Please be patient as the page loads
|
CEBPE_MOUSE
|
||||||
| SwissProt Accessions | Q6PZD9 | Gene names | Cebpe | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CCAAT/enhancer-binding protein epsilon (C/EBP epsilon). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CEBPE_HUMAN
|
||||||
| θ value | 1.67649e-155 (rank : 1) | NC score | 0.945641 (rank : 2) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q15744, Q15745, Q8IYI2, Q99803 | Gene names | CEBPE | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein epsilon (C/EBP epsilon). | |||||
|
CEBPE_MOUSE
|
||||||
| θ value | 1.42253e-146 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 108 | |
| SwissProt Accessions | Q6PZD9 | Gene names | Cebpe | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CCAAT/enhancer-binding protein epsilon (C/EBP epsilon). | |||||
|
CEBPA_HUMAN
|
||||||
| θ value | 1.28434e-38 (rank : 3) | NC score | 0.890027 (rank : 4) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P49715, P78319 | Gene names | CEBPA | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein alpha (C/EBP alpha). | |||||
|
CEBPA_MOUSE
|
||||||
| θ value | 2.86122e-38 (rank : 4) | NC score | 0.894859 (rank : 3) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P53566, Q91XB6 | Gene names | Cebpa | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein alpha (C/EBP alpha). | |||||
|
CEBPB_HUMAN
|
||||||
| θ value | 2.51237e-26 (rank : 5) | NC score | 0.834398 (rank : 6) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P17676, Q96IH2, Q9H4Z5 | Gene names | CEBPB, TCF5 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein beta (C/EBP beta) (Nuclear factor NF- IL6) (Transcription factor 5). | |||||
|
CEBPB_MOUSE
|
||||||
| θ value | 1.99067e-23 (rank : 6) | NC score | 0.850804 (rank : 5) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P28033 | Gene names | Cebpb | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein beta (C/EBP beta) (Interleukin-6- dependent-binding protein) (IL-6DBP) (Liver-enriched transcriptional activator) (LAP) (AGP/EBP). | |||||
|
CEBPD_HUMAN
|
||||||
| θ value | 1.99067e-23 (rank : 7) | NC score | 0.827214 (rank : 7) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49716, Q14937 | Gene names | CEBPD | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein delta (C/EBP delta) (Nuclear factor NF- IL6-beta) (NF-IL6-beta). | |||||
|
CEBPD_MOUSE
|
||||||
| θ value | 1.29031e-22 (rank : 8) | NC score | 0.821607 (rank : 8) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q00322 | Gene names | Cebpd, Crp3 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein delta (C/EBP delta) (C/EBP-related protein 3). | |||||
|
CEBPG_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 9) | NC score | 0.726445 (rank : 10) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P53567, Q5U052 | Gene names | CEBPG | |||
|
Domain Architecture |
|
|||||
| Description | CCAAT/enhancer-binding protein gamma (C/EBP gamma). | |||||
|
CEBPG_MOUSE
|
||||||
| θ value | 1.02475e-11 (rank : 10) | NC score | 0.734002 (rank : 9) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P53568 | Gene names | Cebpg | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein gamma (C/EBP gamma) (Immunoglobulin enhancer-binding protein 1) (IG/EBP-1) (Granulocyte colony-stimulating factor promoter element 1-binding protein) (GPE1-binding protein) (GPE1-BP). | |||||
|
CREB3_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 11) | NC score | 0.239362 (rank : 18) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61817, Q99M21, Q9CVK9 | Gene names | Creb3, Lzip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Transcription factor LZIP). | |||||
|
DDIT3_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 12) | NC score | 0.422276 (rank : 11) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P35639, Q91YW9 | Gene names | Ddit3, Chop, Gadd153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
DDIT3_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 13) | NC score | 0.374760 (rank : 12) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |
|
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
JUNB_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 14) | NC score | 0.172180 (rank : 29) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P09450, Q8C2G9 | Gene names | Junb, Jun-b | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor jun-B. | |||||
|
CREB3_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 15) | NC score | 0.183619 (rank : 28) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O43889, O14671, O14919, Q96GK8, Q9H2W3, Q9UE77 | Gene names | CREB3, LZIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Luman protein) (Transcription factor LZIP-alpha). | |||||
|
ATF6A_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 16) | NC score | 0.116445 (rank : 42) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P18850, O15139, Q5VW62, Q6IPB5, Q9UEC9 | Gene names | ATF6 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-6 alpha (Activating transcription factor 6 alpha) (ATF6-alpha). | |||||
|
HLF_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 17) | NC score | 0.240015 (rank : 16) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16534 | Gene names | HLF | |||
|
Domain Architecture |

|
|||||
| Description | Hepatic leukemia factor. | |||||
|
HLF_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 18) | NC score | 0.239599 (rank : 17) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BW74, Q6PF83 | Gene names | Hlf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatic leukemia factor. | |||||
|
CREB5_HUMAN
|
||||||
| θ value | 0.125558 (rank : 19) | NC score | 0.144276 (rank : 38) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q02930, Q05886, Q06246 | Gene names | CREB5, CREBPA | |||
|
Domain Architecture |

|
|||||
| Description | cAMP response element-binding protein 5 (CRE-BPa). | |||||
|
JUNB_HUMAN
|
||||||
| θ value | 0.125558 (rank : 20) | NC score | 0.166705 (rank : 30) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P17275, Q96GH3 | Gene names | JUNB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-B. | |||||
|
ACHA4_MOUSE
|
||||||
| θ value | 0.21417 (rank : 21) | NC score | 0.022249 (rank : 88) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70174, Q8BHE9, Q8VI10, Q9ET51 | Gene names | Chrna4, Acra4 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal acetylcholine receptor protein subunit alpha-4 precursor. | |||||
|
BATF_HUMAN
|
||||||
| θ value | 0.21417 (rank : 22) | NC score | 0.302528 (rank : 13) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16520 | Gene names | BATF | |||
|
Domain Architecture |
|
|||||
| Description | ATF-like basic leucine zipper transcriptional factor B-ATF (SF-HT- activated gene 2) (SFA-2). | |||||
|
BATF_MOUSE
|
||||||
| θ value | 0.21417 (rank : 23) | NC score | 0.288558 (rank : 14) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O35284 | Gene names | Batf | |||
|
Domain Architecture |
|
|||||
| Description | ATF-like basic leucine zipper transcription factor B-ATF. | |||||
|
CO1A2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 24) | NC score | 0.053209 (rank : 56) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q01149, Q8CGA5 | Gene names | Col1a2, Cola2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
KCNH2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 25) | NC score | 0.025920 (rank : 85) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
KLF2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 26) | NC score | 0.007898 (rank : 111) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5W3, Q9UJS5, Q9UKR6 | Gene names | KLF2, LKLF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 2 (Lung krueppel-like factor). | |||||
|
NGN3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 27) | NC score | 0.071604 (rank : 49) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P70661 | Gene names | Neurog3, Ath4b, Atoh5, Ngn3 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 3 (Atonal protein homolog 5) (Helix-loop-helix protein mATH-4B) (MATH4B). | |||||
|
XBP1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 28) | NC score | 0.215718 (rank : 23) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P17861, Q8WYK6, Q969P1, Q96BD7 | Gene names | XBP1, TREB5, XBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X box-binding protein 1 (XBP-1) (Tax-responsive element-binding protein 5). | |||||
|
ATF4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.145838 (rank : 37) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P18848, Q9UH31 | Gene names | ATF4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (Activating transcription factor 4) (DNA-binding protein TAXREB67) (Cyclic AMP response element-binding protein 2) (CREB2). | |||||
|
COBA2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.047756 (rank : 61) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
EMID1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.063499 (rank : 51) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96A84, Q6ICG1, Q86SS7 | Gene names | EMID1, EMU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMI domain-containing protein 1 precursor (Protein Emu1) (Emilin and multimerin domain-containing protein 1). | |||||
|
NGN3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.068781 (rank : 50) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y4Z2, Q9BY24 | Gene names | NEUROG3, NGN3 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 3. | |||||
|
TEF_HUMAN
|
||||||
| θ value | 0.47712 (rank : 33) | NC score | 0.232537 (rank : 20) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q10587, Q15729, Q7Z3J7, Q8IU94, Q96TG4 | Gene names | TEF | |||
|
Domain Architecture |

|
|||||
| Description | Thyrotroph embryonic factor. | |||||
|
TEF_MOUSE
|
||||||
| θ value | 0.47712 (rank : 34) | NC score | 0.232505 (rank : 21) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9JLC6, Q3U426, Q3UGF4, Q3UM49, Q3URC8, Q6QHT6, Q8C6I0, Q8VD02 | Gene names | Tef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyrotroph embryonic factor. | |||||
|
XBP1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.218190 (rank : 22) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35426, Q8VHM0, Q922G5, Q9ESS3 | Gene names | Xbp1, Treb5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X box-binding protein 1 (XBP-1) (Tax-responsive element-binding protein 5 homolog). | |||||
|
ATF3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 36) | NC score | 0.241386 (rank : 15) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q60765 | Gene names | Atf3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-3 (Activating transcription factor 3) (Transcription factor LRG-21). | |||||
|
ATF4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 37) | NC score | 0.141389 (rank : 39) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q06507, Q61906 | Gene names | Atf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (C/EBP-related ATF) (C/ATF) (TAXREB67 homolog). | |||||
|
JUN_HUMAN
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.163061 (rank : 33) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P05412, Q96G93 | Gene names | JUN | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (p39). | |||||
|
JUN_MOUSE
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.163414 (rank : 32) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P05627 | Gene names | Jun | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (Jun A) (AH119). | |||||
|
ATF3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.234873 (rank : 19) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P18847 | Gene names | ATF3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-3 (Activating transcription factor 3). | |||||
|
CO5A3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.048600 (rank : 59) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P25940, Q9NZQ6 | Gene names | COL5A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(V) chain precursor. | |||||
|
COBA1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.045404 (rank : 63) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P12107, Q14034, Q9UIT4, Q9UIT5, Q9UIT6 | Gene names | COL11A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XI) chain precursor. | |||||
|
LDB3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.015194 (rank : 99) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O75112, Q5K6N9, Q5K6P0, Q5K6P1, Q96FH2, Q9Y4Z3, Q9Y4Z4, Q9Y4Z5 | Gene names | LDB3, KIAA0613, ZASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher). | |||||
|
SMC3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 44) | NC score | 0.033043 (rank : 78) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1186 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UQE7, O60464 | Gene names | CSPG6, BAM, BMH, SMC3, SMC3L1 | |||
|
Domain Architecture |
|
|||||
| Description | Structural maintenance of chromosome 3 (Chondroitin sulfate proteoglycan 6) (Chromosome-associated polypeptide) (hCAP) (Bamacan) (Basement membrane-associated chondroitin proteoglycan). | |||||
|
SMC3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 45) | NC score | 0.033180 (rank : 76) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1187 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9CW03, O35667, Q9QUS3 | Gene names | Cspg6, Bam, Bmh, Mmip1, Smc3, Smc3l1, Smcd | |||
|
Domain Architecture |
|
|||||
| Description | Structural maintenance of chromosome 3 (Chondroitin sulfate proteoglycan 6) (Chromosome segregation protein SmcD) (Bamacan) (Basement membrane-associated chondroitin proteoglycan) (Mad member- interacting protein 1). | |||||
|
CAR14_HUMAN
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.037965 (rank : 72) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BXL6, Q9BVB5 | Gene names | CARD14, CARMA2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 14 (CARD-containing MAGUK protein 2) (Carma 2). | |||||
|
CO3A1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.050721 (rank : 58) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P08121, Q61429, Q9CRN7 | Gene names | Col3a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
COIA1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.040892 (rank : 68) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P39060, Q9UK38, Q9Y6Q7, Q9Y6Q8 | Gene names | COL18A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
CPSF7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.060458 (rank : 54) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N684, Q7Z3H9, Q9H025, Q9H9V1 | Gene names | CPSF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 7 (Cleavage and polyadenylation specificity factor 59 kDa subunit) (CPSF 59 kDa subunit) (Pre-mRNA cleavage factor Im 59 kDa subunit). | |||||
|
CPSF7_MOUSE
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.060497 (rank : 53) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BTV2, Q3TNF1, Q8BKE7, Q8CFS8 | Gene names | Cpsf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 7. | |||||
|
FOSL2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.189570 (rank : 27) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P15408 | Gene names | FOSL2, FRA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
FOSL2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.189786 (rank : 26) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P47930 | Gene names | Fosl2, Fra-2, Fra2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
NGN2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.062119 (rank : 52) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P70447, P70237 | Gene names | Neurog2, Ath4a, Atoh4, Ngn2 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 2 (Atonal protein homolog 4) (Helix-loop-helix protein mATH-4A) (MATH4A). | |||||
|
CAR14_MOUSE
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.035886 (rank : 74) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99KF0 | Gene names | Card14, Bimp2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 14 (Bcl10-interacting MAGUK protein 2) (Bimp2). | |||||
|
EAF2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.044516 (rank : 64) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96CJ1, Q9NZ82 | Gene names | EAF2, TRAITS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELL-associated factor 2 (Testosterone-regulated apoptosis inducer and tumor suppressor protein). | |||||
|
EMID2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.059451 (rank : 55) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96A83 | Gene names | EMID2, COL26A1, EMU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.031714 (rank : 79) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
HOOK3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.021968 (rank : 89) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q86VS8, Q8NBH0, Q9BY13 | Gene names | HOOK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3 (hHK3). | |||||
|
SSH1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.013648 (rank : 101) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q76I79, Q3TDG3, Q69ZM4, Q811E5 | Gene names | Ssh1, Kiaa1298, Ssh1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 1 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-1L) (mSSH-1L). | |||||
|
C102B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 60) | NC score | 0.033177 (rank : 77) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q68D86, Q7Z467, Q8NDK7, Q9H5C1 | Gene names | CCDC102B, C18orf14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 102B. | |||||
|
CNOT3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.028960 (rank : 81) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75175, Q9NZN7, Q9UF76 | Gene names | CNOT3, KIAA0691, NOT3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
COJA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.045637 (rank : 62) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14993, Q00559, Q05850, Q12885, Q13676, Q5JUF0, Q5T424, Q9H572, Q9NPZ2, Q9NQP2 | Gene names | COL19A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XIX) chain precursor (Collagen alpha-1(Y) chain). | |||||
|
JUND_HUMAN
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.137924 (rank : 40) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P17535 | Gene names | JUND | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
NGN2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.048222 (rank : 60) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H2A3, Q8N416 | Gene names | NEUROG2, NGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurogenin 2. | |||||
|
DBP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 65) | NC score | 0.193121 (rank : 24) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q10586 | Gene names | DBP | |||
|
Domain Architecture |

|
|||||
| Description | D site-binding protein (Albumin D box-binding protein) (Albumin D- element-binding protein) (TAXREB302). | |||||
|
DBP_MOUSE
|
||||||
| θ value | 3.0926 (rank : 66) | NC score | 0.192565 (rank : 25) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q60925, Q8VCX3 | Gene names | Dbp | |||
|
Domain Architecture |

|
|||||
| Description | D site-binding protein (Albumin D box-binding protein) (Albumin D- element-binding protein). | |||||
|
EAF2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.052802 (rank : 57) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91ZD6, Q7TN80, Q99KD2 | Gene names | Eaf2, Festa, Traits | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELL-associated factor 2 (Testosterone-regulated apoptosis inducer and tumor suppressor protein) (Ehrlich S-II transcriptional activator factor). | |||||
|
JUND_MOUSE
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.134370 (rank : 41) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P15066 | Gene names | Jund, Jun-d, Jund1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
ROCK1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.006487 (rank : 113) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
AP2A1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.010557 (rank : 107) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95782, Q96CI7, Q96PP6, Q96PP7, Q9H070 | Gene names | AP2A1, ADTAA | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-1 (Adapter-related protein complex 2 alpha- 1 subunit) (Alpha-adaptin A) (Adaptor protein complex AP-2 alpha-1 subunit) (Clathrin assembly protein complex 2 alpha-A large chain) (100 kDa coated vesicle protein A) (Plasma membrane adaptor HA2/AP2 adaptin alpha A subunit). | |||||
|
CO7A1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.033968 (rank : 75) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.029841 (rank : 80) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
AZI1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.021386 (rank : 92) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
CENPU_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.021612 (rank : 91) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q71F23, Q32Q71, Q9H5G1 | Gene names | MLF1IP, CENPU, ICEN24, KLIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein U (CENP-U) (CENP-U(50)) (CENP-50) (Interphase centromere complex protein 24) (MLF1-interacting protein) (KSHV latent nuclear antigen-interacting protein 1). | |||||
|
CO1A2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.044021 (rank : 65) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P08123, P02464, Q9UEB6, Q9UPH0 | Gene names | COL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
CO3A1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.043346 (rank : 67) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
CO4A1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.039227 (rank : 70) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
FBX31_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.024108 (rank : 86) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5XUX0, Q96D73, Q9UFV4 | Gene names | FBXO31, FBX14, FBX31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 31. | |||||
|
GLI1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.001787 (rank : 114) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P08151, Q8TDN9 | Gene names | GLI1, GLI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI1 (Glioma-associated oncogene) (Oncogene GLI). | |||||
|
IFT74_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.020221 (rank : 93) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96LB3, Q6PGQ8, Q9H643, Q9H8G7 | Gene names | IFT74, CCDC2, CMG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
ITSN1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.008970 (rank : 109) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
NGN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.043807 (rank : 66) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92886, Q96HE1 | Gene names | NEUROG1, NEUROD3, NGN, NGN1 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein). | |||||
|
PEPL_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.014924 (rank : 100) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |
|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.015450 (rank : 98) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
RBM25_HUMAN
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.016436 (rank : 95) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
ALX3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.006522 (rank : 112) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95076, O95075 | Gene names | ALX3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein aristaless-like 3 (Proline-rich transcription factor ALX3). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.028423 (rank : 82) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
CO5A2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.038726 (rank : 71) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
CT2NL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.015861 (rank : 97) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
DRD4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.009616 (rank : 108) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P21917 | Gene names | DRD4 | |||
|
Domain Architecture |
|
|||||
| Description | D(4) dopamine receptor (Dopamine D4 receptor) (D(2C) dopamine receptor). | |||||
|
FOSB_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.165677 (rank : 31) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P53539 | Gene names | FOSB, G0S3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein fosB (G0/G1 switch regulatory protein 3). | |||||
|
FOSB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.154803 (rank : 36) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P13346 | Gene names | Fosb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein fosB. | |||||
|
FOSL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.154954 (rank : 35) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P15407 | Gene names | FOSL1, FRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Fos-related antigen 1 (FRA-1). | |||||
|
IF39_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.027958 (rank : 84) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55884, Q9UMF9 | Gene names | EIF3S9 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 9 (eIF-3 eta) (eIF3 p116) (eIF3 p110) (eIF3b) (Prt1 homolog) (hPrt1). | |||||
|
PEPL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.015943 (rank : 96) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
USF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.021853 (rank : 90) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
AKAP9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.012606 (rank : 102) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
CNOT3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.028282 (rank : 83) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8K0V4 | Gene names | Cnot3, Not3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
CO4A4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.039643 (rank : 69) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
CO9A2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.037321 (rank : 73) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q14055 | Gene names | COL9A2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
CT2NL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.018846 (rank : 94) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
DLX5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.001550 (rank : 115) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P56178, Q9UPL1 | Gene names | DLX5 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-5. | |||||
|
GOG8A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.023900 (rank : 87) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 635 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZW0, O94937, Q9NZG8, Q9NZW3 | Gene names | GOLGA8A, KIAA0855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 8A/B (Golgi autoantigen golgin-67) (88 kDa Golgi protein) (Gm88 autoantigen). | |||||
|
HOME2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.010909 (rank : 106) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 532 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NSB8, O95269, O95349, Q9NSB6, Q9NSB7, Q9UNT7 | Gene names | HOMER2 | |||
|
Domain Architecture |
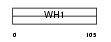
|
|||||
| Description | Homer protein homolog 2 (Homer-2) (Cupidin). | |||||
|
NEB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.011962 (rank : 103) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ULJ8, O76059, Q9NXT2 | Gene names | PPP1R9A, KIAA1222 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-1 (Neurabin-I) (Neural tissue-specific F-actin-binding protein I) (Protein phosphatase 1 regulatory subunit 9A). | |||||
|
OTU7B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.008717 (rank : 110) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6GQQ9, Q8WWA7, Q9NQ53, Q9UFF4 | Gene names | OTUD7B, ZA20D1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | OTU domain-containing protein 7B (EC 3.-.-.-) (Zinc finger protein Cezanne) (Zinc finger A20 domain-containing protein 1) (Cellular zinc finger anti-NF-kappa B protein). | |||||
|
RB6I2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.011494 (rank : 105) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
SL9A5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.011830 (rank : 104) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14940, Q9Y626 | Gene names | SLC9A5, NHE5 | |||
|
Domain Architecture |
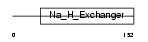
|
|||||
| Description | Sodium/hydrogen exchanger 5 (Na(+)/H(+) exchanger 5) (NHE-5) (Solute carrier family 9 member 5). | |||||
|
ATF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.084236 (rank : 47) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P15336, Q13000 | Gene names | ATF2, CREB2, CREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (HB16). | |||||
|
ATF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.083700 (rank : 48) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P16951, Q64089, Q64090, Q64091 | Gene names | Atf2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (MXBP protein). | |||||
|
ATF7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.102151 (rank : 46) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
ATF7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.103620 (rank : 45) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8R0S1 | Gene names | Atf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
FOSL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.159831 (rank : 34) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P48755, O35285 | Gene names | Fosl1, Fra1 | |||
|
Domain Architecture |

|
|||||
| Description | Fos-related antigen 1 (FRA-1). | |||||
|
FOS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.110401 (rank : 43) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P01100, P18849 | Gene names | FOS, G0S7 | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene protein c-fos (Cellular oncogene fos) (G0/G1 switch regulatory protein 7). | |||||
|
FOS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.110125 (rank : 44) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P01101 | Gene names | Fos | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene protein c-fos (Cellular oncogene fos). | |||||
|
CEBPE_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.42253e-146 (rank : 2) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 108 | |
| SwissProt Accessions | Q6PZD9 | Gene names | Cebpe | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CCAAT/enhancer-binding protein epsilon (C/EBP epsilon). | |||||
|
CEBPE_HUMAN
|
||||||
| NC score | 0.945641 (rank : 2) | θ value | 1.67649e-155 (rank : 1) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q15744, Q15745, Q8IYI2, Q99803 | Gene names | CEBPE | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein epsilon (C/EBP epsilon). | |||||
|
CEBPA_MOUSE
|
||||||
| NC score | 0.894859 (rank : 3) | θ value | 2.86122e-38 (rank : 4) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P53566, Q91XB6 | Gene names | Cebpa | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein alpha (C/EBP alpha). | |||||
|
CEBPA_HUMAN
|
||||||
| NC score | 0.890027 (rank : 4) | θ value | 1.28434e-38 (rank : 3) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P49715, P78319 | Gene names | CEBPA | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein alpha (C/EBP alpha). | |||||
|
CEBPB_MOUSE
|
||||||
| NC score | 0.850804 (rank : 5) | θ value | 1.99067e-23 (rank : 6) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P28033 | Gene names | Cebpb | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein beta (C/EBP beta) (Interleukin-6- dependent-binding protein) (IL-6DBP) (Liver-enriched transcriptional activator) (LAP) (AGP/EBP). | |||||
|
CEBPB_HUMAN
|
||||||
| NC score | 0.834398 (rank : 6) | θ value | 2.51237e-26 (rank : 5) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P17676, Q96IH2, Q9H4Z5 | Gene names | CEBPB, TCF5 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein beta (C/EBP beta) (Nuclear factor NF- IL6) (Transcription factor 5). | |||||
|
CEBPD_HUMAN
|
||||||
| NC score | 0.827214 (rank : 7) | θ value | 1.99067e-23 (rank : 7) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49716, Q14937 | Gene names | CEBPD | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein delta (C/EBP delta) (Nuclear factor NF- IL6-beta) (NF-IL6-beta). | |||||
|
CEBPD_MOUSE
|
||||||
| NC score | 0.821607 (rank : 8) | θ value | 1.29031e-22 (rank : 8) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q00322 | Gene names | Cebpd, Crp3 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein delta (C/EBP delta) (C/EBP-related protein 3). | |||||
|
CEBPG_MOUSE
|
||||||
| NC score | 0.734002 (rank : 9) | θ value | 1.02475e-11 (rank : 10) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P53568 | Gene names | Cebpg | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein gamma (C/EBP gamma) (Immunoglobulin enhancer-binding protein 1) (IG/EBP-1) (Granulocyte colony-stimulating factor promoter element 1-binding protein) (GPE1-binding protein) (GPE1-BP). | |||||
|
CEBPG_HUMAN
|
||||||
| NC score | 0.726445 (rank : 10) | θ value | 1.02475e-11 (rank : 9) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P53567, Q5U052 | Gene names | CEBPG | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein gamma (C/EBP gamma). | |||||
|
DDIT3_MOUSE
|
||||||
| NC score | 0.422276 (rank : 11) | θ value | 0.00298849 (rank : 12) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P35639, Q91YW9 | Gene names | Ddit3, Chop, Gadd153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
DDIT3_HUMAN
|
||||||
| NC score | 0.374760 (rank : 12) | θ value | 0.00869519 (rank : 13) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |

|
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
BATF_HUMAN
|
||||||
| NC score | 0.302528 (rank : 13) | θ value | 0.21417 (rank : 22) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q16520 | Gene names | BATF | |||
|
Domain Architecture |

|
|||||
| Description | ATF-like basic leucine zipper transcriptional factor B-ATF (SF-HT- activated gene 2) (SFA-2). | |||||
|
BATF_MOUSE
|
||||||
| NC score | 0.288558 (rank : 14) | θ value | 0.21417 (rank : 23) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O35284 | Gene names | Batf | |||
|
Domain Architecture |

|
|||||
| Description | ATF-like basic leucine zipper transcription factor B-ATF. | |||||
|
ATF3_MOUSE
|
||||||
| NC score | 0.241386 (rank : 15) | θ value | 0.813845 (rank : 36) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q60765 | Gene names | Atf3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-3 (Activating transcription factor 3) (Transcription factor LRG-21). | |||||
|
HLF_HUMAN
|
||||||
| NC score | 0.240015 (rank : 16) | θ value | 0.0736092 (rank : 17) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16534 | Gene names | HLF | |||
|
Domain Architecture |

|
|||||
| Description | Hepatic leukemia factor. | |||||
|
HLF_MOUSE
|
||||||
| NC score | 0.239599 (rank : 17) | θ value | 0.0736092 (rank : 18) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BW74, Q6PF83 | Gene names | Hlf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatic leukemia factor. | |||||
|
CREB3_MOUSE
|
||||||
| NC score | 0.239362 (rank : 18) | θ value | 0.000461057 (rank : 11) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61817, Q99M21, Q9CVK9 | Gene names | Creb3, Lzip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Transcription factor LZIP). | |||||
|
ATF3_HUMAN
|
||||||
| NC score | 0.234873 (rank : 19) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P18847 | Gene names | ATF3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-3 (Activating transcription factor 3). | |||||
|
TEF_HUMAN
|
||||||
| NC score | 0.232537 (rank : 20) | θ value | 0.47712 (rank : 33) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q10587, Q15729, Q7Z3J7, Q8IU94, Q96TG4 | Gene names | TEF | |||
|
Domain Architecture |

|
|||||
| Description | Thyrotroph embryonic factor. | |||||
|
TEF_MOUSE
|
||||||
| NC score | 0.232505 (rank : 21) | θ value | 0.47712 (rank : 34) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9JLC6, Q3U426, Q3UGF4, Q3UM49, Q3URC8, Q6QHT6, Q8C6I0, Q8VD02 | Gene names | Tef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyrotroph embryonic factor. | |||||
|
XBP1_MOUSE
|
||||||
| NC score | 0.218190 (rank : 22) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35426, Q8VHM0, Q922G5, Q9ESS3 | Gene names | Xbp1, Treb5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X box-binding protein 1 (XBP-1) (Tax-responsive element-binding protein 5 homolog). | |||||
|
XBP1_HUMAN
|
||||||
| NC score | 0.215718 (rank : 23) | θ value | 0.365318 (rank : 28) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P17861, Q8WYK6, Q969P1, Q96BD7 | Gene names | XBP1, TREB5, XBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X box-binding protein 1 (XBP-1) (Tax-responsive element-binding protein 5). | |||||
|
DBP_HUMAN
|
||||||
| NC score | 0.193121 (rank : 24) | θ value | 3.0926 (rank : 65) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q10586 | Gene names | DBP | |||
|
Domain Architecture |

|
|||||
| Description | D site-binding protein (Albumin D box-binding protein) (Albumin D- element-binding protein) (TAXREB302). | |||||
|
DBP_MOUSE
|
||||||
| NC score | 0.192565 (rank : 25) | θ value | 3.0926 (rank : 66) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q60925, Q8VCX3 | Gene names | Dbp | |||
|
Domain Architecture |

|
|||||
| Description | D site-binding protein (Albumin D box-binding protein) (Albumin D- element-binding protein). | |||||
|
FOSL2_MOUSE
|
||||||
| NC score | 0.189786 (rank : 26) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P47930 | Gene names | Fosl2, Fra-2, Fra2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
FOSL2_HUMAN
|
||||||
| NC score | 0.189570 (rank : 27) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P15408 | Gene names | FOSL2, FRA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
CREB3_HUMAN
|
||||||
| NC score | 0.183619 (rank : 28) | θ value | 0.0330416 (rank : 15) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O43889, O14671, O14919, Q96GK8, Q9H2W3, Q9UE77 | Gene names | CREB3, LZIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Luman protein) (Transcription factor LZIP-alpha). | |||||
|
JUNB_MOUSE
|
||||||
| NC score | 0.172180 (rank : 29) | θ value | 0.0113563 (rank : 14) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P09450, Q8C2G9 | Gene names | Junb, Jun-b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-B. | |||||
|
JUNB_HUMAN
|
||||||
| NC score | 0.166705 (rank : 30) | θ value | 0.125558 (rank : 20) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P17275, Q96GH3 | Gene names | JUNB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-B. | |||||
|
FOSB_HUMAN
|
||||||
| NC score | 0.165677 (rank : 31) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P53539 | Gene names | FOSB, G0S3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein fosB (G0/G1 switch regulatory protein 3). | |||||
|
JUN_MOUSE
|
||||||
| NC score | 0.163414 (rank : 32) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P05627 | Gene names | Jun | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (Jun A) (AH119). | |||||
|
JUN_HUMAN
|
||||||
| NC score | 0.163061 (rank : 33) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P05412, Q96G93 | Gene names | JUN | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (p39). | |||||
|
FOSL1_MOUSE
|
||||||
| NC score | 0.159831 (rank : 34) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P48755, O35285 | Gene names | Fosl1, Fra1 | |||
|
Domain Architecture |

|
|||||
| Description | Fos-related antigen 1 (FRA-1). | |||||
|
FOSL1_HUMAN
|
||||||
| NC score | 0.154954 (rank : 35) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P15407 | Gene names | FOSL1, FRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Fos-related antigen 1 (FRA-1). | |||||
|
FOSB_MOUSE
|
||||||
| NC score | 0.154803 (rank : 36) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P13346 | Gene names | Fosb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein fosB. | |||||
|
ATF4_HUMAN
|
||||||
| NC score | 0.145838 (rank : 37) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P18848, Q9UH31 | Gene names | ATF4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (Activating transcription factor 4) (DNA-binding protein TAXREB67) (Cyclic AMP response element-binding protein 2) (CREB2). | |||||
|
CREB5_HUMAN
|
||||||
| NC score | 0.144276 (rank : 38) | θ value | 0.125558 (rank : 19) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q02930, Q05886, Q06246 | Gene names | CREB5, CREBPA | |||
|
Domain Architecture |

|
|||||
| Description | cAMP response element-binding protein 5 (CRE-BPa). | |||||
|
ATF4_MOUSE
|
||||||
| NC score | 0.141389 (rank : 39) | θ value | 0.813845 (rank : 37) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q06507, Q61906 | Gene names | Atf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (C/EBP-related ATF) (C/ATF) (TAXREB67 homolog). | |||||
|
JUND_HUMAN
|
||||||
| NC score | 0.137924 (rank : 40) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P17535 | Gene names | JUND | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
JUND_MOUSE
|
||||||
| NC score | 0.134370 (rank : 41) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P15066 | Gene names | Jund, Jun-d, Jund1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
ATF6A_HUMAN
|
||||||
| NC score | 0.116445 (rank : 42) | θ value | 0.0736092 (rank : 16) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P18850, O15139, Q5VW62, Q6IPB5, Q9UEC9 | Gene names | ATF6 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-6 alpha (Activating transcription factor 6 alpha) (ATF6-alpha). | |||||
|
FOS_HUMAN
|
||||||
| NC score | 0.110401 (rank : 43) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P01100, P18849 | Gene names | FOS, G0S7 | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene protein c-fos (Cellular oncogene fos) (G0/G1 switch regulatory protein 7). | |||||
|
FOS_MOUSE
|
||||||
| NC score | 0.110125 (rank : 44) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P01101 | Gene names | Fos | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene protein c-fos (Cellular oncogene fos). | |||||
|
ATF7_MOUSE
|
||||||
| NC score | 0.103620 (rank : 45) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8R0S1 | Gene names | Atf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
ATF7_HUMAN
|
||||||
| NC score | 0.102151 (rank : 46) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
ATF2_HUMAN
|
||||||
| NC score | 0.084236 (rank : 47) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P15336, Q13000 | Gene names | ATF2, CREB2, CREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (HB16). | |||||
|
ATF2_MOUSE
|
||||||
| NC score | 0.083700 (rank : 48) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P16951, Q64089, Q64090, Q64091 | Gene names | Atf2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (MXBP protein). | |||||
|
NGN3_MOUSE
|
||||||
| NC score | 0.071604 (rank : 49) | θ value | 0.365318 (rank : 27) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P70661 | Gene names | Neurog3, Ath4b, Atoh5, Ngn3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 3 (Atonal protein homolog 5) (Helix-loop-helix protein mATH-4B) (MATH4B). | |||||
|
NGN3_HUMAN
|
||||||
| NC score | 0.068781 (rank : 50) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y4Z2, Q9BY24 | Gene names | NEUROG3, NGN3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 3. | |||||
|
EMID1_HUMAN
|
||||||
| NC score | 0.063499 (rank : 51) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96A84, Q6ICG1, Q86SS7 | Gene names | EMID1, EMU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMI domain-containing protein 1 precursor (Protein Emu1) (Emilin and multimerin domain-containing protein 1). | |||||
|
NGN2_MOUSE
|
||||||
| NC score | 0.062119 (rank : 52) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P70447, P70237 | Gene names | Neurog2, Ath4a, Atoh4, Ngn2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 2 (Atonal protein homolog 4) (Helix-loop-helix protein mATH-4A) (MATH4A). | |||||
|
CPSF7_MOUSE
|
||||||
| NC score | 0.060497 (rank : 53) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BTV2, Q3TNF1, Q8BKE7, Q8CFS8 | Gene names | Cpsf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 7. | |||||
|
CPSF7_HUMAN
|
||||||
| NC score | 0.060458 (rank : 54) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N684, Q7Z3H9, Q9H025, Q9H9V1 | Gene names | CPSF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 7 (Cleavage and polyadenylation specificity factor 59 kDa subunit) (CPSF 59 kDa subunit) (Pre-mRNA cleavage factor Im 59 kDa subunit). | |||||
|
EMID2_HUMAN
|
||||||
| NC score | 0.059451 (rank : 55) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96A83 | Gene names | EMID2, COL26A1, EMU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
CO1A2_MOUSE
|
||||||
| NC score | 0.053209 (rank : 56) | θ value | 0.21417 (rank : 24) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q01149, Q8CGA5 | Gene names | Col1a2, Cola2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
EAF2_MOUSE
|
||||||
| NC score | 0.052802 (rank : 57) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91ZD6, Q7TN80, Q99KD2 | Gene names | Eaf2, Festa, Traits | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELL-associated factor 2 (Testosterone-regulated apoptosis inducer and tumor suppressor protein) (Ehrlich S-II transcriptional activator factor). | |||||
|
CO3A1_MOUSE
|
||||||
| NC score | 0.050721 (rank : 58) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P08121, Q61429, Q9CRN7 | Gene names | Col3a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
CO5A3_HUMAN
|
||||||
| NC score | 0.048600 (rank : 59) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P25940, Q9NZQ6 | Gene names | COL5A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(V) chain precursor. | |||||
|
NGN2_HUMAN
|
||||||
| NC score | 0.048222 (rank : 60) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H2A3, Q8N416 | Gene names | NEUROG2, NGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurogenin 2. | |||||
|
COBA2_HUMAN
|
||||||
| NC score | 0.047756 (rank : 61) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
COJA1_HUMAN
|
||||||
| NC score | 0.045637 (rank : 62) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14993, Q00559, Q05850, Q12885, Q13676, Q5JUF0, Q5T424, Q9H572, Q9NPZ2, Q9NQP2 | Gene names | COL19A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XIX) chain precursor (Collagen alpha-1(Y) chain). | |||||
|
COBA1_HUMAN
|
||||||
| NC score | 0.045404 (rank : 63) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P12107, Q14034, Q9UIT4, Q9UIT5, Q9UIT6 | Gene names | COL11A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XI) chain precursor. | |||||
|
EAF2_HUMAN
|
||||||
| NC score | 0.044516 (rank : 64) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96CJ1, Q9NZ82 | Gene names | EAF2, TRAITS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELL-associated factor 2 (Testosterone-regulated apoptosis inducer and tumor suppressor protein). | |||||
|
CO1A2_HUMAN
|
||||||
| NC score | 0.044021 (rank : 65) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P08123, P02464, Q9UEB6, Q9UPH0 | Gene names | COL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
NGN1_HUMAN
|
||||||
| NC score | 0.043807 (rank : 66) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92886, Q96HE1 | Gene names | NEUROG1, NEUROD3, NGN, NGN1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein). | |||||
|
CO3A1_HUMAN
|
||||||
| NC score | 0.043346 (rank : 67) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
COIA1_HUMAN
|
||||||
| NC score | 0.040892 (rank : 68) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P39060, Q9UK38, Q9Y6Q7, Q9Y6Q8 | Gene names | COL18A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
CO4A4_HUMAN
|
||||||
| NC score | 0.039643 (rank : 69) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
CO4A1_MOUSE
|
||||||
| NC score | 0.039227 (rank : 70) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CO5A2_HUMAN
|
||||||
| NC score | 0.038726 (rank : 71) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
CAR14_HUMAN
|
||||||
| NC score | 0.037965 (rank : 72) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BXL6, Q9BVB5 | Gene names | CARD14, CARMA2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 14 (CARD-containing MAGUK protein 2) (Carma 2). | |||||
|
CO9A2_HUMAN
|
||||||
| NC score | 0.037321 (rank : 73) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q14055 | Gene names | COL9A2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
CAR14_MOUSE
|
||||||
| NC score | 0.035886 (rank : 74) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99KF0 | Gene names | Card14, Bimp2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 14 (Bcl10-interacting MAGUK protein 2) (Bimp2). | |||||
|
CO7A1_HUMAN
|
||||||
| NC score | 0.033968 (rank : 75) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
SMC3_MOUSE
|
||||||
| NC score | 0.033180 (rank : 76) | θ value | 1.06291 (rank : 45) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1187 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9CW03, O35667, Q9QUS3 | Gene names | Cspg6, Bam, Bmh, Mmip1, Smc3, Smc3l1, Smcd | |||
|
Domain Architecture |
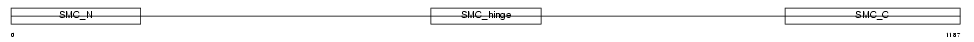
|
|||||
| Description | Structural maintenance of chromosome 3 (Chondroitin sulfate proteoglycan 6) (Chromosome segregation protein SmcD) (Bamacan) (Basement membrane-associated chondroitin proteoglycan) (Mad member- interacting protein 1). | |||||
|
C102B_HUMAN
|
||||||
| NC score | 0.033177 (rank : 77) | θ value | 2.36792 (rank : 60) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q68D86, Q7Z467, Q8NDK7, Q9H5C1 | Gene names | CCDC102B, C18orf14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 102B. | |||||
|
SMC3_HUMAN
|
||||||
| NC score | 0.033043 (rank : 78) | θ value | 1.06291 (rank : 44) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1186 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UQE7, O60464 | Gene names | CSPG6, BAM, BMH, SMC3, SMC3L1 | |||
|
Domain Architecture |
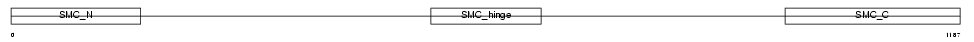
|
|||||
| Description | Structural maintenance of chromosome 3 (Chondroitin sulfate proteoglycan 6) (Chromosome-associated polypeptide) (hCAP) (Bamacan) (Basement membrane-associated chondroitin proteoglycan). | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.031714 (rank : 79) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.029841 (rank : 80) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
CNOT3_HUMAN
|
||||||
| NC score | 0.028960 (rank : 81) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75175, Q9NZN7, Q9UF76 | Gene names | CNOT3, KIAA0691, NOT3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.028423 (rank : 82) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
CNOT3_MOUSE
|
||||||
| NC score | 0.028282 (rank : 83) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8K0V4 | Gene names | Cnot3, Not3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
IF39_HUMAN
|
||||||
| NC score | 0.027958 (rank : 84) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55884, Q9UMF9 | Gene names | EIF3S9 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 9 (eIF-3 eta) (eIF3 p116) (eIF3 p110) (eIF3b) (Prt1 homolog) (hPrt1). | |||||
|
KCNH2_HUMAN
|
||||||
| NC score | 0.025920 (rank : 85) | θ value | 0.21417 (rank : 25) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
FBX31_HUMAN
|
||||||
| NC score | 0.024108 (rank : 86) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5XUX0, Q96D73, Q9UFV4 | Gene names | FBXO31, FBX14, FBX31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 31. | |||||
|
GOG8A_HUMAN
|
||||||
| NC score | 0.023900 (rank : 87) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 635 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZW0, O94937, Q9NZG8, Q9NZW3 | Gene names | GOLGA8A, KIAA0855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 8A/B (Golgi autoantigen golgin-67) (88 kDa Golgi protein) (Gm88 autoantigen). | |||||
|
ACHA4_MOUSE
|
||||||
| NC score | 0.022249 (rank : 88) | θ value | 0.21417 (rank : 21) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70174, Q8BHE9, Q8VI10, Q9ET51 | Gene names | Chrna4, Acra4 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal acetylcholine receptor protein subunit alpha-4 precursor. | |||||
|
HOOK3_HUMAN
|
||||||
| NC score | 0.021968 (rank : 89) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q86VS8, Q8NBH0, Q9BY13 | Gene names | HOOK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3 (hHK3). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.021853 (rank : 90) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
CENPU_HUMAN
|
||||||
| NC score | 0.021612 (rank : 91) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q71F23, Q32Q71, Q9H5G1 | Gene names | MLF1IP, CENPU, ICEN24, KLIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein U (CENP-U) (CENP-U(50)) (CENP-50) (Interphase centromere complex protein 24) (MLF1-interacting protein) (KSHV latent nuclear antigen-interacting protein 1). | |||||
|
AZI1_MOUSE
|
||||||
| NC score | 0.021386 (rank : 92) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
IFT74_HUMAN
|
||||||
| NC score | 0.020221 (rank : 93) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96LB3, Q6PGQ8, Q9H643, Q9H8G7 | Gene names | IFT74, CCDC2, CMG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
CT2NL_HUMAN
|
||||||
| NC score | 0.018846 (rank : 94) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
RBM25_HUMAN
|
||||||
| NC score | 0.016436 (rank : 95) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
PEPL_MOUSE
|
||||||
| NC score | 0.015943 (rank : 96) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
CT2NL_MOUSE
|
||||||
| NC score | 0.015861 (rank : 97) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.015450 (rank : 98) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
LDB3_HUMAN
|
||||||
| NC score | 0.015194 (rank : 99) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O75112, Q5K6N9, Q5K6P0, Q5K6P1, Q96FH2, Q9Y4Z3, Q9Y4Z4, Q9Y4Z5 | Gene names | LDB3, KIAA0613, ZASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher). | |||||
|
PEPL_HUMAN
|
||||||
| NC score | 0.014924 (rank : 100) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
SSH1_MOUSE
|
||||||
| NC score | 0.013648 (rank : 101) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q76I79, Q3TDG3, Q69ZM4, Q811E5 | Gene names | Ssh1, Kiaa1298, Ssh1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 1 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-1L) (mSSH-1L). | |||||
|
AKAP9_HUMAN
|
||||||
| NC score | 0.012606 (rank : 102) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
NEB1_HUMAN
|
||||||
| NC score | 0.011962 (rank : 103) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ULJ8, O76059, Q9NXT2 | Gene names | PPP1R9A, KIAA1222 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-1 (Neurabin-I) (Neural tissue-specific F-actin-binding protein I) (Protein phosphatase 1 regulatory subunit 9A). | |||||
|
SL9A5_HUMAN
|
||||||
| NC score | 0.011830 (rank : 104) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14940, Q9Y626 | Gene names | SLC9A5, NHE5 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium/hydrogen exchanger 5 (Na(+)/H(+) exchanger 5) (NHE-5) (Solute carrier family 9 member 5). | |||||
|
RB6I2_HUMAN
|
||||||
| NC score | 0.011494 (rank : 105) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
HOME2_HUMAN
|
||||||
| NC score | 0.010909 (rank : 106) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 532 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NSB8, O95269, O95349, Q9NSB6, Q9NSB7, Q9UNT7 | Gene names | HOMER2 | |||
|
Domain Architecture |

|
|||||
| Description | Homer protein homolog 2 (Homer-2) (Cupidin). | |||||
|
AP2A1_HUMAN
|
||||||
| NC score | 0.010557 (rank : 107) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95782, Q96CI7, Q96PP6, Q96PP7, Q9H070 | Gene names | AP2A1, ADTAA | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-1 (Adapter-related protein complex 2 alpha- 1 subunit) (Alpha-adaptin A) (Adaptor protein complex AP-2 alpha-1 subunit) (Clathrin assembly protein complex 2 alpha-A large chain) (100 kDa coated vesicle protein A) (Plasma membrane adaptor HA2/AP2 adaptin alpha A subunit). | |||||
|
DRD4_HUMAN
|
||||||
| NC score | 0.009616 (rank : 108) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P21917 | Gene names | DRD4 | |||
|
Domain Architecture |

|
|||||
| Description | D(4) dopamine receptor (Dopamine D4 receptor) (D(2C) dopamine receptor). | |||||
|
ITSN1_MOUSE
|
||||||
| NC score | 0.008970 (rank : 109) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
OTU7B_HUMAN
|
||||||
| NC score | 0.008717 (rank : 110) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6GQQ9, Q8WWA7, Q9NQ53, Q9UFF4 | Gene names | OTUD7B, ZA20D1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | OTU domain-containing protein 7B (EC 3.-.-.-) (Zinc finger protein Cezanne) (Zinc finger A20 domain-containing protein 1) (Cellular zinc finger anti-NF-kappa B protein). | |||||
|
KLF2_HUMAN
|
||||||
| NC score | 0.007898 (rank : 111) | θ value | 0.365318 (rank : 26) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5W3, Q9UJS5, Q9UKR6 | Gene names | KLF2, LKLF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 2 (Lung krueppel-like factor). | |||||
|
ALX3_HUMAN
|
||||||
| NC score | 0.006522 (rank : 112) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95076, O95075 | Gene names | ALX3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein aristaless-like 3 (Proline-rich transcription factor ALX3). | |||||
|
ROCK1_MOUSE
|
||||||
| NC score | 0.006487 (rank : 113) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
GLI1_HUMAN
|
||||||
| NC score | 0.001787 (rank : 114) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P08151, Q8TDN9 | Gene names | GLI1, GLI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI1 (Glioma-associated oncogene) (Oncogene GLI). | |||||
|
DLX5_HUMAN
|
||||||
| NC score | 0.001550 (rank : 115) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P56178, Q9UPL1 | Gene names | DLX5 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-5. | |||||