Please be patient as the page loads
|
MUS81_HUMAN
|
||||||
| SwissProt Accessions | Q96NY9, Q9H7D9 | Gene names | MUS81 | |||
|
Domain Architecture |

|
|||||
| Description | Crossover junction endonuclease MUS81 (EC 3.1.22.-). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MUS81_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96NY9, Q9H7D9 | Gene names | MUS81 | |||
|
Domain Architecture |

|
|||||
| Description | Crossover junction endonuclease MUS81 (EC 3.1.22.-). | |||||
|
MUS81_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.993650 (rank : 2) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91ZJ0, Q8R356 | Gene names | Mus81 | |||
|
Domain Architecture |

|
|||||
| Description | Crossover junction endonuclease MUS81 (EC 3.1.22.-). | |||||
|
EME1_MOUSE
|
||||||
| θ value | 1.69304e-06 (rank : 3) | NC score | 0.340597 (rank : 3) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BJW7 | Gene names | Eme1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crossover junction endonuclease EME1 (EC 3.1.22.-). | |||||
|
EME1_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 4) | NC score | 0.325996 (rank : 4) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96AY2, Q96N62 | Gene names | EME1, MMS4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crossover junction endonuclease EME1 (EC 3.1.22.-) (hMMS4). | |||||
|
UN45B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 5) | NC score | 0.038755 (rank : 5) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IWX7, Q495Q8, Q495Q9 | Gene names | UNC45B, CMYA4, UNC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog B (UNC-45B). | |||||
|
UN45B_MOUSE
|
||||||
| θ value | 1.38821 (rank : 6) | NC score | 0.037602 (rank : 6) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CGY6, Q5XG72, Q8BHC5, Q8BWK3 | Gene names | Unc45b, Cmya4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog B (UNC-45B). | |||||
|
MYO5B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 7) | NC score | 0.009591 (rank : 17) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULV0 | Gene names | MYO5B, KIAA1119 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5B (Myosin Vb). | |||||
|
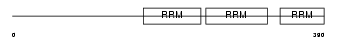
HNRPR_HUMAN
|
||||||
| θ value | 3.0926 (rank : 8) | NC score | 0.014267 (rank : 11) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43390 | Gene names | HNRPR | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein R (hnRNP R). | |||||
|
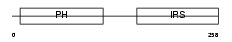
IRS1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 9) | NC score | 0.021827 (rank : 7) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35568 | Gene names | IRS1 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin receptor substrate 1 (IRS-1). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 10) | NC score | 0.012242 (rank : 13) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
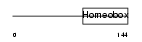
PHX2A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 11) | NC score | 0.010205 (rank : 15) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14813, Q8IVZ2 | Gene names | PHOX2A, ARIX, PMX2A | |||
|
Domain Architecture |
|
|||||
| Description | Paired mesoderm homeobox protein 2A (Paired-like homeobox 2A) (Aristaless homeobox protein homolog) (ARIX1 homeodomain protein). | |||||
|
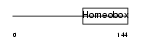
PHX2A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 12) | NC score | 0.010406 (rank : 14) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62066 | Gene names | Phox2a, Arix, Phox2, Pmx2, Pmx2a | |||
|
Domain Architecture |
|
|||||
| Description | Paired mesoderm homeobox protein 2A (Paired-like homeobox 2A) (PHOX2A homeodomain protein) (Aristaless homeobox protein homolog). | |||||
|
MYH14_HUMAN
|
||||||
| θ value | 5.27518 (rank : 13) | NC score | 0.008581 (rank : 18) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
WNK4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 14) | NC score | 0.003781 (rank : 20) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
ACK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 15) | NC score | 0.001975 (rank : 21) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1140 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07912, Q8N6U7, Q96H59 | Gene names | TNK2, ACK1 | |||
|
Domain Architecture |
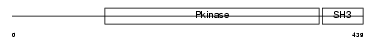
|
|||||
| Description | Activated CDC42 kinase 1 (EC 2.7.10.2) (ACK-1) (Tyrosine kinase non- receptor protein 2). | |||||
|
B4GN4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 16) | NC score | 0.013761 (rank : 12) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q76KP1, Q96LV2 | Gene names | B4GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
TAF6L_HUMAN
|
||||||
| θ value | 6.88961 (rank : 17) | NC score | 0.017284 (rank : 9) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6J9, Q96HA6 | Gene names | TAF6L, PAF65A | |||
|
Domain Architecture |
|
|||||
| Description | TAF6-like RNA polymerase II p300/CBP-associated factor-associated factor 65 kDa subunit 6L (PCAF-associated factor 65 alpha) (PAF65- alpha). | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 18) | NC score | 0.007425 (rank : 19) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.016971 (rank : 10) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 20) | NC score | 0.017617 (rank : 8) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
SCAM3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 21) | NC score | 0.010102 (rank : 16) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35609, Q99M48, Q9ERM9 | Gene names | Scamp3 | |||
|
Domain Architecture |

|
|||||
| Description | Secretory carrier-associated membrane protein 3 (Secretory carrier membrane protein 3). | |||||
|
MUS81_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96NY9, Q9H7D9 | Gene names | MUS81 | |||
|
Domain Architecture |

|
|||||
| Description | Crossover junction endonuclease MUS81 (EC 3.1.22.-). | |||||
|
MUS81_MOUSE
|
||||||
| NC score | 0.993650 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91ZJ0, Q8R356 | Gene names | Mus81 | |||
|
Domain Architecture |

|
|||||
| Description | Crossover junction endonuclease MUS81 (EC 3.1.22.-). | |||||
|
EME1_MOUSE
|
||||||
| NC score | 0.340597 (rank : 3) | θ value | 1.69304e-06 (rank : 3) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BJW7 | Gene names | Eme1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crossover junction endonuclease EME1 (EC 3.1.22.-). | |||||
|
EME1_HUMAN
|
||||||
| NC score | 0.325996 (rank : 4) | θ value | 8.40245e-06 (rank : 4) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96AY2, Q96N62 | Gene names | EME1, MMS4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crossover junction endonuclease EME1 (EC 3.1.22.-) (hMMS4). | |||||
|
UN45B_HUMAN
|
||||||
| NC score | 0.038755 (rank : 5) | θ value | 1.06291 (rank : 5) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IWX7, Q495Q8, Q495Q9 | Gene names | UNC45B, CMYA4, UNC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog B (UNC-45B). | |||||
|
UN45B_MOUSE
|
||||||
| NC score | 0.037602 (rank : 6) | θ value | 1.38821 (rank : 6) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CGY6, Q5XG72, Q8BHC5, Q8BWK3 | Gene names | Unc45b, Cmya4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog B (UNC-45B). | |||||
|
IRS1_HUMAN
|
||||||
| NC score | 0.021827 (rank : 7) | θ value | 3.0926 (rank : 9) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35568 | Gene names | IRS1 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin receptor substrate 1 (IRS-1). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.017617 (rank : 8) | θ value | 8.99809 (rank : 20) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
TAF6L_HUMAN
|
||||||
| NC score | 0.017284 (rank : 9) | θ value | 6.88961 (rank : 17) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6J9, Q96HA6 | Gene names | TAF6L, PAF65A | |||
|
Domain Architecture |

|
|||||
| Description | TAF6-like RNA polymerase II p300/CBP-associated factor-associated factor 65 kDa subunit 6L (PCAF-associated factor 65 alpha) (PAF65- alpha). | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.016971 (rank : 10) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
HNRPR_HUMAN
|
||||||
| NC score | 0.014267 (rank : 11) | θ value | 3.0926 (rank : 8) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43390 | Gene names | HNRPR | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein R (hnRNP R). | |||||
|
B4GN4_HUMAN
|
||||||
| NC score | 0.013761 (rank : 12) | θ value | 6.88961 (rank : 16) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q76KP1, Q96LV2 | Gene names | B4GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.012242 (rank : 13) | θ value | 4.03905 (rank : 10) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
PHX2A_MOUSE
|
||||||
| NC score | 0.010406 (rank : 14) | θ value | 4.03905 (rank : 12) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62066 | Gene names | Phox2a, Arix, Phox2, Pmx2, Pmx2a | |||
|
Domain Architecture |

|
|||||
| Description | Paired mesoderm homeobox protein 2A (Paired-like homeobox 2A) (PHOX2A homeodomain protein) (Aristaless homeobox protein homolog). | |||||
|
PHX2A_HUMAN
|
||||||
| NC score | 0.010205 (rank : 15) | θ value | 4.03905 (rank : 11) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14813, Q8IVZ2 | Gene names | PHOX2A, ARIX, PMX2A | |||
|
Domain Architecture |

|
|||||
| Description | Paired mesoderm homeobox protein 2A (Paired-like homeobox 2A) (Aristaless homeobox protein homolog) (ARIX1 homeodomain protein). | |||||
|
SCAM3_MOUSE
|
||||||
| NC score | 0.010102 (rank : 16) | θ value | 8.99809 (rank : 21) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35609, Q99M48, Q9ERM9 | Gene names | Scamp3 | |||
|
Domain Architecture |

|
|||||
| Description | Secretory carrier-associated membrane protein 3 (Secretory carrier membrane protein 3). | |||||
|
MYO5B_HUMAN
|
||||||
| NC score | 0.009591 (rank : 17) | θ value | 2.36792 (rank : 7) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULV0 | Gene names | MYO5B, KIAA1119 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5B (Myosin Vb). | |||||
|
MYH14_HUMAN
|
||||||
| NC score | 0.008581 (rank : 18) | θ value | 5.27518 (rank : 13) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.007425 (rank : 19) | θ value | 8.99809 (rank : 18) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
WNK4_HUMAN
|
||||||
| NC score | 0.003781 (rank : 20) | θ value | 5.27518 (rank : 14) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
ACK1_HUMAN
|
||||||
| NC score | 0.001975 (rank : 21) | θ value | 6.88961 (rank : 15) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1140 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07912, Q8N6U7, Q96H59 | Gene names | TNK2, ACK1 | |||
|
Domain Architecture |

|
|||||
| Description | Activated CDC42 kinase 1 (EC 2.7.10.2) (ACK-1) (Tyrosine kinase non- receptor protein 2). | |||||