Please be patient as the page loads
|
NUDT8_MOUSE
|
||||||
| SwissProt Accessions | Q9CR24, Q3TDQ2 | Gene names | Nudt8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoside diphosphate-linked moiety X motif 8, mitochondrial precursor (EC 3.6.1.-) (Nudix motif 8). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NUDT8_MOUSE
|
||||||
| θ value | 1.18001e-100 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CR24, Q3TDQ2 | Gene names | Nudt8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoside diphosphate-linked moiety X motif 8, mitochondrial precursor (EC 3.6.1.-) (Nudix motif 8). | |||||
|
NUDT8_HUMAN
|
||||||
| θ value | 1.95407e-87 (rank : 2) | NC score | 0.993048 (rank : 2) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WV74, Q6ZW59 | Gene names | NUDT8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoside diphosphate-linked moiety X motif 8, mitochondrial precursor (EC 3.6.1.-) (Nudix motif 8). | |||||
|
NUDT7_MOUSE
|
||||||
| θ value | 8.38298e-14 (rank : 3) | NC score | 0.679700 (rank : 3) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99P30, Q6IS65, Q8BU08, Q8K260, Q9D0Q3, Q9DBI9 | Gene names | Nudt7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisomal coenzyme A diphosphatase NUDT7 (EC 3.6.1.-) (Nucleoside diphosphate-linked moiety X motif 7) (Nudix motif 7). | |||||
|
NUDT7_HUMAN
|
||||||
| θ value | 1.42992e-13 (rank : 4) | NC score | 0.678468 (rank : 4) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P0C024 | Gene names | NUDT7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisomal coenzyme A diphosphatase NUDT7 (EC 3.6.1.-) (Nucleoside diphosphate-linked moiety X motif 7) (Nudix motif 7). | |||||
|
AP4A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 5) | NC score | 0.087349 (rank : 6) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50583 | Gene names | NUDT2, APAH1 | |||
|
Domain Architecture |
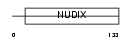
|
|||||
| Description | Bis(5'-nucleosyl)-tetraphosphatase [Asymmetrical] (EC 3.6.1.17) (Diadenosine 5',5'''-P1,P4-tetraphosphate asymmetrical hydrolase) (Diadenosine tetraphosphatase) (Ap4A hydrolase) (Ap4Aase) (Nucleoside diphosphate-linked moiety X motif 2) (Nudix motif 2). | |||||
|
NUD17_MOUSE
|
||||||
| θ value | 1.81305 (rank : 6) | NC score | 0.051645 (rank : 7) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CWD3, Q3URR9 | Gene names | Nudt17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoside diphosphate-linked moiety X motif 17, mitochondrial precursor (EC 3.6.1.-) (Nudix motif 17). | |||||
|
AP4A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 7) | NC score | 0.093956 (rank : 5) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P56380, Q9D2U6, Q9D6V2 | Gene names | Nudt2, Apah1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bis(5'-nucleosyl)-tetraphosphatase [Asymmetrical] (EC 3.6.1.17) (Diadenosine 5',5'''-P1,P4-tetraphosphate asymmetrical hydrolase) (Diadenosine tetraphosphatase) (Ap4A hydrolase) (Ap4Aase) (Nucleoside diphosphate-linked moiety X motif 2) (Nudix motif 2). | |||||
|
NUD11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 8) | NC score | 0.014823 (rank : 8) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96G61, Q9NVN0 | Gene names | NUDT11, APS1, DIPP3B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diphosphoinositol polyphosphate phosphohydrolase 3 beta (EC 3.6.1.52) (DIPP-3 beta) (DIPP3 beta) (hDIPP3beta) (Diadenosine 5',5'''-P1,P6- hexaphosphate hydrolase 3 beta) (EC 3.6.1.-) (Nucleoside diphosphate- linked moiety X motif 11) (Nudix motif 11) (hAps1). | |||||
|
NUDT8_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.18001e-100 (rank : 1) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CR24, Q3TDQ2 | Gene names | Nudt8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoside diphosphate-linked moiety X motif 8, mitochondrial precursor (EC 3.6.1.-) (Nudix motif 8). | |||||
|
NUDT8_HUMAN
|
||||||
| NC score | 0.993048 (rank : 2) | θ value | 1.95407e-87 (rank : 2) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WV74, Q6ZW59 | Gene names | NUDT8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoside diphosphate-linked moiety X motif 8, mitochondrial precursor (EC 3.6.1.-) (Nudix motif 8). | |||||
|
NUDT7_MOUSE
|
||||||
| NC score | 0.679700 (rank : 3) | θ value | 8.38298e-14 (rank : 3) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99P30, Q6IS65, Q8BU08, Q8K260, Q9D0Q3, Q9DBI9 | Gene names | Nudt7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisomal coenzyme A diphosphatase NUDT7 (EC 3.6.1.-) (Nucleoside diphosphate-linked moiety X motif 7) (Nudix motif 7). | |||||
|
NUDT7_HUMAN
|
||||||
| NC score | 0.678468 (rank : 4) | θ value | 1.42992e-13 (rank : 4) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P0C024 | Gene names | NUDT7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisomal coenzyme A diphosphatase NUDT7 (EC 3.6.1.-) (Nucleoside diphosphate-linked moiety X motif 7) (Nudix motif 7). | |||||
|
AP4A_MOUSE
|
||||||
| NC score | 0.093956 (rank : 5) | θ value | 2.36792 (rank : 7) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P56380, Q9D2U6, Q9D6V2 | Gene names | Nudt2, Apah1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bis(5'-nucleosyl)-tetraphosphatase [Asymmetrical] (EC 3.6.1.17) (Diadenosine 5',5'''-P1,P4-tetraphosphate asymmetrical hydrolase) (Diadenosine tetraphosphatase) (Ap4A hydrolase) (Ap4Aase) (Nucleoside diphosphate-linked moiety X motif 2) (Nudix motif 2). | |||||
|
AP4A_HUMAN
|
||||||
| NC score | 0.087349 (rank : 6) | θ value | 1.81305 (rank : 5) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50583 | Gene names | NUDT2, APAH1 | |||
|
Domain Architecture |

|
|||||
| Description | Bis(5'-nucleosyl)-tetraphosphatase [Asymmetrical] (EC 3.6.1.17) (Diadenosine 5',5'''-P1,P4-tetraphosphate asymmetrical hydrolase) (Diadenosine tetraphosphatase) (Ap4A hydrolase) (Ap4Aase) (Nucleoside diphosphate-linked moiety X motif 2) (Nudix motif 2). | |||||
|
NUD17_MOUSE
|
||||||
| NC score | 0.051645 (rank : 7) | θ value | 1.81305 (rank : 6) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CWD3, Q3URR9 | Gene names | Nudt17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoside diphosphate-linked moiety X motif 17, mitochondrial precursor (EC 3.6.1.-) (Nudix motif 17). | |||||
|
NUD11_HUMAN
|
||||||
| NC score | 0.014823 (rank : 8) | θ value | 8.99809 (rank : 8) | |||
| Query Neighborhood Hits | 8 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96G61, Q9NVN0 | Gene names | NUDT11, APS1, DIPP3B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diphosphoinositol polyphosphate phosphohydrolase 3 beta (EC 3.6.1.52) (DIPP-3 beta) (DIPP3 beta) (hDIPP3beta) (Diadenosine 5',5'''-P1,P6- hexaphosphate hydrolase 3 beta) (EC 3.6.1.-) (Nucleoside diphosphate- linked moiety X motif 11) (Nudix motif 11) (hAps1). | |||||