Please be patient as the page loads
|
ITAE_HUMAN
|
||||||
| SwissProt Accessions | P38570, Q9NZU9 | Gene names | ITGAE | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-E precursor (Mucosal lymphocyte 1 antigen) (HML-1 antigen) (Integrin alpha-IEL) (CD103 antigen) [Contains: Integrin alpha-E light chain; Integrin alpha-E heavy chain]. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ITAE_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P38570, Q9NZU9 | Gene names | ITGAE | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-E precursor (Mucosal lymphocyte 1 antigen) (HML-1 antigen) (Integrin alpha-IEL) (CD103 antigen) [Contains: Integrin alpha-E light chain; Integrin alpha-E heavy chain]. | |||||
|
ITAE_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.996418 (rank : 2) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q60677 | Gene names | Itgae | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-E precursor (Integrin alpha M290) (CD103 antigen) [Contains: Integrin alpha-E light chain; Integrin alpha-E heavy chain]. | |||||
|
ITAD_MOUSE
|
||||||
| θ value | 1.39067e-109 (rank : 3) | NC score | 0.960998 (rank : 7) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q3V0T4 | Gene names | Itgad | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrin alpha-D precursor (CD11d antigen). | |||||
|
ITAM_HUMAN
|
||||||
| θ value | 3.42535e-108 (rank : 4) | NC score | 0.962048 (rank : 5) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P11215 | Gene names | ITGAM, CD11B, CR3A | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-M precursor (Cell surface glycoprotein MAC-1 alpha subunit) (CR-3 alpha chain) (Leukocyte adhesion receptor MO1) (Neutrophil adherence receptor) (CD11b antigen). | |||||
|
ITAD_HUMAN
|
||||||
| θ value | 2.07809e-105 (rank : 5) | NC score | 0.957937 (rank : 11) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q13349, Q15575, Q15576 | Gene names | ITGAD | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-D precursor (Leukointegrin alpha D) (ADB2) (CD11d antigen). | |||||
|
ITAM_MOUSE
|
||||||
| θ value | 2.2976e-104 (rank : 6) | NC score | 0.960365 (rank : 8) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P05555, Q8CA73 | Gene names | Itgam | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-M precursor (Cell surface glycoprotein MAC-1 alpha subunit) (CR-3 alpha chain) (Leukocyte adhesion receptor MO1) (CD11b antigen). | |||||
|
ITAX_HUMAN
|
||||||
| θ value | 2.46047e-98 (rank : 7) | NC score | 0.957685 (rank : 12) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P20702, Q8IVA6 | Gene names | ITGAX, CD11C | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-X precursor (Leukocyte adhesion glycoprotein p150,95 alpha chain) (Leukocyte adhesion receptor p150,95) (Leu M5) (CD11c antigen). | |||||
|
ITAX_MOUSE
|
||||||
| θ value | 2.08291e-97 (rank : 8) | NC score | 0.961054 (rank : 6) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9QXH4 | Gene names | Itgax | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-X precursor (Leukocyte adhesion glycoprotein p150,95 alpha chain) (Leukocyte adhesion receptor p150,95) (CD11c antigen). | |||||
|
ITAL_MOUSE
|
||||||
| θ value | 1.49618e-87 (rank : 9) | NC score | 0.962773 (rank : 3) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P24063 | Gene names | Itgal, Lfa-1, Ly-15 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-L precursor (Leukocyte adhesion glycoprotein LFA-1 alpha chain) (LFA-1A) (Leukocyte function-associated molecule 1 alpha chain) (Lymphocyte antigen Ly-15) (CD11a antigen). | |||||
|
ITAL_HUMAN
|
||||||
| θ value | 1.2666e-86 (rank : 10) | NC score | 0.962481 (rank : 4) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P20701, O43746, Q45H73, Q9UBC8 | Gene names | ITGAL, CD11A | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-L precursor (Leukocyte adhesion glycoprotein LFA-1 alpha chain) (LFA-1A) (Leukocyte function-associated molecule 1 alpha chain) (CD11a antigen). | |||||
|
ITA10_HUMAN
|
||||||
| θ value | 2.23575e-83 (rank : 11) | NC score | 0.958468 (rank : 9) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O75578, Q6UXJ6, Q9UHZ8 | Gene names | ITGA10 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-10 precursor. | |||||
|
ITA11_HUMAN
|
||||||
| θ value | 2.0926e-81 (rank : 12) | NC score | 0.954089 (rank : 14) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9UKX5, Q8WYI8, Q9UKQ1 | Gene names | ITGA11 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-11 precursor. | |||||
|
ITA2_HUMAN
|
||||||
| θ value | 3.94648e-80 (rank : 13) | NC score | 0.954230 (rank : 13) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P17301, Q14595 | Gene names | ITGA2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
ITA2_MOUSE
|
||||||
| θ value | 5.15425e-80 (rank : 14) | NC score | 0.953904 (rank : 15) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q62469, Q62163 | Gene names | Itga2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
ITA11_MOUSE
|
||||||
| θ value | 1.26953e-78 (rank : 15) | NC score | 0.952491 (rank : 16) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P61622 | Gene names | Itga11 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-11 precursor. | |||||
|
ITA1_HUMAN
|
||||||
| θ value | 5.8972e-76 (rank : 16) | NC score | 0.958106 (rank : 10) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P56199 | Gene names | ITGA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrin alpha-1 (Laminin and collagen receptor) (VLA-1) (CD49a antigen). | |||||
|
ITA9_HUMAN
|
||||||
| θ value | 2.25659e-51 (rank : 17) | NC score | 0.864656 (rank : 17) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13797, Q14638 | Gene names | ITGA9 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-9 precursor (Integrin alpha-RLC). | |||||
|
ITA4_MOUSE
|
||||||
| θ value | 5.20228e-48 (rank : 18) | NC score | 0.858059 (rank : 18) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q00651 | Gene names | Itga4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-4 precursor (Integrin alpha-IV) (VLA-4) (Lymphocyte Peyer patch adhesion molecules subunit alpha) (LPAM subunit alpha) (CD49d antigen). | |||||
|
ITA4_HUMAN
|
||||||
| θ value | 2.41655e-45 (rank : 19) | NC score | 0.855075 (rank : 19) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P13612 | Gene names | ITGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-4 precursor (Integrin alpha-IV) (VLA-4) (CD49d antigen). | |||||
|
ITA3_MOUSE
|
||||||
| θ value | 2.95404e-43 (rank : 20) | NC score | 0.831334 (rank : 20) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62470, Q08441, Q08442 | Gene names | Itga3 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-3 precursor (Galactoprotein B3) (GAPB3) (VLA-3 alpha chain) (CD49c antigen) [Contains: Integrin alpha-3 heavy chain; Integrin alpha-3 light chain]. | |||||
|
ITA3_HUMAN
|
||||||
| θ value | 1.79215e-40 (rank : 21) | NC score | 0.830138 (rank : 21) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P26006 | Gene names | ITGA3 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-3 precursor (Galactoprotein B3) (GAPB3) (VLA-3 alpha chain) (FRP-2) (CD49c antigen) [Contains: Integrin alpha-3 heavy chain; Integrin alpha-3 light chain]. | |||||
|
ITAV_MOUSE
|
||||||
| θ value | 5.76516e-39 (rank : 22) | NC score | 0.806254 (rank : 22) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P43406 | Gene names | Itgav | |||
|
Domain Architecture |
|
|||||
| Description | Integrin alpha-V precursor (Vitronectin receptor subunit alpha) (CD51 antigen) [Contains: Integrin alpha-V heavy chain; Integrin alpha-V light chain]. | |||||
|
ITAV_HUMAN
|
||||||
| θ value | 1.6774e-38 (rank : 23) | NC score | 0.802471 (rank : 24) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P06756 | Gene names | ITGAV, VNRA | |||
|
Domain Architecture |
|
|||||
| Description | Integrin alpha-V precursor (Vitronectin receptor subunit alpha) (CD51 antigen) [Contains: Integrin alpha-V heavy chain; Integrin alpha-V light chain]. | |||||
|
ITA6_HUMAN
|
||||||
| θ value | 6.59618e-35 (rank : 24) | NC score | 0.799801 (rank : 28) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P23229, Q08443, Q14646, Q16508, Q9UN03 | Gene names | ITGA6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-6 precursor (VLA-6) (CD49f antigen) [Contains: Integrin alpha-6 heavy chain; Integrin alpha-6 light chain]. | |||||
|
ITA7_MOUSE
|
||||||
| θ value | 9.52487e-34 (rank : 25) | NC score | 0.794667 (rank : 30) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61738, O88731, O88732, P70350, Q61737, Q61741 | Gene names | Itga7 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-7 precursor [Contains: Integrin alpha-7 heavy chain; Integrin alpha-7 light chain]. | |||||
|
ITA7_HUMAN
|
||||||
| θ value | 1.6247e-33 (rank : 26) | NC score | 0.794803 (rank : 29) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13683, O43197, Q9NY89, Q9UET0, Q9UEV2 | Gene names | ITGA7 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-7 precursor [Contains: Integrin alpha-7 heavy chain; Integrin alpha-7 light chain]. | |||||
|
ITA6_MOUSE
|
||||||
| θ value | 2.12192e-33 (rank : 27) | NC score | 0.803375 (rank : 23) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61739 | Gene names | Itga6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-6 precursor (VLA-6) (CD49f antigen) [Contains: Integrin alpha-6 heavy chain; Integrin alpha-6 light chain]. | |||||
|
ITA8_HUMAN
|
||||||
| θ value | 2.77131e-33 (rank : 28) | NC score | 0.800112 (rank : 27) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P53708, Q5VX94 | Gene names | ITGA8 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-8 precursor [Contains: Integrin alpha-8 heavy chain; Integrin alpha-8 light chain]. | |||||
|
ITA5_HUMAN
|
||||||
| θ value | 4.00176e-32 (rank : 29) | NC score | 0.801158 (rank : 25) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P08648, Q96HA5 | Gene names | ITGA5, FNRA | |||
|
Domain Architecture |
|
|||||
| Description | Integrin alpha-5 precursor (Fibronectin receptor subunit alpha) (Integrin alpha-F) (VLA-5) (CD49e antigen) [Contains: Integrin alpha-5 heavy chain; Integrin alpha-5 light chain]. | |||||
|
ITA5_MOUSE
|
||||||
| θ value | 1.28732e-30 (rank : 30) | NC score | 0.800586 (rank : 26) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P11688 | Gene names | Itga5 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-5 precursor (Fibronectin receptor subunit alpha) (Integrin alpha-F) (VLA-5) (CD49e antigen) [Contains: Integrin alpha-5 heavy chain; Integrin alpha-5 light chain]. | |||||
|
ITA2B_MOUSE
|
||||||
| θ value | 4.14116e-29 (rank : 31) | NC score | 0.789136 (rank : 32) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9QUM0, Q64229, Q9Z2M0 | Gene names | Itga2b | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-IIb precursor (Platelet membrane glycoprotein IIb) (GPalpha IIb) (GPIIb) (CD41 antigen) [Contains: Integrin alpha-IIb heavy chain; Integrin alpha-IIb light chain]. | |||||
|
ITA2B_HUMAN
|
||||||
| θ value | 9.2256e-29 (rank : 32) | NC score | 0.793019 (rank : 31) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P08514, O95366, Q14443 | Gene names | ITGA2B, GP2B, ITGAB | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-IIb precursor (Platelet membrane glycoprotein IIb) (GPalpha IIb) (GPIIb) (CD41 antigen) [Contains: Integrin alpha-IIb heavy chain; Integrin alpha-IIb light chain]. | |||||
|
COCA1_MOUSE
|
||||||
| θ value | 1.33526e-19 (rank : 33) | NC score | 0.394238 (rank : 50) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q60847, P70322 | Gene names | Col12a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
COCA1_HUMAN
|
||||||
| θ value | 1.47631e-18 (rank : 34) | NC score | 0.395648 (rank : 49) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
MATN1_MOUSE
|
||||||
| θ value | 1.80466e-16 (rank : 35) | NC score | 0.570612 (rank : 33) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P51942 | Gene names | Matn1, Cmp, Crtm | |||
|
Domain Architecture |
|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
COKA1_HUMAN
|
||||||
| θ value | 2.35696e-16 (rank : 36) | NC score | 0.408584 (rank : 42) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9P218, Q4VXQ4, Q8WUT2, Q96CY9, Q9BQU6, Q9BQU7 | Gene names | COL20A1, KIAA1510 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XX) chain precursor. | |||||
|
MATN1_HUMAN
|
||||||
| θ value | 6.85773e-16 (rank : 37) | NC score | 0.569478 (rank : 34) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P21941 | Gene names | MATN1, CMP, CRTM | |||
|
Domain Architecture |
|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
COEA1_HUMAN
|
||||||
| θ value | 1.52774e-15 (rank : 38) | NC score | 0.400243 (rank : 46) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q05707, O00260, O00261, O00262, Q05708, Q5XJ18, Q96C67 | Gene names | COL14A1, UND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor (Undulin). | |||||
|
MATN2_HUMAN
|
||||||
| θ value | 1.29331e-14 (rank : 39) | NC score | 0.330337 (rank : 53) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00339, Q6UWA5, Q7Z5X1, Q8NDE6, Q96FT5, Q9NSZ1 | Gene names | MATN2 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-2 precursor. | |||||
|
CO6A3_HUMAN
|
||||||
| θ value | 2.20605e-14 (rank : 40) | NC score | 0.396578 (rank : 48) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P12111, Q16501 | Gene names | COL6A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(VI) chain precursor. | |||||
|
COEA1_MOUSE
|
||||||
| θ value | 4.91457e-14 (rank : 41) | NC score | 0.408463 (rank : 43) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q80X19, Q8C6X3, Q9WV05 | Gene names | Col14a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor. | |||||
|
MATN3_HUMAN
|
||||||
| θ value | 4.91457e-14 (rank : 42) | NC score | 0.403055 (rank : 45) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O15232 | Gene names | MATN3 | |||
|
Domain Architecture |
|
|||||
| Description | Matrilin-3 precursor. | |||||
|
MATN2_MOUSE
|
||||||
| θ value | 6.41864e-14 (rank : 43) | NC score | 0.337224 (rank : 52) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O08746 | Gene names | Matn2 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-2 precursor. | |||||
|
MATN3_MOUSE
|
||||||
| θ value | 2.43908e-13 (rank : 44) | NC score | 0.399691 (rank : 47) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O35701, Q9JHM0 | Gene names | Matn3 | |||
|
Domain Architecture |
|
|||||
| Description | Matrilin-3 precursor. | |||||
|
MATN4_HUMAN
|
||||||
| θ value | 2.0648e-12 (rank : 45) | NC score | 0.391172 (rank : 51) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95460, Q5QPU2, Q5QPU3, Q5QPU4, Q8N2M5, Q8N2M7, Q9H1F8, Q9H1F9 | Gene names | MATN4 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-4 precursor. | |||||
|
MATN4_MOUSE
|
||||||
| θ value | 1.02475e-11 (rank : 46) | NC score | 0.405921 (rank : 44) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O89029, O89030, Q9QWS3 | Gene names | Matn4 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-4 precursor (MAT-4). | |||||
|
VITRN_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 47) | NC score | 0.568752 (rank : 35) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VHI5, Q3TZ47, Q8BQ41, Q8K047, Q9CYZ1 | Gene names | Vit | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vitrin precursor. | |||||
|
CO7A1_HUMAN
|
||||||
| θ value | 3.29651e-10 (rank : 48) | NC score | 0.257157 (rank : 56) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
COCH_HUMAN
|
||||||
| θ value | 3.29651e-10 (rank : 49) | NC score | 0.546783 (rank : 37) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O43405 | Gene names | COCH, COCH5B2 | |||
|
Domain Architecture |

|
|||||
| Description | Cochlin precursor (COCH-5B2). | |||||
|
COCH_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 50) | NC score | 0.546038 (rank : 38) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62507, Q3TAF5, Q9QWK6 | Gene names | Coch | |||
|
Domain Architecture |

|
|||||
| Description | Cochlin precursor (COCH-5B2). | |||||
|
VITRN_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 51) | NC score | 0.560386 (rank : 36) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6UXI7, Q6P7T3, Q96DT1 | Gene names | VIT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vitrin precursor. | |||||
|
PHLD_MOUSE
|
||||||
| θ value | 1.99992e-07 (rank : 52) | NC score | 0.506019 (rank : 39) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O70362 | Gene names | Gpld1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 1 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
VWF_HUMAN
|
||||||
| θ value | 5.81887e-07 (rank : 53) | NC score | 0.300228 (rank : 55) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P04275 | Gene names | VWF, F8VWF | |||
|
Domain Architecture |

|
|||||
| Description | von Willebrand factor precursor (vWF) [Contains: von Willebrand antigen 2 (von Willebrand antigen II)]. | |||||
|
VWF_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 54) | NC score | 0.321224 (rank : 54) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8CIZ8, Q60863, Q6XUV6, Q8BIU9, Q8CGN0, Q9JK16 | Gene names | Vwf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | von Willebrand factor precursor (vWF) [Contains: von Willebrand antigen 2 (von Willebrand antigen II)]. | |||||
|
PHLD1_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 55) | NC score | 0.487544 (rank : 40) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P80108, Q15128 | Gene names | GPLD1, PIGPLD1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 1 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
PHLD2_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 56) | NC score | 0.473847 (rank : 41) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q15127 | Gene names | GPLD2, PIGPLD2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 2 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
CO6A2_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 57) | NC score | 0.219167 (rank : 58) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P12110, Q13909, Q13910, Q13911, Q14049, Q16259, Q16597, Q9UML3 | Gene names | COL6A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
CO6A1_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 58) | NC score | 0.206162 (rank : 59) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q04857 | Gene names | Col6a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VI) chain precursor. | |||||
|
CO6A2_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 59) | NC score | 0.219673 (rank : 57) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q02788, Q05505 | Gene names | Col6a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
CO6A1_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 60) | NC score | 0.201721 (rank : 60) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P12109, O00117, O00118, Q14040, Q14041, Q16258 | Gene names | COL6A1 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-1(VI) chain precursor. | |||||
|
KRHB1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 61) | NC score | 0.004870 (rank : 98) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14533, Q14846, Q16274, Q8WU52, Q9BR74 | Gene names | KRTHB1, MLN137 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type II cuticular Hb1 (Hair keratin, type II Hb1) (ghHKb1) (ghHb1) (MLN 137). | |||||
|
IRK10_HUMAN
|
||||||
| θ value | 0.813845 (rank : 62) | NC score | 0.010233 (rank : 93) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78508, Q8N4I7, Q92808 | Gene names | KCNJ10 | |||
|
Domain Architecture |
|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 10 (Potassium channel, inwardly rectifying subfamily J member 10) (Inward rectifier K(+) channel Kir1.2) (ATP-dependent inwardly rectifying potassium channel Kir4.1). | |||||
|
IRK10_MOUSE
|
||||||
| θ value | 0.813845 (rank : 63) | NC score | 0.010242 (rank : 92) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JM63 | Gene names | Kcnj10 | |||
|
Domain Architecture |
|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 10 (Potassium channel, inwardly rectifying subfamily J member 10) (Inward rectifier K(+) channel Kir4.1). | |||||
|
AFF2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 64) | NC score | 0.010843 (rank : 91) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
MYH10_HUMAN
|
||||||
| θ value | 1.06291 (rank : 65) | NC score | 0.003045 (rank : 101) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH10_MOUSE
|
||||||
| θ value | 1.06291 (rank : 66) | NC score | 0.003072 (rank : 100) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 1.06291 (rank : 67) | NC score | 0.002973 (rank : 102) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 1.06291 (rank : 68) | NC score | 0.002526 (rank : 103) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
KRHB3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 69) | NC score | 0.004039 (rank : 99) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P78385, Q9NSB3 | Gene names | KRTHB3 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type II cuticular Hb3 (Hair keratin, type II Hb3). | |||||
|
PSMD2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.013317 (rank : 89) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13200, Q12932, Q15321, Q96I12 | Gene names | PSMD2, TRAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 2 (26S proteasome regulatory subunit RPN1) (26S proteasome regulatory subunit S2) (26S proteasome subunit p97) (Tumor necrosis factor type 1 receptor- associated protein 2) (55.11 protein). | |||||
|
PSMD2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.013161 (rank : 90) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VDM4 | Gene names | Psmd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 2 (26S proteasome regulatory subunit RPN1) (26S proteasome regulatory subunit S2) (26S proteasome subunit p97). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.001714 (rank : 104) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
ADFP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.007734 (rank : 94) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99541, Q9BSC3 | Gene names | ADFP | |||
|
Domain Architecture |

|
|||||
| Description | Adipophilin (Adipose differentiation-related protein) (ADRP). | |||||
|
ADFP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.007181 (rank : 95) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P43883 | Gene names | Adfp, Adrp | |||
|
Domain Architecture |

|
|||||
| Description | Adipophilin (Adipose differentiation-related protein) (ADRP). | |||||
|
DDX21_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | -0.000770 (rank : 105) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
IRK14_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.005630 (rank : 97) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UNX9 | Gene names | KCNJ14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-sensitive inward rectifier potassium channel 14 (Potassium channel, inwardly rectifying subfamily J member 14) (Inward rectifier K(+) channel Kir2.4) (IRK4). | |||||
|
IRK14_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.005638 (rank : 96) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8JZN3, Q8BMK3, Q8BXM0 | Gene names | Kcnj14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-sensitive inward rectifier potassium channel 14 (Potassium channel, inwardly rectifying subfamily J member 14) (Inward rectifier K(+) channel Kir2.4) (IRK4). | |||||
|
ANTR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.057404 (rank : 72) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H6X2, Q96P02, Q9NVP3 | Gene names | ANTXR1, ATR, TEM8 | |||
|
Domain Architecture |
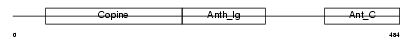
|
|||||
| Description | Anthrax toxin receptor 1 precursor (Tumor endothelial marker 8). | |||||
|
ANTR1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.054236 (rank : 85) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CZ52 | Gene names | Antxr1, Atr, Tem8 | |||
|
Domain Architecture |
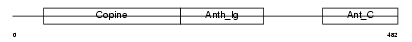
|
|||||
| Description | Anthrax toxin receptor 1 precursor (Tumor endothelial marker 8). | |||||
|
ANTR2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.064083 (rank : 65) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P58335, Q59E98, Q5JPE9, Q86UI1, Q8N4J8, Q8NB13, Q96NC7 | Gene names | ANTXR2, CMG2 | |||
|
Domain Architecture |
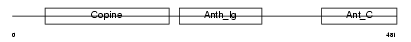
|
|||||
| Description | Anthrax toxin receptor 2 precursor (Capillary morphogenesis gene 2 protein) (CMG-2). | |||||
|
CFAB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.055405 (rank : 80) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P04186 | Gene names | Cfb, Bf, H2-Bf | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor B precursor (EC 3.4.21.47) (C3/C5 convertase) [Contains: Complement factor B Ba fragment; Complement factor B Bb fragment]. | |||||
|
CO4A1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.055647 (rank : 78) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02462 | Gene names | COL4A1 | |||
|
Domain Architecture |
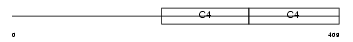
|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CO4A1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.055237 (rank : 81) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CO4A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.057048 (rank : 74) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P08572 | Gene names | COL4A2 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
CO4A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.056285 (rank : 76) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P08122, Q61375 | Gene names | Col4a2 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
CO4A3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.057014 (rank : 75) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
CO4A4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.054820 (rank : 83) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
CO4A6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.055558 (rank : 79) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14031, Q12823, Q14053, Q5JYH6, Q9NQM5, Q9NTX3, Q9UJ76, Q9UMG6, Q9Y4L4 | Gene names | COL4A6 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-6(IV) chain precursor. | |||||
|
CO5A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.054113 (rank : 86) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
CO5A3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.051737 (rank : 87) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P25940, Q9NZQ6 | Gene names | COL5A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(V) chain precursor. | |||||
|
CO9A1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.060300 (rank : 68) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P20849, Q13699, Q13700, Q5TF52, Q6P467, Q96BM8, Q99225, Q9H151, Q9H152, Q9Y6P2, Q9Y6P3 | Gene names | COL9A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
CO9A1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.060534 (rank : 67) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
CO9A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.054830 (rank : 82) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14055 | Gene names | COL9A2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
CO9A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.057615 (rank : 71) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q07643 | Gene names | Col9a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
CO9A3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.062681 (rank : 66) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14050, Q13681, Q9H4G9, Q9UPE2 | Gene names | COL9A3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-3(IX) chain precursor. | |||||
|
COBA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.054339 (rank : 84) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
COBA2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.056161 (rank : 77) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
COGA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.058267 (rank : 70) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q07092 | Gene names | COL16A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVI) chain precursor. | |||||
|
COJA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.064641 (rank : 64) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14993, Q00559, Q05850, Q12885, Q13676, Q5JUF0, Q5T424, Q9H572, Q9NPZ2, Q9NQP2 | Gene names | COL19A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XIX) chain precursor (Collagen alpha-1(Y) chain). | |||||
|
CONA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.065531 (rank : 61) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86Y22, Q8IVR4, Q9NT93 | Gene names | COL23A1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
CONA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.057204 (rank : 73) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K4G2, Q5SUQ0 | Gene names | Col23a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
FINC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.065338 (rank : 63) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P02751, O95609, O95610, Q14312, Q14325, Q14326, Q6LDP6, Q86T27, Q8IVI8, Q96KP7, Q96KP8, Q96KP9, Q9H1B8, Q9HAP3, Q9UMK2 | Gene names | FN1, FN | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN) (Cold-insoluble globulin) (CIG). | |||||
|
FINC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.065361 (rank : 62) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11276, Q61567, Q61568, Q61569, Q64233, Q80UI4 | Gene names | Fn1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN). | |||||
|
MARCO_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.058456 (rank : 69) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60754 | Gene names | Marco | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage receptor MARCO (Macrophage receptor with collagenous structure). | |||||
|
TENN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.051011 (rank : 88) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80Z71 | Gene names | Tnn | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-N precursor (TN-N). | |||||
|
ITAE_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P38570, Q9NZU9 | Gene names | ITGAE | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-E precursor (Mucosal lymphocyte 1 antigen) (HML-1 antigen) (Integrin alpha-IEL) (CD103 antigen) [Contains: Integrin alpha-E light chain; Integrin alpha-E heavy chain]. | |||||
|
ITAE_MOUSE
|
||||||
| NC score | 0.996418 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q60677 | Gene names | Itgae | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-E precursor (Integrin alpha M290) (CD103 antigen) [Contains: Integrin alpha-E light chain; Integrin alpha-E heavy chain]. | |||||
|
ITAL_MOUSE
|
||||||
| NC score | 0.962773 (rank : 3) | θ value | 1.49618e-87 (rank : 9) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P24063 | Gene names | Itgal, Lfa-1, Ly-15 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-L precursor (Leukocyte adhesion glycoprotein LFA-1 alpha chain) (LFA-1A) (Leukocyte function-associated molecule 1 alpha chain) (Lymphocyte antigen Ly-15) (CD11a antigen). | |||||
|
ITAL_HUMAN
|
||||||
| NC score | 0.962481 (rank : 4) | θ value | 1.2666e-86 (rank : 10) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P20701, O43746, Q45H73, Q9UBC8 | Gene names | ITGAL, CD11A | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-L precursor (Leukocyte adhesion glycoprotein LFA-1 alpha chain) (LFA-1A) (Leukocyte function-associated molecule 1 alpha chain) (CD11a antigen). | |||||
|
ITAM_HUMAN
|
||||||
| NC score | 0.962048 (rank : 5) | θ value | 3.42535e-108 (rank : 4) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P11215 | Gene names | ITGAM, CD11B, CR3A | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-M precursor (Cell surface glycoprotein MAC-1 alpha subunit) (CR-3 alpha chain) (Leukocyte adhesion receptor MO1) (Neutrophil adherence receptor) (CD11b antigen). | |||||
|
ITAX_MOUSE
|
||||||
| NC score | 0.961054 (rank : 6) | θ value | 2.08291e-97 (rank : 8) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9QXH4 | Gene names | Itgax | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-X precursor (Leukocyte adhesion glycoprotein p150,95 alpha chain) (Leukocyte adhesion receptor p150,95) (CD11c antigen). | |||||
|
ITAD_MOUSE
|
||||||
| NC score | 0.960998 (rank : 7) | θ value | 1.39067e-109 (rank : 3) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q3V0T4 | Gene names | Itgad | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrin alpha-D precursor (CD11d antigen). | |||||
|
ITAM_MOUSE
|
||||||
| NC score | 0.960365 (rank : 8) | θ value | 2.2976e-104 (rank : 6) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P05555, Q8CA73 | Gene names | Itgam | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-M precursor (Cell surface glycoprotein MAC-1 alpha subunit) (CR-3 alpha chain) (Leukocyte adhesion receptor MO1) (CD11b antigen). | |||||
|
ITA10_HUMAN
|
||||||
| NC score | 0.958468 (rank : 9) | θ value | 2.23575e-83 (rank : 11) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O75578, Q6UXJ6, Q9UHZ8 | Gene names | ITGA10 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-10 precursor. | |||||
|
ITA1_HUMAN
|
||||||
| NC score | 0.958106 (rank : 10) | θ value | 5.8972e-76 (rank : 16) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P56199 | Gene names | ITGA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrin alpha-1 (Laminin and collagen receptor) (VLA-1) (CD49a antigen). | |||||
|
ITAD_HUMAN
|
||||||
| NC score | 0.957937 (rank : 11) | θ value | 2.07809e-105 (rank : 5) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q13349, Q15575, Q15576 | Gene names | ITGAD | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-D precursor (Leukointegrin alpha D) (ADB2) (CD11d antigen). | |||||
|
ITAX_HUMAN
|
||||||
| NC score | 0.957685 (rank : 12) | θ value | 2.46047e-98 (rank : 7) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P20702, Q8IVA6 | Gene names | ITGAX, CD11C | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-X precursor (Leukocyte adhesion glycoprotein p150,95 alpha chain) (Leukocyte adhesion receptor p150,95) (Leu M5) (CD11c antigen). | |||||
|
ITA2_HUMAN
|
||||||
| NC score | 0.954230 (rank : 13) | θ value | 3.94648e-80 (rank : 13) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P17301, Q14595 | Gene names | ITGA2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
ITA11_HUMAN
|
||||||
| NC score | 0.954089 (rank : 14) | θ value | 2.0926e-81 (rank : 12) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9UKX5, Q8WYI8, Q9UKQ1 | Gene names | ITGA11 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-11 precursor. | |||||
|
ITA2_MOUSE
|
||||||
| NC score | 0.953904 (rank : 15) | θ value | 5.15425e-80 (rank : 14) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q62469, Q62163 | Gene names | Itga2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
ITA11_MOUSE
|
||||||
| NC score | 0.952491 (rank : 16) | θ value | 1.26953e-78 (rank : 15) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P61622 | Gene names | Itga11 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-11 precursor. | |||||
|
ITA9_HUMAN
|
||||||
| NC score | 0.864656 (rank : 17) | θ value | 2.25659e-51 (rank : 17) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13797, Q14638 | Gene names | ITGA9 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-9 precursor (Integrin alpha-RLC). | |||||
|
ITA4_MOUSE
|
||||||
| NC score | 0.858059 (rank : 18) | θ value | 5.20228e-48 (rank : 18) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q00651 | Gene names | Itga4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-4 precursor (Integrin alpha-IV) (VLA-4) (Lymphocyte Peyer patch adhesion molecules subunit alpha) (LPAM subunit alpha) (CD49d antigen). | |||||
|
ITA4_HUMAN
|
||||||
| NC score | 0.855075 (rank : 19) | θ value | 2.41655e-45 (rank : 19) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P13612 | Gene names | ITGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-4 precursor (Integrin alpha-IV) (VLA-4) (CD49d antigen). | |||||
|
ITA3_MOUSE
|
||||||
| NC score | 0.831334 (rank : 20) | θ value | 2.95404e-43 (rank : 20) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62470, Q08441, Q08442 | Gene names | Itga3 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-3 precursor (Galactoprotein B3) (GAPB3) (VLA-3 alpha chain) (CD49c antigen) [Contains: Integrin alpha-3 heavy chain; Integrin alpha-3 light chain]. | |||||
|
ITA3_HUMAN
|
||||||
| NC score | 0.830138 (rank : 21) | θ value | 1.79215e-40 (rank : 21) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P26006 | Gene names | ITGA3 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-3 precursor (Galactoprotein B3) (GAPB3) (VLA-3 alpha chain) (FRP-2) (CD49c antigen) [Contains: Integrin alpha-3 heavy chain; Integrin alpha-3 light chain]. | |||||
|
ITAV_MOUSE
|
||||||
| NC score | 0.806254 (rank : 22) | θ value | 5.76516e-39 (rank : 22) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P43406 | Gene names | Itgav | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-V precursor (Vitronectin receptor subunit alpha) (CD51 antigen) [Contains: Integrin alpha-V heavy chain; Integrin alpha-V light chain]. | |||||
|
ITA6_MOUSE
|
||||||
| NC score | 0.803375 (rank : 23) | θ value | 2.12192e-33 (rank : 27) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61739 | Gene names | Itga6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-6 precursor (VLA-6) (CD49f antigen) [Contains: Integrin alpha-6 heavy chain; Integrin alpha-6 light chain]. | |||||
|
ITAV_HUMAN
|
||||||
| NC score | 0.802471 (rank : 24) | θ value | 1.6774e-38 (rank : 23) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P06756 | Gene names | ITGAV, VNRA | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-V precursor (Vitronectin receptor subunit alpha) (CD51 antigen) [Contains: Integrin alpha-V heavy chain; Integrin alpha-V light chain]. | |||||
|
ITA5_HUMAN
|
||||||
| NC score | 0.801158 (rank : 25) | θ value | 4.00176e-32 (rank : 29) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P08648, Q96HA5 | Gene names | ITGA5, FNRA | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-5 precursor (Fibronectin receptor subunit alpha) (Integrin alpha-F) (VLA-5) (CD49e antigen) [Contains: Integrin alpha-5 heavy chain; Integrin alpha-5 light chain]. | |||||
|
ITA5_MOUSE
|
||||||
| NC score | 0.800586 (rank : 26) | θ value | 1.28732e-30 (rank : 30) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P11688 | Gene names | Itga5 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-5 precursor (Fibronectin receptor subunit alpha) (Integrin alpha-F) (VLA-5) (CD49e antigen) [Contains: Integrin alpha-5 heavy chain; Integrin alpha-5 light chain]. | |||||
|
ITA8_HUMAN
|
||||||
| NC score | 0.800112 (rank : 27) | θ value | 2.77131e-33 (rank : 28) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P53708, Q5VX94 | Gene names | ITGA8 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-8 precursor [Contains: Integrin alpha-8 heavy chain; Integrin alpha-8 light chain]. | |||||
|
ITA6_HUMAN
|
||||||
| NC score | 0.799801 (rank : 28) | θ value | 6.59618e-35 (rank : 24) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P23229, Q08443, Q14646, Q16508, Q9UN03 | Gene names | ITGA6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-6 precursor (VLA-6) (CD49f antigen) [Contains: Integrin alpha-6 heavy chain; Integrin alpha-6 light chain]. | |||||
|
ITA7_HUMAN
|
||||||
| NC score | 0.794803 (rank : 29) | θ value | 1.6247e-33 (rank : 26) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13683, O43197, Q9NY89, Q9UET0, Q9UEV2 | Gene names | ITGA7 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-7 precursor [Contains: Integrin alpha-7 heavy chain; Integrin alpha-7 light chain]. | |||||
|
ITA7_MOUSE
|
||||||
| NC score | 0.794667 (rank : 30) | θ value | 9.52487e-34 (rank : 25) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61738, O88731, O88732, P70350, Q61737, Q61741 | Gene names | Itga7 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-7 precursor [Contains: Integrin alpha-7 heavy chain; Integrin alpha-7 light chain]. | |||||
|
ITA2B_HUMAN
|
||||||
| NC score | 0.793019 (rank : 31) | θ value | 9.2256e-29 (rank : 32) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P08514, O95366, Q14443 | Gene names | ITGA2B, GP2B, ITGAB | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-IIb precursor (Platelet membrane glycoprotein IIb) (GPalpha IIb) (GPIIb) (CD41 antigen) [Contains: Integrin alpha-IIb heavy chain; Integrin alpha-IIb light chain]. | |||||
|
ITA2B_MOUSE
|
||||||
| NC score | 0.789136 (rank : 32) | θ value | 4.14116e-29 (rank : 31) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9QUM0, Q64229, Q9Z2M0 | Gene names | Itga2b | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-IIb precursor (Platelet membrane glycoprotein IIb) (GPalpha IIb) (GPIIb) (CD41 antigen) [Contains: Integrin alpha-IIb heavy chain; Integrin alpha-IIb light chain]. | |||||
|
MATN1_MOUSE
|
||||||
| NC score | 0.570612 (rank : 33) | θ value | 1.80466e-16 (rank : 35) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P51942 | Gene names | Matn1, Cmp, Crtm | |||
|
Domain Architecture |

|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
MATN1_HUMAN
|
||||||
| NC score | 0.569478 (rank : 34) | θ value | 6.85773e-16 (rank : 37) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P21941 | Gene names | MATN1, CMP, CRTM | |||
|
Domain Architecture |

|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
VITRN_MOUSE
|
||||||
| NC score | 0.568752 (rank : 35) | θ value | 1.133e-10 (rank : 47) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VHI5, Q3TZ47, Q8BQ41, Q8K047, Q9CYZ1 | Gene names | Vit | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vitrin precursor. | |||||
|
VITRN_HUMAN
|
||||||
| NC score | 0.560386 (rank : 36) | θ value | 1.25267e-09 (rank : 51) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6UXI7, Q6P7T3, Q96DT1 | Gene names | VIT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vitrin precursor. | |||||
|
COCH_HUMAN
|
||||||
| NC score | 0.546783 (rank : 37) | θ value | 3.29651e-10 (rank : 49) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O43405 | Gene names | COCH, COCH5B2 | |||
|
Domain Architecture |

|
|||||
| Description | Cochlin precursor (COCH-5B2). | |||||
|
COCH_MOUSE
|
||||||
| NC score | 0.546038 (rank : 38) | θ value | 9.59137e-10 (rank : 50) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62507, Q3TAF5, Q9QWK6 | Gene names | Coch | |||
|
Domain Architecture |

|
|||||
| Description | Cochlin precursor (COCH-5B2). | |||||
|
PHLD_MOUSE
|
||||||
| NC score | 0.506019 (rank : 39) | θ value | 1.99992e-07 (rank : 52) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O70362 | Gene names | Gpld1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 1 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
PHLD1_HUMAN
|
||||||
| NC score | 0.487544 (rank : 40) | θ value | 9.29e-05 (rank : 55) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P80108, Q15128 | Gene names | GPLD1, PIGPLD1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 1 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
PHLD2_HUMAN
|
||||||
| NC score | 0.473847 (rank : 41) | θ value | 0.000270298 (rank : 56) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q15127 | Gene names | GPLD2, PIGPLD2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 2 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
COKA1_HUMAN
|
||||||
| NC score | 0.408584 (rank : 42) | θ value | 2.35696e-16 (rank : 36) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9P218, Q4VXQ4, Q8WUT2, Q96CY9, Q9BQU6, Q9BQU7 | Gene names | COL20A1, KIAA1510 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XX) chain precursor. | |||||
|
COEA1_MOUSE
|
||||||
| NC score | 0.408463 (rank : 43) | θ value | 4.91457e-14 (rank : 41) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q80X19, Q8C6X3, Q9WV05 | Gene names | Col14a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor. | |||||
|
MATN4_MOUSE
|
||||||
| NC score | 0.405921 (rank : 44) | θ value | 1.02475e-11 (rank : 46) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O89029, O89030, Q9QWS3 | Gene names | Matn4 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-4 precursor (MAT-4). | |||||
|
MATN3_HUMAN
|
||||||
| NC score | 0.403055 (rank : 45) | θ value | 4.91457e-14 (rank : 42) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O15232 | Gene names | MATN3 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-3 precursor. | |||||
|
COEA1_HUMAN
|
||||||
| NC score | 0.400243 (rank : 46) | θ value | 1.52774e-15 (rank : 38) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q05707, O00260, O00261, O00262, Q05708, Q5XJ18, Q96C67 | Gene names | COL14A1, UND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor (Undulin). | |||||
|
MATN3_MOUSE
|
||||||
| NC score | 0.399691 (rank : 47) | θ value | 2.43908e-13 (rank : 44) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O35701, Q9JHM0 | Gene names | Matn3 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-3 precursor. | |||||
|
CO6A3_HUMAN
|
||||||
| NC score | 0.396578 (rank : 48) | θ value | 2.20605e-14 (rank : 40) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P12111, Q16501 | Gene names | COL6A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(VI) chain precursor. | |||||
|
COCA1_HUMAN
|
||||||
| NC score | 0.395648 (rank : 49) | θ value | 1.47631e-18 (rank : 34) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
COCA1_MOUSE
|
||||||
| NC score | 0.394238 (rank : 50) | θ value | 1.33526e-19 (rank : 33) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q60847, P70322 | Gene names | Col12a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
MATN4_HUMAN
|
||||||
| NC score | 0.391172 (rank : 51) | θ value | 2.0648e-12 (rank : 45) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95460, Q5QPU2, Q5QPU3, Q5QPU4, Q8N2M5, Q8N2M7, Q9H1F8, Q9H1F9 | Gene names | MATN4 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-4 precursor. | |||||
|
MATN2_MOUSE
|
||||||
| NC score | 0.337224 (rank : 52) | θ value | 6.41864e-14 (rank : 43) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O08746 | Gene names | Matn2 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-2 precursor. | |||||
|
MATN2_HUMAN
|
||||||
| NC score | 0.330337 (rank : 53) | θ value | 1.29331e-14 (rank : 39) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00339, Q6UWA5, Q7Z5X1, Q8NDE6, Q96FT5, Q9NSZ1 | Gene names | MATN2 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-2 precursor. | |||||
|
VWF_MOUSE
|
||||||
| NC score | 0.321224 (rank : 54) | θ value | 3.77169e-06 (rank : 54) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8CIZ8, Q60863, Q6XUV6, Q8BIU9, Q8CGN0, Q9JK16 | Gene names | Vwf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | von Willebrand factor precursor (vWF) [Contains: von Willebrand antigen 2 (von Willebrand antigen II)]. | |||||
|
VWF_HUMAN
|
||||||
| NC score | 0.300228 (rank : 55) | θ value | 5.81887e-07 (rank : 53) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P04275 | Gene names | VWF, F8VWF | |||
|
Domain Architecture |

|
|||||
| Description | von Willebrand factor precursor (vWF) [Contains: von Willebrand antigen 2 (von Willebrand antigen II)]. | |||||
|
CO7A1_HUMAN
|
||||||
| NC score | 0.257157 (rank : 56) | θ value | 3.29651e-10 (rank : 48) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
CO6A2_MOUSE
|
||||||
| NC score | 0.219673 (rank : 57) | θ value | 0.00298849 (rank : 59) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q02788, Q05505 | Gene names | Col6a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
CO6A2_HUMAN
|
||||||
| NC score | 0.219167 (rank : 58) | θ value | 0.000602161 (rank : 57) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P12110, Q13909, Q13910, Q13911, Q14049, Q16259, Q16597, Q9UML3 | Gene names | COL6A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
CO6A1_MOUSE
|
||||||
| NC score | 0.206162 (rank : 59) | θ value | 0.00298849 (rank : 58) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q04857 | Gene names | Col6a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VI) chain precursor. | |||||
|
CO6A1_HUMAN
|
||||||
| NC score | 0.201721 (rank : 60) | θ value | 0.00869519 (rank : 60) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P12109, O00117, O00118, Q14040, Q14041, Q16258 | Gene names | COL6A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VI) chain precursor. | |||||
|
CONA1_HUMAN
|
||||||
| NC score | 0.065531 (rank : 61) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86Y22, Q8IVR4, Q9NT93 | Gene names | COL23A1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
FINC_MOUSE
|
||||||
| NC score | 0.065361 (rank : 62) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11276, Q61567, Q61568, Q61569, Q64233, Q80UI4 | Gene names | Fn1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN). | |||||
|
FINC_HUMAN
|
||||||
| NC score | 0.065338 (rank : 63) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P02751, O95609, O95610, Q14312, Q14325, Q14326, Q6LDP6, Q86T27, Q8IVI8, Q96KP7, Q96KP8, Q96KP9, Q9H1B8, Q9HAP3, Q9UMK2 | Gene names | FN1, FN | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN) (Cold-insoluble globulin) (CIG). | |||||
|
COJA1_HUMAN
|
||||||
| NC score | 0.064641 (rank : 64) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14993, Q00559, Q05850, Q12885, Q13676, Q5JUF0, Q5T424, Q9H572, Q9NPZ2, Q9NQP2 | Gene names | COL19A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XIX) chain precursor (Collagen alpha-1(Y) chain). | |||||
|
ANTR2_HUMAN
|
||||||
| NC score | 0.064083 (rank : 65) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P58335, Q59E98, Q5JPE9, Q86UI1, Q8N4J8, Q8NB13, Q96NC7 | Gene names | ANTXR2, CMG2 | |||
|
Domain Architecture |

|
|||||
| Description | Anthrax toxin receptor 2 precursor (Capillary morphogenesis gene 2 protein) (CMG-2). | |||||
|
CO9A3_HUMAN
|
||||||
| NC score | 0.062681 (rank : 66) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14050, Q13681, Q9H4G9, Q9UPE2 | Gene names | COL9A3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-3(IX) chain precursor. | |||||
|
CO9A1_MOUSE
|
||||||
| NC score | 0.060534 (rank : 67) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
CO9A1_HUMAN
|
||||||
| NC score | 0.060300 (rank : 68) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P20849, Q13699, Q13700, Q5TF52, Q6P467, Q96BM8, Q99225, Q9H151, Q9H152, Q9Y6P2, Q9Y6P3 | Gene names | COL9A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
MARCO_MOUSE
|
||||||
| NC score | 0.058456 (rank : 69) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60754 | Gene names | Marco | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage receptor MARCO (Macrophage receptor with collagenous structure). | |||||
|
COGA1_HUMAN
|
||||||
| NC score | 0.058267 (rank : 70) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q07092 | Gene names | COL16A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVI) chain precursor. | |||||
|
CO9A2_MOUSE
|
||||||
| NC score | 0.057615 (rank : 71) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q07643 | Gene names | Col9a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
ANTR1_HUMAN
|
||||||
| NC score | 0.057404 (rank : 72) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H6X2, Q96P02, Q9NVP3 | Gene names | ANTXR1, ATR, TEM8 | |||
|
Domain Architecture |

|
|||||
| Description | Anthrax toxin receptor 1 precursor (Tumor endothelial marker 8). | |||||
|
CONA1_MOUSE
|
||||||
| NC score | 0.057204 (rank : 73) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K4G2, Q5SUQ0 | Gene names | Col23a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
CO4A2_HUMAN
|
||||||
| NC score | 0.057048 (rank : 74) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P08572 | Gene names | COL4A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
CO4A3_HUMAN
|
||||||
| NC score | 0.057014 (rank : 75) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
CO4A2_MOUSE
|
||||||
| NC score | 0.056285 (rank : 76) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P08122, Q61375 | Gene names | Col4a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
COBA2_MOUSE
|
||||||
| NC score | 0.056161 (rank : 77) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
CO4A1_HUMAN
|
||||||
| NC score | 0.055647 (rank : 78) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02462 | Gene names | COL4A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CO4A6_HUMAN
|
||||||
| NC score | 0.055558 (rank : 79) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14031, Q12823, Q14053, Q5JYH6, Q9NQM5, Q9NTX3, Q9UJ76, Q9UMG6, Q9Y4L4 | Gene names | COL4A6 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-6(IV) chain precursor. | |||||
|
CFAB_MOUSE
|
||||||
| NC score | 0.055405 (rank : 80) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P04186 | Gene names | Cfb, Bf, H2-Bf | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor B precursor (EC 3.4.21.47) (C3/C5 convertase) [Contains: Complement factor B Ba fragment; Complement factor B Bb fragment]. | |||||
|
CO4A1_MOUSE
|
||||||
| NC score | 0.055237 (rank : 81) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CO9A2_HUMAN
|
||||||
| NC score | 0.054830 (rank : 82) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14055 | Gene names | COL9A2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
CO4A4_HUMAN
|
||||||
| NC score | 0.054820 (rank : 83) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
COBA2_HUMAN
|
||||||
| NC score | 0.054339 (rank : 84) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
ANTR1_MOUSE
|
||||||
| NC score | 0.054236 (rank : 85) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CZ52 | Gene names | Antxr1, Atr, Tem8 | |||
|
Domain Architecture |

|
|||||
| Description | Anthrax toxin receptor 1 precursor (Tumor endothelial marker 8). | |||||
|
CO5A2_HUMAN
|
||||||
| NC score | 0.054113 (rank : 86) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
CO5A3_HUMAN
|
||||||
| NC score | 0.051737 (rank : 87) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P25940, Q9NZQ6 | Gene names | COL5A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(V) chain precursor. | |||||
|
TENN_MOUSE
|
||||||
| NC score | 0.051011 (rank : 88) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80Z71 | Gene names | Tnn | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-N precursor (TN-N). | |||||
|
PSMD2_HUMAN
|
||||||
| NC score | 0.013317 (rank : 89) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13200, Q12932, Q15321, Q96I12 | Gene names | PSMD2, TRAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 2 (26S proteasome regulatory subunit RPN1) (26S proteasome regulatory subunit S2) (26S proteasome subunit p97) (Tumor necrosis factor type 1 receptor- associated protein 2) (55.11 protein). | |||||
|
PSMD2_MOUSE
|
||||||
| NC score | 0.013161 (rank : 90) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VDM4 | Gene names | Psmd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 2 (26S proteasome regulatory subunit RPN1) (26S proteasome regulatory subunit S2) (26S proteasome subunit p97). | |||||
|
AFF2_HUMAN
|
||||||
| NC score | 0.010843 (rank : 91) | θ value | 1.06291 (rank : 64) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
IRK10_MOUSE
|
||||||
| NC score | 0.010242 (rank : 92) | θ value | 0.813845 (rank : 63) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JM63 | Gene names | Kcnj10 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 10 (Potassium channel, inwardly rectifying subfamily J member 10) (Inward rectifier K(+) channel Kir4.1). | |||||
|
IRK10_HUMAN
|
||||||
| NC score | 0.010233 (rank : 93) | θ value | 0.813845 (rank : 62) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78508, Q8N4I7, Q92808 | Gene names | KCNJ10 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 10 (Potassium channel, inwardly rectifying subfamily J member 10) (Inward rectifier K(+) channel Kir1.2) (ATP-dependent inwardly rectifying potassium channel Kir4.1). | |||||
|
ADFP_HUMAN
|
||||||
| NC score | 0.007734 (rank : 94) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99541, Q9BSC3 | Gene names | ADFP | |||
|
Domain Architecture |

|
|||||
| Description | Adipophilin (Adipose differentiation-related protein) (ADRP). | |||||
|
ADFP_MOUSE
|
||||||
| NC score | 0.007181 (rank : 95) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P43883 | Gene names | Adfp, Adrp | |||
|
Domain Architecture |

|
|||||
| Description | Adipophilin (Adipose differentiation-related protein) (ADRP). | |||||
|
IRK14_MOUSE
|
||||||
| NC score | 0.005638 (rank : 96) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8JZN3, Q8BMK3, Q8BXM0 | Gene names | Kcnj14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-sensitive inward rectifier potassium channel 14 (Potassium channel, inwardly rectifying subfamily J member 14) (Inward rectifier K(+) channel Kir2.4) (IRK4). | |||||
|
IRK14_HUMAN
|
||||||
| NC score | 0.005630 (rank : 97) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UNX9 | Gene names | KCNJ14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-sensitive inward rectifier potassium channel 14 (Potassium channel, inwardly rectifying subfamily J member 14) (Inward rectifier K(+) channel Kir2.4) (IRK4). | |||||
|
KRHB1_HUMAN
|
||||||
| NC score | 0.004870 (rank : 98) | θ value | 0.365318 (rank : 61) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14533, Q14846, Q16274, Q8WU52, Q9BR74 | Gene names | KRTHB1, MLN137 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cuticular Hb1 (Hair keratin, type II Hb1) (ghHKb1) (ghHb1) (MLN 137). | |||||
|
KRHB3_HUMAN
|
||||||
| NC score | 0.004039 (rank : 99) | θ value | 1.38821 (rank : 69) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P78385, Q9NSB3 | Gene names | KRTHB3 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cuticular Hb3 (Hair keratin, type II Hb3). | |||||
|
MYH10_MOUSE
|
||||||
| NC score | 0.003072 (rank : 100) | θ value | 1.06291 (rank : 66) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH10_HUMAN
|
||||||
| NC score | 0.003045 (rank : 101) | θ value | 1.06291 (rank : 65) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.002973 (rank : 102) | θ value | 1.06291 (rank : 67) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.002526 (rank : 103) | θ value | 1.06291 (rank : 68) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.001714 (rank : 104) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
DDX21_MOUSE
|
||||||
| NC score | -0.000770 (rank : 105) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||