Please be patient as the page loads
|
SYIM_HUMAN
|
||||||
| SwissProt Accessions | Q9NSE4, Q6PI85, Q7L439, Q86WU9, Q96D91 | Gene names | IARS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Isoleucyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.5) (Isoleucine--tRNA ligase) (IleRS). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SYIM_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NSE4, Q6PI85, Q7L439, Q86WU9, Q96D91 | Gene names | IARS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Isoleucyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.5) (Isoleucine--tRNA ligase) (IleRS). | |||||
|
SYIM_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.996656 (rank : 2) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BIJ6, Q3TKT8, Q8BIJ0, Q8BIP3 | Gene names | Iars2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Isoleucyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.5) (Isoleucine--tRNA ligase) (IleRS). | |||||
|
SYIC_HUMAN
|
||||||
| θ value | 5.8972e-76 (rank : 3) | NC score | 0.847861 (rank : 4) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P41252, Q5TCD0, Q9H588 | Gene names | IARS | |||
|
Domain Architecture |

|
|||||
| Description | Isoleucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.5) (Isoleucine--tRNA ligase) (IleRS) (IRS). | |||||
|
SYIC_MOUSE
|
||||||
| θ value | 1.22964e-73 (rank : 4) | NC score | 0.847879 (rank : 3) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BU30 | Gene names | Iars | |||
|
Domain Architecture |

|
|||||
| Description | Isoleucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.5) (Isoleucine--tRNA ligase) (IleRS) (IRS). | |||||
|
SYV_HUMAN
|
||||||
| θ value | 4.70527e-49 (rank : 5) | NC score | 0.753741 (rank : 6) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26640, Q5JQ90, Q96E77, Q9UQM2 | Gene names | VARS, G7A, VARS2 | |||
|
Domain Architecture |

|
|||||
| Description | Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--tRNA ligase) (ValRS) (Protein G7a). | |||||
|
SYV_MOUSE
|
||||||
| θ value | 6.7944e-48 (rank : 6) | NC score | 0.756982 (rank : 5) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Z1Q9, Q9QUN2 | Gene names | Vars, Bat6, G7a, Vars2 | |||
|
Domain Architecture |
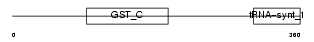
|
|||||
| Description | Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--tRNA ligase) (ValRS) (Protein G7a). | |||||
|
SYLM_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 7) | NC score | 0.477728 (rank : 7) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VDC0 | Gene names | Lars2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable leucyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
SYLM_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 8) | NC score | 0.471072 (rank : 8) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15031 | Gene names | LARS2, KIAA0028 | |||
|
Domain Architecture |

|
|||||
| Description | Probable leucyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
SYLC_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 9) | NC score | 0.266768 (rank : 9) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P2J5, Q9NSE1 | Gene names | LARS, KIAA1352 | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
SYLC_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 10) | NC score | 0.252071 (rank : 10) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BMJ2, Q8BKW9 | Gene names | Lars | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
SYMM_MOUSE
|
||||||
| θ value | 0.163984 (rank : 11) | NC score | 0.173616 (rank : 11) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q499X9, Q3T9W7, Q8BWR9, Q8BXY1 | Gene names | Mars2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methionyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.10) (Methionine--tRNA ligase 2) (Mitochondrial methionine--tRNA ligase) (MtMetRS). | |||||
|
SYMM_HUMAN
|
||||||
| θ value | 0.279714 (rank : 12) | NC score | 0.163492 (rank : 12) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96GW9, Q76E79, Q8IW62, Q8N7N4 | Gene names | MARS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methionyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.10) (Methionine--tRNA ligase 2) (Mitochondrial methionine--tRNA ligase) (MtMetRS). | |||||
|
SYM_MOUSE
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.081659 (rank : 16) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q68FL6 | Gene names | Mars | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methionyl-tRNA synthetase (EC 6.1.1.10) (Methionine--tRNA ligase) (MetRS). | |||||
|
SYM_HUMAN
|
||||||
| θ value | 1.81305 (rank : 14) | NC score | 0.110915 (rank : 15) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P56192, Q14895, Q53H14, Q96A15, Q96BZ0, Q9NSE0 | Gene names | MARS | |||
|
Domain Architecture |

|
|||||
| Description | Methionyl-tRNA synthetase (EC 6.1.1.10) (Methionine--tRNA ligase) (MetRS). | |||||
|
DUS8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 15) | NC score | 0.007648 (rank : 19) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13202 | Gene names | DUSP8, VH5 | |||
|
Domain Architecture |
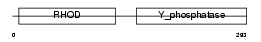
|
|||||
| Description | Dual specificity protein phosphatase 8 (EC 3.1.3.48) (EC 3.1.3.16) (Dual specificity protein phosphatase hVH-5). | |||||
|
U520_HUMAN
|
||||||
| θ value | 5.27518 (rank : 16) | NC score | 0.013757 (rank : 17) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75643, O94884, Q6PX59, Q96IF2, Q9H7S0 | Gene names | ASCC3L1, HELIC2, KIAA0788 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U5 small nuclear ribonucleoprotein 200 kDa helicase (EC 3.6.1.-) (U5 snRNP-specific 200 kDa protein) (U5-200KD) (Activating signal cointegrator 1 complex subunit 3-like 1) (BRR2 homolog). | |||||
|
KI13B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 17) | NC score | 0.007751 (rank : 18) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
LAMC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 18) | NC score | 0.001218 (rank : 22) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
PDE7B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.004410 (rank : 21) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NP56 | Gene names | PDE7B | |||
|
Domain Architecture |
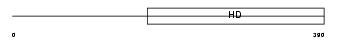
|
|||||
| Description | cAMP-specific 3',5'-cyclic phosphodiesterase 7B (EC 3.1.4.17). | |||||
|
ST1B1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 20) | NC score | 0.004675 (rank : 20) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QWG7, Q8C301, Q9Z2T0 | Gene names | Sult1b1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sulfotransferase family cytosolic 1B member 1 (EC 2.8.2.-) (Sulfotransferase 1B1) (DOPA/tyrosine sulfotransferase). | |||||
|
EF1G_HUMAN
|
||||||
| θ value | θ > 10 (rank : 21) | NC score | 0.141125 (rank : 13) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P26641, Q6PJ62, Q6PK31, Q96CU2, Q9P196 | Gene names | EEF1G, EF1G | |||
|
Domain Architecture |
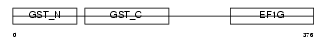
|
|||||
| Description | Elongation factor 1-gamma (EF-1-gamma) (eEF-1B gamma). | |||||
|
EF1G_MOUSE
|
||||||
| θ value | θ > 10 (rank : 22) | NC score | 0.140461 (rank : 14) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D8N0, Q920C5, Q9CRT5, Q9CSU3, Q9D004 | Gene names | Eef1g | |||
|
Domain Architecture |
|
|||||
| Description | Elongation factor 1-gamma (EF-1-gamma) (eEF-1B gamma). | |||||
|
SYIM_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NSE4, Q6PI85, Q7L439, Q86WU9, Q96D91 | Gene names | IARS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Isoleucyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.5) (Isoleucine--tRNA ligase) (IleRS). | |||||
|
SYIM_MOUSE
|
||||||
| NC score | 0.996656 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BIJ6, Q3TKT8, Q8BIJ0, Q8BIP3 | Gene names | Iars2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Isoleucyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.5) (Isoleucine--tRNA ligase) (IleRS). | |||||
|
SYIC_MOUSE
|
||||||
| NC score | 0.847879 (rank : 3) | θ value | 1.22964e-73 (rank : 4) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BU30 | Gene names | Iars | |||
|
Domain Architecture |

|
|||||
| Description | Isoleucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.5) (Isoleucine--tRNA ligase) (IleRS) (IRS). | |||||
|
SYIC_HUMAN
|
||||||
| NC score | 0.847861 (rank : 4) | θ value | 5.8972e-76 (rank : 3) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P41252, Q5TCD0, Q9H588 | Gene names | IARS | |||
|
Domain Architecture |

|
|||||
| Description | Isoleucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.5) (Isoleucine--tRNA ligase) (IleRS) (IRS). | |||||
|
SYV_MOUSE
|
||||||
| NC score | 0.756982 (rank : 5) | θ value | 6.7944e-48 (rank : 6) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Z1Q9, Q9QUN2 | Gene names | Vars, Bat6, G7a, Vars2 | |||
|
Domain Architecture |

|
|||||
| Description | Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--tRNA ligase) (ValRS) (Protein G7a). | |||||
|
SYV_HUMAN
|
||||||
| NC score | 0.753741 (rank : 6) | θ value | 4.70527e-49 (rank : 5) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26640, Q5JQ90, Q96E77, Q9UQM2 | Gene names | VARS, G7A, VARS2 | |||
|
Domain Architecture |

|
|||||
| Description | Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--tRNA ligase) (ValRS) (Protein G7a). | |||||
|
SYLM_MOUSE
|
||||||
| NC score | 0.477728 (rank : 7) | θ value | 3.29651e-10 (rank : 7) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VDC0 | Gene names | Lars2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable leucyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
SYLM_HUMAN
|
||||||
| NC score | 0.471072 (rank : 8) | θ value | 2.79066e-09 (rank : 8) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15031 | Gene names | LARS2, KIAA0028 | |||
|
Domain Architecture |

|
|||||
| Description | Probable leucyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
SYLC_HUMAN
|
||||||
| NC score | 0.266768 (rank : 9) | θ value | 0.00228821 (rank : 9) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P2J5, Q9NSE1 | Gene names | LARS, KIAA1352 | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
SYLC_MOUSE
|
||||||
| NC score | 0.252071 (rank : 10) | θ value | 0.00665767 (rank : 10) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BMJ2, Q8BKW9 | Gene names | Lars | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
SYMM_MOUSE
|
||||||
| NC score | 0.173616 (rank : 11) | θ value | 0.163984 (rank : 11) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q499X9, Q3T9W7, Q8BWR9, Q8BXY1 | Gene names | Mars2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methionyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.10) (Methionine--tRNA ligase 2) (Mitochondrial methionine--tRNA ligase) (MtMetRS). | |||||
|
SYMM_HUMAN
|
||||||
| NC score | 0.163492 (rank : 12) | θ value | 0.279714 (rank : 12) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96GW9, Q76E79, Q8IW62, Q8N7N4 | Gene names | MARS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methionyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.10) (Methionine--tRNA ligase 2) (Mitochondrial methionine--tRNA ligase) (MtMetRS). | |||||
|
EF1G_HUMAN
|
||||||
| NC score | 0.141125 (rank : 13) | θ value | θ > 10 (rank : 21) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P26641, Q6PJ62, Q6PK31, Q96CU2, Q9P196 | Gene names | EEF1G, EF1G | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 1-gamma (EF-1-gamma) (eEF-1B gamma). | |||||
|
EF1G_MOUSE
|
||||||
| NC score | 0.140461 (rank : 14) | θ value | θ > 10 (rank : 22) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D8N0, Q920C5, Q9CRT5, Q9CSU3, Q9D004 | Gene names | Eef1g | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 1-gamma (EF-1-gamma) (eEF-1B gamma). | |||||
|
SYM_HUMAN
|
||||||
| NC score | 0.110915 (rank : 15) | θ value | 1.81305 (rank : 14) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P56192, Q14895, Q53H14, Q96A15, Q96BZ0, Q9NSE0 | Gene names | MARS | |||
|
Domain Architecture |

|
|||||
| Description | Methionyl-tRNA synthetase (EC 6.1.1.10) (Methionine--tRNA ligase) (MetRS). | |||||
|
SYM_MOUSE
|
||||||
| NC score | 0.081659 (rank : 16) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q68FL6 | Gene names | Mars | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methionyl-tRNA synthetase (EC 6.1.1.10) (Methionine--tRNA ligase) (MetRS). | |||||
|
U520_HUMAN
|
||||||
| NC score | 0.013757 (rank : 17) | θ value | 5.27518 (rank : 16) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75643, O94884, Q6PX59, Q96IF2, Q9H7S0 | Gene names | ASCC3L1, HELIC2, KIAA0788 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U5 small nuclear ribonucleoprotein 200 kDa helicase (EC 3.6.1.-) (U5 snRNP-specific 200 kDa protein) (U5-200KD) (Activating signal cointegrator 1 complex subunit 3-like 1) (BRR2 homolog). | |||||
|
KI13B_HUMAN
|
||||||
| NC score | 0.007751 (rank : 18) | θ value | 6.88961 (rank : 17) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
DUS8_HUMAN
|
||||||
| NC score | 0.007648 (rank : 19) | θ value | 5.27518 (rank : 15) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13202 | Gene names | DUSP8, VH5 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein phosphatase 8 (EC 3.1.3.48) (EC 3.1.3.16) (Dual specificity protein phosphatase hVH-5). | |||||
|
ST1B1_MOUSE
|
||||||
| NC score | 0.004675 (rank : 20) | θ value | 8.99809 (rank : 20) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QWG7, Q8C301, Q9Z2T0 | Gene names | Sult1b1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sulfotransferase family cytosolic 1B member 1 (EC 2.8.2.-) (Sulfotransferase 1B1) (DOPA/tyrosine sulfotransferase). | |||||
|
PDE7B_HUMAN
|
||||||
| NC score | 0.004410 (rank : 21) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NP56 | Gene names | PDE7B | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-specific 3',5'-cyclic phosphodiesterase 7B (EC 3.1.4.17). | |||||
|
LAMC1_HUMAN
|
||||||
| NC score | 0.001218 (rank : 22) | θ value | 8.99809 (rank : 18) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||