | 0.000158464 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P15089 | Gene names | Cpa3 | |||
|
Domain Architecture |

|
|||||
| Description | Mast cell carboxypeptidase A precursor (EC 3.4.17.1) (MC-CPA) (Carboxypeptidase A3). | |||||
|
CBPA5_MOUSE
|
||||||
| NC score | 0.219339 (rank : 46) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R4H4 | Gene names | Cpa5 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase A5 precursor (EC 3.4.17.1). | |||||
|
CBPA3_HUMAN
|
||||||
| NC score | 0.214515 (rank : 47) | θ value | 0.00228821 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P15088, Q96E94 | Gene names | CPA3 | |||
|
Domain Architecture |

|
|||||
| Description | Mast cell carboxypeptidase A precursor (EC 3.4.17.1) (MC-CPA) (Carboxypeptidase A3). | |||||
|
CBPO_HUMAN
|
||||||
| NC score | 0.192764 (rank : 48) | θ value | 0.62314 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IVL8, Q2M277, Q7RTW7 | Gene names | CPO | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase O precursor (EC 3.4.17.-) (CPO). | |||||
|
CBPB1_HUMAN
|
||||||
| NC score | 0.188548 (rank : 49) | θ value | 0.0736092 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15086, O60834, Q96BQ8 | Gene names | CPB1, CPB, PCPB | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase B precursor (EC 3.4.17.2) (Pancreas-specific protein) (PASP). | |||||
|
CBPA5_HUMAN
|
||||||
| NC score | 0.186849 (rank : 50) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WXQ8, Q86SE2, Q86XM3, Q8NA08 | Gene names | CPA5 | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase A5 precursor (EC 3.4.17.1). | |||||
|
CBPA1_MOUSE
|
||||||
| NC score | 0.163008 (rank : 51) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7TPZ8 | Gene names | Cpa1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase A1 precursor (EC 3.4.17.1). | |||||
|
CBPA1_HUMAN
|
||||||
| NC score | 0.162257 (rank : 52) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P15085, Q9BS67 | Gene names | CPA1, CPA | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase A1 precursor (EC 3.4.17.1). | |||||
|
DDR2_MOUSE
|
||||||
| NC score | 0.131890 (rank : 53) | θ value | 3.89403e-11 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q62371 | Gene names | Ddr2, Ntrkr3, Tkt, Tyro10 | |||
|
Domain Architecture |

|
|||||
| Description | Discoidin domain-containing receptor 2 precursor (EC 2.7.10.1) (Discoidin domain receptor 2) (Receptor protein-tyrosine kinase TKT) (Tyrosine-protein kinase TYRO 10) (Neurotrophic tyrosine kinase, receptor-related 3) (CD167b antigen). | |||||
|
DDR2_HUMAN
|
||||||
| NC score | 0.130787 (rank : 54) | θ value | 3.89403e-11 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 818 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q16832 | Gene names | DDR2, NTRKR3, TKT, TYRO10 | |||
|
Domain Architecture |

|
|||||
| Description | Discoidin domain-containing receptor 2 precursor (EC 2.7.10.1) (Discoidin domain receptor 2) (Receptor protein-tyrosine kinase TKT) (Tyrosine-protein kinase TYRO 10) (Neurotrophic tyrosine kinase, receptor-related 3) (CD167b antigen). | |||||
|
DDR1_HUMAN
|
||||||
| NC score | 0.121572 (rank : 55) | θ value | 6.64225e-11 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 819 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08345, Q14196, Q16562, Q5ST12, Q9UD37 | Gene names | DDR1, CAK, EDDR1, RTK6, TRKE | |||
|
Domain Architecture |

|
|||||
| Description | Epithelial discoidin domain-containing receptor 1 precursor (EC 2.7.10.1) (Epithelial discoidin domain receptor 1) (Tyrosine kinase DDR) (Discoidin receptor tyrosine kinase) (Tyrosine-protein kinase CAK) (Cell adhesion kinase) (TRK E) (Protein-tyrosine kinase RTK 6) (HGK2) (CD167a antigen). | |||||
|
DDR1_MOUSE
|
||||||
| NC score | 0.115395 (rank : 56) | θ value | 1.06045e-08 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 818 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q03146 | Gene names | Ddr1, Cak, Eddr1, Mpk6 | |||
|
Domain Architecture |

|
|||||
| Description | Epithelial discoidin domain-containing receptor 1 precursor (EC 2.7.10.1) (Epithelial discoidin domain receptor 1) (Tyrosine kinase DDR) (Discoidin receptor tyrosine kinase) (Tyrosine-protein kinase CAK) (Cell adhesion kinase) (Protein-tyrosine kinase MPK-6) (CD167a antigen). | |||||
|
DCBD1_MOUSE
|
||||||
| NC score | 0.098672 (rank : 57) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D4J3, Q8R327, Q9D696 | Gene names | Dcbld1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 1 precursor. | |||||
|
GLPC_HUMAN
|
||||||
| NC score | 0.088511 (rank : 58) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P04921, Q92642 | Gene names | GYPC, GPC | |||
|
Domain Architecture |

|
|||||
| Description | Glycophorin C (PAS-2') (Glycoprotein beta) (GLPC) (Glycoconnectin) (Sialoglycoprotein D) (Glycophorin D) (GPD) (CD236 antigen). | |||||
|
HEPH_MOUSE
|
||||||
| NC score | 0.079007 (rank : 59) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z0Z4, Q6ZQ65, Q80Y80, Q8C4S2 | Gene names | Heph, Kiaa0698 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hephaestin precursor (EC 1.-.-.-). | |||||
|
CERU_HUMAN
|
||||||
| NC score | 0.078425 (rank : 60) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P00450, Q14063, Q2PP18, Q9UKS4 | Gene names | CP | |||
|
Domain Architecture |
|
|||||
| Description | Ceruloplasmin precursor (EC 1.16.3.1) (Ferroxidase). | |||||
|
HEPH_HUMAN
|
||||||
| NC score | 0.077949 (rank : 61) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BQS7, O75180, Q6UW45, Q9C058 | Gene names | HEPH, KIAA0698 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hephaestin precursor (EC 1.-.-.-). | |||||
|
CERU_MOUSE
|
||||||
| NC score | 0.077628 (rank : 62) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61147 | Gene names | Cp | |||
|
Domain Architecture |

|
|||||
| Description | Ceruloplasmin precursor (EC 1.16.3.1) (Ferroxidase). | |||||
|
NRX1A_HUMAN
|
||||||
| NC score | 0.073288 (rank : 63) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9ULB1, O60323, Q53TJ9, Q53TQ1, Q9C079, Q9C080, Q9C081, Q9H3M2, Q9UDM6 | Gene names | NRXN1, KIAA0578 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
NRX1A_MOUSE
|
||||||
| NC score | 0.072363 (rank : 64) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CS84, O88722, Q80Y87, Q8CHE6 | Gene names | Nrxn1, Kiaa0578 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
NRX3A_HUMAN
|
||||||
| NC score | 0.071421 (rank : 65) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y4C0, O95378, Q9NS47, Q9P1V3, Q9P1V6, Q9UIE2, Q9UIE3, Q9ULA5, Q9Y486 | Gene names | NRXN3, KIAA0743 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-3-alpha precursor (Neurexin III-alpha). | |||||
|
MFRP_MOUSE
|
||||||
| NC score | 0.069322 (rank : 66) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K480, Q8BPP4 | Gene names | Mfrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane frizzled-related protein (Membrane-type frizzled-related protein). | |||||
|
MFRP_HUMAN
|
||||||
| NC score | 0.068152 (rank : 67) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BY79, Q335M3, Q96DQ9 | Gene names | MFRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane frizzled-related protein (Membrane-type frizzled-related protein). | |||||
|
PCOC2_MOUSE
|
||||||
| NC score | 0.065521 (rank : 68) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4W6, Q3V1K6, Q9CX06 | Gene names | Pcolce2, Pcpe2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Procollagen C-endopeptidase enhancer 2 precursor (Procollagen COOH- terminal proteinase enhancer 2) (Procollagen C-proteinase enhancer 2) (PCPE-2). | |||||
|
PCOC1_MOUSE
|
||||||
| NC score | 0.064853 (rank : 69) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61398, O35113 | Gene names | Pcolce, Pcpe1 | |||
|
Domain Architecture |

|
|||||
| Description | Procollagen C-endopeptidase enhancer 1 precursor (Procollagen COOH- terminal proteinase enhancer 1) (Procollagen C-proteinase enhancer 1) (PCPE-1) (Type I procollagen COOH-terminal proteinase enhancer) (Type 1 procollagen C-proteinase enhancer protein) (P14). | |||||
|
PCOC2_HUMAN
|
||||||
| NC score | 0.063716 (rank : 70) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UKZ9, Q9BRH3 | Gene names | PCOLCE2, PCPE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Procollagen C-endopeptidase enhancer 2 precursor (Procollagen COOH- terminal proteinase enhancer 2) (Procollagen C-proteinase enhancer 2) (PCPE-2). | |||||
|
PCOC1_HUMAN
|
||||||
| NC score | 0.063654 (rank : 71) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15113, O14550 | Gene names | PCOLCE, PCPE1 | |||
|
Domain Architecture |

|
|||||
| Description | Procollagen C-endopeptidase enhancer 1 precursor (Procollagen COOH- terminal proteinase enhancer 1) (Procollagen C-proteinase enhancer 1) (PCPE-1) (Type I procollagen COOH-terminal proteinase enhancer) (Type 1 procollagen C-proteinase enhancer protein). | |||||
|
NRX2A_HUMAN
|
||||||
| NC score | 0.061890 (rank : 72) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P2S2, Q9Y2D6 | Gene names | NRXN2, KIAA0921 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-2-alpha precursor (Neurexin II-alpha). | |||||
|
NETO2_HUMAN
|
||||||
| NC score | 0.057666 (rank : 73) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NC67, Q7Z381, Q8ND51, Q96SP4, Q9NVY8 | Gene names | NETO2, BTCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 2 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 2). | |||||
|
NETO2_MOUSE
|
||||||
| NC score | 0.056608 (rank : 74) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BNJ6, Q5VM49, Q8C4Q8 | Gene names | Neto2, Btcl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 2 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 2). | |||||
|
TSG6_MOUSE
|
||||||
| NC score | 0.056576 (rank : 75) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O08859 | Gene names | Tnfaip6, Tnfip6, Tsg6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor-inducible protein TSG-6 precursor (TNF- stimulated gene 6 protein). | |||||
|
TSG6_HUMAN
|
||||||
| NC score | 0.056320 (rank : 76) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P98066, Q8WWI9 | Gene names | TNFAIP6, TSG6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor-inducible protein TSG-6 precursor (TNF- stimulated gene 6 protein) (Hyaluronate-binding protein). | |||||
|
NETO1_MOUSE
|
||||||
| NC score | 0.055084 (rank : 77) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4I7, Q80X39, Q8C4S3, Q8CCM2 | Gene names | Neto1, Btcl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 1 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 1). | |||||
|
NETO1_HUMAN
|
||||||
| NC score | 0.053536 (rank : 78) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TDF5, Q86W85, Q8ND78, Q8TDF4 | Gene names | NETO1, BTCL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 1 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 1). | |||||
|
CUBN_HUMAN
|
||||||
| NC score | 0.050747 (rank : 79) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60494, Q5VTA6, Q96RU9 | Gene names | CUBN, IFCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor) (Intrinsic factor-vitamin B12 receptor) (460 kDa receptor) (Intestinal intrinsic factor receptor). | |||||
|
PDGFD_HUMAN
|
||||||
| NC score | 0.050518 (rank : 80) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9GZP0, Q9BWV5 | Gene names | PDGFD, IEGF, SCDGFB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor D precursor (PDGF D) (Iris-expressed growth factor) (Spinal cord-derived growth factor B) (SCDGF-B). | |||||
|
CUBN_MOUSE
|
||||||
| NC score | 0.050484 (rank : 81) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JLB4 | Gene names | Cubn, Ifcr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor). | |||||
|
CUZD1_HUMAN
|
||||||
| NC score | 0.050408 (rank : 82) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86UP6, Q7Z660, Q7Z661, Q86SG1, Q86UP5, Q9HAR7 | Gene names | CUZD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Transmembrane protein UO-44). | |||||
|
CSPG5_MOUSE
|
||||||
| NC score | 0.020761 (rank : 83) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
ENTK_MOUSE
|
||||||
| NC score | 0.019539 (rank : 84) | θ value | 0.62314 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97435 | Gene names | Prss7, Entk | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
SSF1_HUMAN
|
||||||
| NC score | 0.016876 (rank : 85) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQ55, Q9BW97, Q9H170 | Gene names | PPAN, SSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of SWI4 1 homolog (Ssf-1) (Peter Pan homolog). | |||||
|
SSF1_MOUSE
|
||||||
| NC score | 0.016690 (rank : 86) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91YU8 | Gene names | Ppan | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of SWI4 1 homolog (Ssf-1) (Peter Pan homolog). | |||||
|
TROAP_HUMAN
|
||||||
| NC score | 0.010927 (rank : 87) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
PRGC2_MOUSE
|
||||||
| NC score | 0.008364 (rank : 88) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
ZN318_HUMAN
|
||||||
| NC score | 0.008191 (rank : 89) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VUA4, O94796, Q4G0E4, Q8NEM6, Q9UNU8, Q9Y2W9 | Gene names | ZNF318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Endocrine regulatory protein). | |||||
|
TRIM8_MOUSE
|
||||||
| NC score | 0.007179 (rank : 90) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PJ2, Q8C508, Q8C700, Q8CGI2, Q99PQ4 | Gene names | Trim8, Gerp, Rnf27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
PTC2_HUMAN
|
||||||
| NC score | 0.006859 (rank : 91) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6C5, O95341, O95856, Q5QP87, Q6UX14 | Gene names | PTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein patched homolog 2 (PTC2). | |||||
|
TRIM8_HUMAN
|
||||||
| NC score | 0.005538 (rank : 92) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BZR9, Q9C028 | Gene names | TRIM8, GERP, RNF27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
FOXC2_HUMAN
|
||||||
| NC score | 0.002276 (rank : 93) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99958 | Gene names | FOXC2, FKHL14, MFH1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
MATN3_HUMAN
|
||||||
| NC score | 0.001008 (rank : 94) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15232 | Gene names | MATN3 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-3 precursor. | |||||
|
PCD12_HUMAN
|
||||||
| NC score | 0.000290 (rank : 95) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NPG4, Q6UXB6, Q96KB8, Q9H7Y6, Q9H8E0 | Gene names | PCDH12 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin-12 precursor (Vascular cadherin-2) (Vascular endothelial cadherin-2) (VE-cadherin-2) (VE-cad-2). | |||||
|
TARA_HUMAN
|
||||||
| NC score | -0.000537 (rank : 96) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
NKX12_MOUSE
|
||||||
| NC score | -0.000979 (rank : 97) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42580, Q9JHF2, Q9JKW5, Q9JKW6, Q9JKW9, Q9JKX0, Q9JKX1 | Gene names | Nkx1-2, Nkx1.1, Sax1 | |||
|
Domain Architecture |

|
|||||
| Description | NK1 transcription factor-related protein 2 (Homeobox protein SAX-1) (NKX-1.1). | |||||