Please be patient as the page loads
|
CPVL_HUMAN
|
||||||
| SwissProt Accessions | Q9H3G5, Q6UX20, Q8NBL7, Q96AR7, Q9HB41 | Gene names | CPVL, VLP | |||
|
Domain Architecture |

|
|||||
| Description | Probable serine carboxypeptidase CPVL precursor (EC 3.4.16.-) (Carboxypeptidase, vitellogenic-like) (Vitellogenic carboxypeptidase- like protein) (VCP-like protein) (HVLP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CPVL_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H3G5, Q6UX20, Q8NBL7, Q96AR7, Q9HB41 | Gene names | CPVL, VLP | |||
|
Domain Architecture |

|
|||||
| Description | Probable serine carboxypeptidase CPVL precursor (EC 3.4.16.-) (Carboxypeptidase, vitellogenic-like) (Vitellogenic carboxypeptidase- like protein) (VCP-like protein) (HVLP). | |||||
|
PPGB_HUMAN
|
||||||
| θ value | 2.58187e-47 (rank : 2) | NC score | 0.855758 (rank : 2) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10619, Q5JZH1, Q96KJ2, Q9BR08, Q9BW68 | Gene names | PPGB | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal protective protein precursor (EC 3.4.16.5) (Cathepsin A) (Carboxypeptidase C) (Protective protein for beta-galactosidase) [Contains: Lysosomal protective protein 32 kDa chain; Lysosomal protective protein 20 kDa chain]. | |||||
|
PPGB_MOUSE
|
||||||
| θ value | 1.08474e-45 (rank : 3) | NC score | 0.850307 (rank : 3) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P16675, Q8VEF6 | Gene names | Ppgb | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal protective protein precursor (EC 3.4.16.5) (Cathepsin A) (Carboxypeptidase C) (Protective protein for beta-galactosidase) [Contains: Lysosomal protective protein 32 kDa chain; Lysosomal protective protein 20 kDa chain]. | |||||
|
RISC_MOUSE
|
||||||
| θ value | 2.05049e-36 (rank : 4) | NC score | 0.817003 (rank : 4) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q920A5 | Gene names | Scpep1, Risc | |||
|
Domain Architecture |

|
|||||
| Description | Retinoid-inducible serine carboxypeptidase precursor (EC 3.4.16.-) (Serine carboxypeptidase 1). | |||||
|
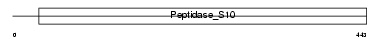
RISC_HUMAN
|
||||||
| θ value | 2.50655e-34 (rank : 5) | NC score | 0.799210 (rank : 5) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9HB40, Q96A94, Q9H3F0 | Gene names | SCPEP1, RISC, SCP1 | |||
|
Domain Architecture |
|
|||||
| Description | Retinoid-inducible serine carboxypeptidase precursor (EC 3.4.16.-) (Serine carboxypeptidase 1). | |||||
|
ENOG_MOUSE
|
||||||
| θ value | 5.27518 (rank : 6) | NC score | 0.018258 (rank : 7) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17183 | Gene names | Eno2, Eno-2 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-enolase (EC 4.2.1.11) (2-phospho-D-glycerate hydro-lyase) (Neural enolase) (Neuron-specific enolase) (NSE) (Enolase 2). | |||||
|
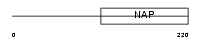
TSPY1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 7) | NC score | 0.018297 (rank : 6) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q01534, O00216, P09002, Q9UNN7 | Gene names | TSPY1, TSPY | |||
|
Domain Architecture |
|
|||||
| Description | Testis-specific Y-encoded protein 1. | |||||
|
ECE2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 8) | NC score | 0.009434 (rank : 9) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60344, Q96NX3, Q96NX4 | Gene names | ECE2, KIAA0604 | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme 2 (EC 3.4.24.71) (ECE-2). | |||||
|
ENOG_HUMAN
|
||||||
| θ value | 6.88961 (rank : 9) | NC score | 0.017207 (rank : 8) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P09104, Q96J33 | Gene names | ENO2 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-enolase (EC 4.2.1.11) (2-phospho-D-glycerate hydro-lyase) (Neural enolase) (Neuron-specific enolase) (NSE) (Enolase 2). | |||||
|
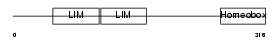
LHX2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 10) | NC score | 0.007943 (rank : 10) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z0S2 | Gene names | Lhx2 | |||
|
Domain Architecture |
|
|||||
| Description | LIM/homeobox protein Lhx2. | |||||
|
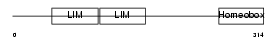
LHX2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 11) | NC score | 0.007262 (rank : 11) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50458, O95860, Q8N1Z3 | Gene names | LHX2, LH2 | |||
|
Domain Architecture |
|
|||||
| Description | LIM/homeobox protein Lhx2 (Homeobox protein LH-2). | |||||
|
CPVL_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H3G5, Q6UX20, Q8NBL7, Q96AR7, Q9HB41 | Gene names | CPVL, VLP | |||
|
Domain Architecture |

|
|||||
| Description | Probable serine carboxypeptidase CPVL precursor (EC 3.4.16.-) (Carboxypeptidase, vitellogenic-like) (Vitellogenic carboxypeptidase- like protein) (VCP-like protein) (HVLP). | |||||
|
PPGB_HUMAN
|
||||||
| NC score | 0.855758 (rank : 2) | θ value | 2.58187e-47 (rank : 2) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10619, Q5JZH1, Q96KJ2, Q9BR08, Q9BW68 | Gene names | PPGB | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal protective protein precursor (EC 3.4.16.5) (Cathepsin A) (Carboxypeptidase C) (Protective protein for beta-galactosidase) [Contains: Lysosomal protective protein 32 kDa chain; Lysosomal protective protein 20 kDa chain]. | |||||
|
PPGB_MOUSE
|
||||||
| NC score | 0.850307 (rank : 3) | θ value | 1.08474e-45 (rank : 3) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P16675, Q8VEF6 | Gene names | Ppgb | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal protective protein precursor (EC 3.4.16.5) (Cathepsin A) (Carboxypeptidase C) (Protective protein for beta-galactosidase) [Contains: Lysosomal protective protein 32 kDa chain; Lysosomal protective protein 20 kDa chain]. | |||||
|
RISC_MOUSE
|
||||||
| NC score | 0.817003 (rank : 4) | θ value | 2.05049e-36 (rank : 4) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q920A5 | Gene names | Scpep1, Risc | |||
|
Domain Architecture |

|
|||||
| Description | Retinoid-inducible serine carboxypeptidase precursor (EC 3.4.16.-) (Serine carboxypeptidase 1). | |||||
|
RISC_HUMAN
|
||||||
| NC score | 0.799210 (rank : 5) | θ value | 2.50655e-34 (rank : 5) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9HB40, Q96A94, Q9H3F0 | Gene names | SCPEP1, RISC, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoid-inducible serine carboxypeptidase precursor (EC 3.4.16.-) (Serine carboxypeptidase 1). | |||||
|
TSPY1_HUMAN
|
||||||
| NC score | 0.018297 (rank : 6) | θ value | 5.27518 (rank : 7) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q01534, O00216, P09002, Q9UNN7 | Gene names | TSPY1, TSPY | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific Y-encoded protein 1. | |||||
|
ENOG_MOUSE
|
||||||
| NC score | 0.018258 (rank : 7) | θ value | 5.27518 (rank : 6) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17183 | Gene names | Eno2, Eno-2 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-enolase (EC 4.2.1.11) (2-phospho-D-glycerate hydro-lyase) (Neural enolase) (Neuron-specific enolase) (NSE) (Enolase 2). | |||||
|
ENOG_HUMAN
|
||||||
| NC score | 0.017207 (rank : 8) | θ value | 6.88961 (rank : 9) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P09104, Q96J33 | Gene names | ENO2 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-enolase (EC 4.2.1.11) (2-phospho-D-glycerate hydro-lyase) (Neural enolase) (Neuron-specific enolase) (NSE) (Enolase 2). | |||||
|
ECE2_HUMAN
|
||||||
| NC score | 0.009434 (rank : 9) | θ value | 6.88961 (rank : 8) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60344, Q96NX3, Q96NX4 | Gene names | ECE2, KIAA0604 | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme 2 (EC 3.4.24.71) (ECE-2). | |||||
|
LHX2_MOUSE
|
||||||
| NC score | 0.007943 (rank : 10) | θ value | 6.88961 (rank : 10) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z0S2 | Gene names | Lhx2 | |||
|
Domain Architecture |

|
|||||
| Description | LIM/homeobox protein Lhx2. | |||||
|
LHX2_HUMAN
|
||||||
| NC score | 0.007262 (rank : 11) | θ value | 8.99809 (rank : 11) | |||
| Query Neighborhood Hits | 11 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50458, O95860, Q8N1Z3 | Gene names | LHX2, LH2 | |||
|
Domain Architecture |

|
|||||
| Description | LIM/homeobox protein Lhx2 (Homeobox protein LH-2). | |||||