Please be patient as the page loads
|
WDR63_HUMAN
|
||||||
| SwissProt Accessions | Q8IWG1, Q96L72, Q96NU4 | Gene names | WDR63 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 63 (Testis development protein NYD-SP29). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
WDR63_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q8IWG1, Q96L72, Q96NU4 | Gene names | WDR63 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 63 (Testis development protein NYD-SP29). | |||||
|
DNAI2_HUMAN
|
||||||
| θ value | 1.99067e-23 (rank : 2) | NC score | 0.576566 (rank : 2) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9GZS0, Q9H179, Q9NT53 | Gene names | DNAI2 | |||
|
Domain Architecture |
|
|||||
| Description | Dynein intermediate chain 2, axonemal (Axonemal dynein intermediate chain 2). | |||||
|
DC1I1_MOUSE
|
||||||
| θ value | 6.21693e-09 (rank : 3) | NC score | 0.438720 (rank : 3) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O88485, O88486 | Gene names | Dync1i1, Dnci1, Dncic1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 1 (Dynein intermediate chain 1, cytosolic) (DH IC-1) (Cytoplasmic dynein intermediate chain 1). | |||||
|
DC1I1_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 4) | NC score | 0.432327 (rank : 4) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O14576, Q9Y2X1 | Gene names | DYNC1I1, DNCI1, DNCIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 1 (Dynein intermediate chain 1, cytosolic) (DH IC-1) (Cytoplasmic dynein intermediate chain 1). | |||||
|
DC1I2_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 5) | NC score | 0.423590 (rank : 5) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13409, Q96NG7, Q96S87, Q9BXZ5, Q9NT58 | Gene names | DYNC1I2, DNCI2, DNCIC2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 2 (Dynein intermediate chain 2, cytosolic) (DH IC-2) (Cytoplasmic dynein intermediate chain 2). | |||||
|
DC1I2_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 6) | NC score | 0.421358 (rank : 6) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O88487, Q3TGH7 | Gene names | Dync1i2, Dnci2, Dncic2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 2 (Dynein intermediate chain 2, cytosolic) (DH IC-2) (Cytoplasmic dynein intermediate chain 2). | |||||
|
WDR34_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 7) | NC score | 0.386341 (rank : 7) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5U4F6 | Gene names | Wdr34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 34. | |||||
|
WDR34_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 8) | NC score | 0.383518 (rank : 8) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96EX3, Q5VXV4, Q9BV46 | Gene names | WDR34 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 34. | |||||
|
WDR60_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 9) | NC score | 0.244611 (rank : 10) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8C761, Q3UNM4 | Gene names | Wdr60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
WDR60_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 10) | NC score | 0.229641 (rank : 11) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WVS4, Q9NW58 | Gene names | WDR60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
ROCK2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 11) | NC score | 0.030410 (rank : 46) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
ROCK2_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 12) | NC score | 0.030318 (rank : 47) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
DNAI1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 13) | NC score | 0.328766 (rank : 9) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UI46, Q9UEZ8 | Gene names | DNAI1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynein intermediate chain 1, axonemal (Axonemal dynein intermediate chain 1). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 0.365318 (rank : 14) | NC score | 0.042134 (rank : 34) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
HELC1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 15) | NC score | 0.050650 (rank : 30) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N3C0, O43738, Q9H1I9, Q9NTR0 | Gene names | ASCC3, HELIC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 3 (EC 3.6.1.-) (ASC-1 complex subunit p200) (Trip4 complex subunit p200) (Helicase, ATP binding 1). | |||||
|
KI21A_HUMAN
|
||||||
| θ value | 0.47712 (rank : 16) | NC score | 0.029822 (rank : 50) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1543 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7Z4S6, Q6UKL9, Q7Z668, Q86WZ5, Q8IVZ8, Q9C0F5, Q9NXU4, Q9Y590 | Gene names | KIF21A, KIAA1708, KIF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21A (Kinesin-like protein KIF2) (NY-REN-62 antigen). | |||||
|
BRD7_MOUSE
|
||||||
| θ value | 0.62314 (rank : 17) | NC score | 0.056608 (rank : 23) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88665, Q3UQ56, Q3UU06, Q9CT78 | Gene names | Brd7, Bp75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 7 (75 kDa bromodomain protein). | |||||
|
CDC20_MOUSE
|
||||||
| θ value | 0.62314 (rank : 18) | NC score | 0.048119 (rank : 31) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JJ66, Q3TGP1, Q8BPG4, Q99LK3 | Gene names | Cdc20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC) (mmCdc20). | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 19) | NC score | 0.039801 (rank : 35) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
TTK_HUMAN
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.017426 (rank : 59) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P33981 | Gene names | TTK, MPS1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein kinase TTK (EC 2.7.12.1) (Phosphotyrosine picked threonine-protein kinase) (PYT). | |||||
|
WDR17_HUMAN
|
||||||
| θ value | 0.62314 (rank : 21) | NC score | 0.083438 (rank : 12) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZU2 | Gene names | WDR17 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 17. | |||||
|
BRD7_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.055037 (rank : 24) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NPI1, Q4VC09, Q8N2L9, Q96KA4, Q9BV48, Q9UH59 | Gene names | BRD7, BP75, CELTIX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 7 (75 kDa bromodomain protein) (Protein CELTIX-1). | |||||
|
GRWD1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.078834 (rank : 13) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q810D6, Q922H3 | Gene names | Grwd1, A301 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich WD repeat-containing protein 1 (Protein A301). | |||||
|
C1QR1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.014577 (rank : 62) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89103, Q3U2X0 | Gene names | Cd93, Aa4, C1qr1, C1qrp, Ly68 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (Cell surface antigen AA4) (Lymphocyte antigen 68). | |||||
|
AKAP9_HUMAN
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.035428 (rank : 39) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
MYH4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.031186 (rank : 42) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
PWP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.054058 (rank : 27) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13610 | Gene names | PWP1 | |||
|
Domain Architecture |
|
|||||
| Description | Periodic tryptophan protein 1 homolog (Keratinocyte protein IEF SSP 9502). | |||||
|
MYH8_MOUSE
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.030019 (rank : 48) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
PRP19_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.045526 (rank : 32) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UMS4 | Gene names | PRPF19, NMP200, PRP19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor 19 (PRP19/PSO4 homolog) (Nuclear matrix protein 200) (hPso4). | |||||
|
PRP19_MOUSE
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.044494 (rank : 33) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99KP6, Q3TP64, Q8BKZ5, Q8BVQ4 | Gene names | Prpf19, Prp19, Snev | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor 19 (PRP19/PSO4 homolog) (Nuclear matrix protein 200) (Nuclear matrix protein SNEV). | |||||
|
JHD1A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.038434 (rank : 36) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
JHD1A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.037470 (rank : 38) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
PRPF3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.054322 (rank : 25) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43395, O43446 | Gene names | PRPF3, HPRP3, PRP3 | |||
|
Domain Architecture |
|
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp3 (Pre-mRNA-splicing factor 3) (U4/U6 snRNP 90 kDa protein) (hPrp3). | |||||
|
PRPF3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.054062 (rank : 26) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q922U1, Q3TJH4, Q9D6C6 | Gene names | Prpf3 | |||
|
Domain Architecture |
|
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp3 (Pre-mRNA-splicing factor 3). | |||||
|
PWP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.051889 (rank : 28) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99LL5, Q9D6T6 | Gene names | Pwp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Periodic tryptophan protein 1 homolog. | |||||
|
VAV2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.018491 (rank : 58) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
JHD1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.032522 (rank : 40) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.029141 (rank : 51) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
UNC5B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.012663 (rank : 64) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K1S3, Q3U4F2, Q6PFH0, Q80Y85, Q9D398 | Gene names | Unc5b, Unc5h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5B precursor (Unc-5 homolog B) (Unc-5 homolog 2). | |||||
|
FA48A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.031059 (rank : 43) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEM7, Q71RF3, Q9Y6A6 | Gene names | FAM48A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM48A (p38-interacting protein) (p38IP). | |||||
|
MCM7_MOUSE
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.015761 (rank : 60) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61881 | Gene names | Mcm7, Cdc47, Mcmd7 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM7 (CDC47 homolog). | |||||
|
ROCK1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.025243 (rank : 57) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2000 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13464 | Gene names | ROCK1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK) (NY-REN-35 antigen). | |||||
|
ROCK1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.025532 (rank : 56) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
MYH3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.029992 (rank : 49) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
ZN473_MOUSE
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | -0.000332 (rank : 67) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 824 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BI67, Q8BI98, Q8BIB7 | Gene names | Znf473, Zfp100, Zfp473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 473 homolog (Zinc finger protein 100) (Zfp-100). | |||||
|
CDC20_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.037643 (rank : 37) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q12834, Q9BW56, Q9UQI9 | Gene names | CDC20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC). | |||||
|
DNJC7_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.009333 (rank : 65) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QYI3, Q3TKR1, Q8BPG3, Q8CIL2, Q9CT29, Q9D026 | Gene names | Dnajc7, Ttc2 | |||
|
Domain Architecture |
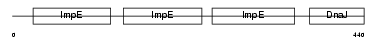
|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2) (MDj11). | |||||
|
PEX7_MOUSE
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.067881 (rank : 19) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P97865 | Gene names | Pex7 | |||
|
Domain Architecture |
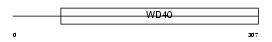
|
|||||
| Description | Peroxisomal targeting signal 2 receptor (PTS2 receptor) (Peroxin-7). | |||||
|
TRI12_MOUSE
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.007360 (rank : 66) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99PQ1, Q9D704 | Gene names | Trim12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 12. | |||||
|
VAV2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.015338 (rank : 61) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
AHI1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.030746 (rank : 44) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K3E5, Q7TNV2, Q9CVY1 | Gene names | Ahi1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein) (AHI-1). | |||||
|
CCD96_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.031851 (rank : 41) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
CT2NL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.028986 (rank : 52) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
GEMI5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.066897 (rank : 20) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 424 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BX17 | Gene names | Gemin5 | |||
|
Domain Architecture |

|
|||||
| Description | Gem-associated protein 5 (Gemin5). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.025616 (rank : 55) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
LUZP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.026524 (rank : 54) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
MYH10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.026834 (rank : 53) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
RFIP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.013640 (rank : 63) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7L804, Q9Y2F0 | Gene names | RAB11FIP2, KIAA0941 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 2 (Rab11-FIP2) (NRip11). | |||||
|
TAXB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.030624 (rank : 45) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q86VP1, O60398, O95770, Q13311, Q9BQG5, Q9UI88 | Gene names | TAX1BP1, T6BP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tax1-binding protein 1 (TRAF6-binding protein). | |||||
|
GEMI5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.058440 (rank : 22) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8TEQ6, Q8WWV4, Q969W4, Q9H9K3, Q9UFI5 | Gene names | GEMIN5 | |||
|
Domain Architecture |

|
|||||
| Description | Gem-associated protein 5 (Gemin5). | |||||
|
GRWD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.060526 (rank : 21) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BQ67 | Gene names | GRWD1, GRWD, WDR28 | |||
|
Domain Architecture |
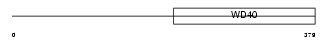
|
|||||
| Description | Glutamate-rich WD repeat-containing protein 1. | |||||
|
PEX7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.068014 (rank : 18) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O00628 | Gene names | PEX7, PTS2R | |||
|
Domain Architecture |
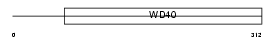
|
|||||
| Description | Peroxisomal targeting signal 2 receptor (PTS2 receptor) (Peroxin-7). | |||||
|
RBBP4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.071019 (rank : 14) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q09028, P31149, Q53H02, Q96BV9 | Gene names | RBBP4, RBAP48 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-binding protein RBBP4 (Retinoblastoma-binding protein 4) (RBBP-4) (Retinoblastoma-binding protein p48) (Chromatin assembly factor 1 subunit C) (CAF-1 subunit C) (Chromatin assembly factor I p48 subunit) (CAF-I 48 kDa subunit) (CAF-I p48) (Nucleosome remodeling factor subunit RBAP48). | |||||
|
RBBP4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.070139 (rank : 15) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60972 | Gene names | Rbbp4, Rbap48 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-binding protein RBBP4 (Retinoblastoma-binding protein 4) (RBBP-4) (Retinoblastoma-binding protein p48) (Chromatin assembly factor 1 subunit C) (CAF-1 subunit C) (Chromatin assembly factor I p48 subunit) (CAF-I 48 kDa subunit) (CAF-I p48) (Nucleosome remodeling factor subunit RBAP48). | |||||
|
RBBP7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.069946 (rank : 16) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q16576 | Gene names | RBBP7, RBAP46 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-binding protein RBBP7 (Retinoblastoma-binding protein 7) (RBBP-7) (Retinoblastoma-binding protein p46) (Histone acetyltransferase type B subunit 2) (Nucleosome remodeling factor subunit RBAP46). | |||||
|
RBBP7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.069913 (rank : 17) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60973, Q3UX20 | Gene names | Rbbp7, Rbap46 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-binding protein RBBP7 (Retinoblastoma-binding protein 7) (RBBP-7) (Retinoblastoma-binding protein p46) (Histone acetyltransferase type B subunit 2) (Nucleosome remodeling factor subunit RBAP46). | |||||
|
THOC3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.051686 (rank : 29) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8VE80, Q9CWI8 | Gene names | Thoc3 | |||
|
Domain Architecture |
|
|||||
| Description | THO complex subunit 3 (Tho3). | |||||
|
WDR63_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q8IWG1, Q96L72, Q96NU4 | Gene names | WDR63 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 63 (Testis development protein NYD-SP29). | |||||
|
DNAI2_HUMAN
|
||||||
| NC score | 0.576566 (rank : 2) | θ value | 1.99067e-23 (rank : 2) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9GZS0, Q9H179, Q9NT53 | Gene names | DNAI2 | |||
|
Domain Architecture |

|
|||||
| Description | Dynein intermediate chain 2, axonemal (Axonemal dynein intermediate chain 2). | |||||
|
DC1I1_MOUSE
|
||||||
| NC score | 0.438720 (rank : 3) | θ value | 6.21693e-09 (rank : 3) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O88485, O88486 | Gene names | Dync1i1, Dnci1, Dncic1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 1 (Dynein intermediate chain 1, cytosolic) (DH IC-1) (Cytoplasmic dynein intermediate chain 1). | |||||
|
DC1I1_HUMAN
|
||||||
| NC score | 0.432327 (rank : 4) | θ value | 4.0297e-08 (rank : 4) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O14576, Q9Y2X1 | Gene names | DYNC1I1, DNCI1, DNCIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 1 (Dynein intermediate chain 1, cytosolic) (DH IC-1) (Cytoplasmic dynein intermediate chain 1). | |||||
|
DC1I2_HUMAN
|
||||||
| NC score | 0.423590 (rank : 5) | θ value | 1.99992e-07 (rank : 5) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13409, Q96NG7, Q96S87, Q9BXZ5, Q9NT58 | Gene names | DYNC1I2, DNCI2, DNCIC2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 2 (Dynein intermediate chain 2, cytosolic) (DH IC-2) (Cytoplasmic dynein intermediate chain 2). | |||||
|
DC1I2_MOUSE
|
||||||
| NC score | 0.421358 (rank : 6) | θ value | 5.81887e-07 (rank : 6) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O88487, Q3TGH7 | Gene names | Dync1i2, Dnci2, Dncic2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 2 (Dynein intermediate chain 2, cytosolic) (DH IC-2) (Cytoplasmic dynein intermediate chain 2). | |||||
|
WDR34_MOUSE
|
||||||
| NC score | 0.386341 (rank : 7) | θ value | 0.000158464 (rank : 7) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5U4F6 | Gene names | Wdr34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 34. | |||||
|
WDR34_HUMAN
|
||||||
| NC score | 0.383518 (rank : 8) | θ value | 0.00035302 (rank : 8) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96EX3, Q5VXV4, Q9BV46 | Gene names | WDR34 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 34. | |||||
|
DNAI1_HUMAN
|
||||||
| NC score | 0.328766 (rank : 9) | θ value | 0.0961366 (rank : 13) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UI46, Q9UEZ8 | Gene names | DNAI1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynein intermediate chain 1, axonemal (Axonemal dynein intermediate chain 1). | |||||
|
WDR60_MOUSE
|
||||||
| NC score | 0.244611 (rank : 10) | θ value | 0.00298849 (rank : 9) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8C761, Q3UNM4 | Gene names | Wdr60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
WDR60_HUMAN
|
||||||
| NC score | 0.229641 (rank : 11) | θ value | 0.0431538 (rank : 10) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WVS4, Q9NW58 | Gene names | WDR60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
WDR17_HUMAN
|
||||||
| NC score | 0.083438 (rank : 12) | θ value | 0.62314 (rank : 21) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZU2 | Gene names | WDR17 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 17. | |||||
|
GRWD1_MOUSE
|
||||||
| NC score | 0.078834 (rank : 13) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q810D6, Q922H3 | Gene names | Grwd1, A301 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich WD repeat-containing protein 1 (Protein A301). | |||||
|
RBBP4_HUMAN
|
||||||
| NC score | 0.071019 (rank : 14) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q09028, P31149, Q53H02, Q96BV9 | Gene names | RBBP4, RBAP48 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-binding protein RBBP4 (Retinoblastoma-binding protein 4) (RBBP-4) (Retinoblastoma-binding protein p48) (Chromatin assembly factor 1 subunit C) (CAF-1 subunit C) (Chromatin assembly factor I p48 subunit) (CAF-I 48 kDa subunit) (CAF-I p48) (Nucleosome remodeling factor subunit RBAP48). | |||||
|
RBBP4_MOUSE
|
||||||
| NC score | 0.070139 (rank : 15) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60972 | Gene names | Rbbp4, Rbap48 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-binding protein RBBP4 (Retinoblastoma-binding protein 4) (RBBP-4) (Retinoblastoma-binding protein p48) (Chromatin assembly factor 1 subunit C) (CAF-1 subunit C) (Chromatin assembly factor I p48 subunit) (CAF-I 48 kDa subunit) (CAF-I p48) (Nucleosome remodeling factor subunit RBAP48). | |||||
|
RBBP7_HUMAN
|
||||||
| NC score | 0.069946 (rank : 16) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q16576 | Gene names | RBBP7, RBAP46 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-binding protein RBBP7 (Retinoblastoma-binding protein 7) (RBBP-7) (Retinoblastoma-binding protein p46) (Histone acetyltransferase type B subunit 2) (Nucleosome remodeling factor subunit RBAP46). | |||||
|
RBBP7_MOUSE
|
||||||
| NC score | 0.069913 (rank : 17) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60973, Q3UX20 | Gene names | Rbbp7, Rbap46 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-binding protein RBBP7 (Retinoblastoma-binding protein 7) (RBBP-7) (Retinoblastoma-binding protein p46) (Histone acetyltransferase type B subunit 2) (Nucleosome remodeling factor subunit RBAP46). | |||||
|
PEX7_HUMAN
|
||||||
| NC score | 0.068014 (rank : 18) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O00628 | Gene names | PEX7, PTS2R | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal targeting signal 2 receptor (PTS2 receptor) (Peroxin-7). | |||||
|
PEX7_MOUSE
|
||||||
| NC score | 0.067881 (rank : 19) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P97865 | Gene names | Pex7 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal targeting signal 2 receptor (PTS2 receptor) (Peroxin-7). | |||||
|
GEMI5_MOUSE
|
||||||
| NC score | 0.066897 (rank : 20) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 424 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BX17 | Gene names | Gemin5 | |||
|
Domain Architecture |

|
|||||
| Description | Gem-associated protein 5 (Gemin5). | |||||
|
GRWD1_HUMAN
|
||||||
| NC score | 0.060526 (rank : 21) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BQ67 | Gene names | GRWD1, GRWD, WDR28 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate-rich WD repeat-containing protein 1. | |||||
|
GEMI5_HUMAN
|
||||||
| NC score | 0.058440 (rank : 22) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8TEQ6, Q8WWV4, Q969W4, Q9H9K3, Q9UFI5 | Gene names | GEMIN5 | |||
|
Domain Architecture |

|
|||||
| Description | Gem-associated protein 5 (Gemin5). | |||||
|
BRD7_MOUSE
|
||||||
| NC score | 0.056608 (rank : 23) | θ value | 0.62314 (rank : 17) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88665, Q3UQ56, Q3UU06, Q9CT78 | Gene names | Brd7, Bp75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 7 (75 kDa bromodomain protein). | |||||
|
BRD7_HUMAN
|
||||||
| NC score | 0.055037 (rank : 24) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NPI1, Q4VC09, Q8N2L9, Q96KA4, Q9BV48, Q9UH59 | Gene names | BRD7, BP75, CELTIX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 7 (75 kDa bromodomain protein) (Protein CELTIX-1). | |||||
|
PRPF3_HUMAN
|
||||||
| NC score | 0.054322 (rank : 25) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43395, O43446 | Gene names | PRPF3, HPRP3, PRP3 | |||
|
Domain Architecture |

|
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp3 (Pre-mRNA-splicing factor 3) (U4/U6 snRNP 90 kDa protein) (hPrp3). | |||||
|
PRPF3_MOUSE
|
||||||
| NC score | 0.054062 (rank : 26) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q922U1, Q3TJH4, Q9D6C6 | Gene names | Prpf3 | |||
|
Domain Architecture |

|
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp3 (Pre-mRNA-splicing factor 3). | |||||
|
PWP1_HUMAN
|
||||||
| NC score | 0.054058 (rank : 27) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13610 | Gene names | PWP1 | |||
|
Domain Architecture |

|
|||||
| Description | Periodic tryptophan protein 1 homolog (Keratinocyte protein IEF SSP 9502). | |||||
|
PWP1_MOUSE
|
||||||
| NC score | 0.051889 (rank : 28) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99LL5, Q9D6T6 | Gene names | Pwp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Periodic tryptophan protein 1 homolog. | |||||
|
THOC3_MOUSE
|
||||||
| NC score | 0.051686 (rank : 29) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8VE80, Q9CWI8 | Gene names | Thoc3 | |||
|
Domain Architecture |

|
|||||
| Description | THO complex subunit 3 (Tho3). | |||||
|
HELC1_HUMAN
|
||||||
| NC score | 0.050650 (rank : 30) | θ value | 0.47712 (rank : 15) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N3C0, O43738, Q9H1I9, Q9NTR0 | Gene names | ASCC3, HELIC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 3 (EC 3.6.1.-) (ASC-1 complex subunit p200) (Trip4 complex subunit p200) (Helicase, ATP binding 1). | |||||
|
CDC20_MOUSE
|
||||||
| NC score | 0.048119 (rank : 31) | θ value | 0.62314 (rank : 18) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JJ66, Q3TGP1, Q8BPG4, Q99LK3 | Gene names | Cdc20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC) (mmCdc20). | |||||
|
PRP19_HUMAN
|
||||||
| NC score | 0.045526 (rank : 32) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UMS4 | Gene names | PRPF19, NMP200, PRP19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor 19 (PRP19/PSO4 homolog) (Nuclear matrix protein 200) (hPso4). | |||||
|
PRP19_MOUSE
|
||||||
| NC score | 0.044494 (rank : 33) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99KP6, Q3TP64, Q8BKZ5, Q8BVQ4 | Gene names | Prpf19, Prp19, Snev | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor 19 (PRP19/PSO4 homolog) (Nuclear matrix protein 200) (Nuclear matrix protein SNEV). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.042134 (rank : 34) | θ value | 0.365318 (rank : 14) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.039801 (rank : 35) | θ value | 0.62314 (rank : 19) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
JHD1A_HUMAN
|
||||||
| NC score | 0.038434 (rank : 36) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
CDC20_HUMAN
|
||||||
| NC score | 0.037643 (rank : 37) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q12834, Q9BW56, Q9UQI9 | Gene names | CDC20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC). | |||||
|
JHD1A_MOUSE
|
||||||
| NC score | 0.037470 (rank : 38) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
AKAP9_HUMAN
|
||||||
| NC score | 0.035428 (rank : 39) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
JHD1B_MOUSE
|
||||||
| NC score | 0.032522 (rank : 40) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
CCD96_MOUSE
|
||||||
| NC score | 0.031851 (rank : 41) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
MYH4_HUMAN
|
||||||
| NC score | 0.031186 (rank : 42) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
FA48A_HUMAN
|
||||||
| NC score | 0.031059 (rank : 43) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEM7, Q71RF3, Q9Y6A6 | Gene names | FAM48A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM48A (p38-interacting protein) (p38IP). | |||||
|
AHI1_MOUSE
|
||||||
| NC score | 0.030746 (rank : 44) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K3E5, Q7TNV2, Q9CVY1 | Gene names | Ahi1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein) (AHI-1). | |||||
|
TAXB1_HUMAN
|
||||||
| NC score | 0.030624 (rank : 45) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q86VP1, O60398, O95770, Q13311, Q9BQG5, Q9UI88 | Gene names | TAX1BP1, T6BP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tax1-binding protein 1 (TRAF6-binding protein). | |||||
|
ROCK2_HUMAN
|
||||||
| NC score | 0.030410 (rank : 46) | θ value | 0.0563607 (rank : 11) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
ROCK2_MOUSE
|
||||||
| NC score | 0.030318 (rank : 47) | θ value | 0.0563607 (rank : 12) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
MYH8_MOUSE
|
||||||
| NC score | 0.030019 (rank : 48) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
MYH3_HUMAN
|
||||||
| NC score | 0.029992 (rank : 49) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
KI21A_HUMAN
|
||||||
| NC score | 0.029822 (rank : 50) | θ value | 0.47712 (rank : 16) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1543 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7Z4S6, Q6UKL9, Q7Z668, Q86WZ5, Q8IVZ8, Q9C0F5, Q9NXU4, Q9Y590 | Gene names | KIF21A, KIAA1708, KIF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21A (Kinesin-like protein KIF2) (NY-REN-62 antigen). | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.029141 (rank : 51) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
CT2NL_HUMAN
|
||||||
| NC score | 0.028986 (rank : 52) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
MYH10_HUMAN
|
||||||
| NC score | 0.026834 (rank : 53) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
LUZP1_MOUSE
|
||||||
| NC score | 0.026524 (rank : 54) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.025616 (rank : 55) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
ROCK1_MOUSE
|
||||||
| NC score | 0.025532 (rank : 56) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
ROCK1_HUMAN
|
||||||
| NC score | 0.025243 (rank : 57) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2000 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13464 | Gene names | ROCK1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK) (NY-REN-35 antigen). | |||||
|
VAV2_MOUSE
|
||||||
| NC score | 0.018491 (rank : 58) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
TTK_HUMAN
|
||||||
| NC score | 0.017426 (rank : 59) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P33981 | Gene names | TTK, MPS1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein kinase TTK (EC 2.7.12.1) (Phosphotyrosine picked threonine-protein kinase) (PYT). | |||||
|
MCM7_MOUSE
|
||||||
| NC score | 0.015761 (rank : 60) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61881 | Gene names | Mcm7, Cdc47, Mcmd7 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM7 (CDC47 homolog). | |||||
|
VAV2_HUMAN
|
||||||
| NC score | 0.015338 (rank : 61) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
C1QR1_MOUSE
|
||||||
| NC score | 0.014577 (rank : 62) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89103, Q3U2X0 | Gene names | Cd93, Aa4, C1qr1, C1qrp, Ly68 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (Cell surface antigen AA4) (Lymphocyte antigen 68). | |||||
|
RFIP2_HUMAN
|
||||||
| NC score | 0.013640 (rank : 63) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7L804, Q9Y2F0 | Gene names | RAB11FIP2, KIAA0941 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 2 (Rab11-FIP2) (NRip11). | |||||
|
UNC5B_MOUSE
|
||||||
| NC score | 0.012663 (rank : 64) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K1S3, Q3U4F2, Q6PFH0, Q80Y85, Q9D398 | Gene names | Unc5b, Unc5h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5B precursor (Unc-5 homolog B) (Unc-5 homolog 2). | |||||
|
DNJC7_MOUSE
|
||||||
| NC score | 0.009333 (rank : 65) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QYI3, Q3TKR1, Q8BPG3, Q8CIL2, Q9CT29, Q9D026 | Gene names | Dnajc7, Ttc2 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 7 (Tetratricopeptide repeat protein 2) (TPR repeat protein 2) (MDj11). | |||||
|
TRI12_MOUSE
|
||||||
| NC score | 0.007360 (rank : 66) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99PQ1, Q9D704 | Gene names | Trim12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 12. | |||||
|
ZN473_MOUSE
|
||||||
| NC score | -0.000332 (rank : 67) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 824 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BI67, Q8BI98, Q8BIB7 | Gene names | Znf473, Zfp100, Zfp473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 473 homolog (Zinc finger protein 100) (Zfp-100). | |||||