Please be patient as the page loads
|
INP5E_MOUSE
|
||||||
| SwissProt Accessions | Q9JII1, Q3TCC9 | Gene names | Inpp5e | |||
|
Domain Architecture |

|
|||||
| Description | 72 kDa inositol polyphosphate 5-phosphatase (EC 3.1.3.36) (Phosphatidylinositol-4,5-bisphosphate 5-phosphatase) (Phosphatidylinositol polyphosphate 5-phosphatase type IV). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
INP5E_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.949204 (rank : 2) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NRR6, Q15734 | Gene names | INPP5E | |||
|
Domain Architecture |
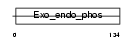
|
|||||
| Description | 72 kDa inositol polyphosphate 5-phosphatase (EC 3.1.3.36) (Phosphatidylinositol-4,5-bisphosphate 5-phosphatase) (Phosphatidylinositol polyphosphate 5-phosphatase type IV). | |||||
|
INP5E_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9JII1, Q3TCC9 | Gene names | Inpp5e | |||
|
Domain Architecture |

|
|||||
| Description | 72 kDa inositol polyphosphate 5-phosphatase (EC 3.1.3.36) (Phosphatidylinositol-4,5-bisphosphate 5-phosphatase) (Phosphatidylinositol polyphosphate 5-phosphatase type IV). | |||||
|
SYNJ2_HUMAN
|
||||||
| θ value | 2.67802e-36 (rank : 3) | NC score | 0.764516 (rank : 4) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15056, Q5TA13, Q5TA16, Q5TA19, Q86XK0, Q8IZA8, Q9H226 | Gene names | SYNJ2, KIAA0348 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptojanin-2 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 2). | |||||
|
SYNJ2_MOUSE
|
||||||
| θ value | 5.0505e-35 (rank : 4) | NC score | 0.768794 (rank : 3) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9D2G5, O35404, O88399, O88400, O88401, O88402, O88403, O88404 | Gene names | Synj2, Kiaa0348 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-2 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 2). | |||||
|
SYNJ1_HUMAN
|
||||||
| θ value | 8.06329e-33 (rank : 5) | NC score | 0.698419 (rank : 11) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
SYNJ1_MOUSE
|
||||||
| θ value | 1.79631e-32 (rank : 6) | NC score | 0.698932 (rank : 10) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
I5P2_HUMAN
|
||||||
| θ value | 5.22648e-32 (rank : 7) | NC score | 0.737969 (rank : 7) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P32019, Q5VSG9, Q5VSH0, Q5VSH1, Q658Q5, Q6P6D4, Q6PD53, Q86YE1 | Gene names | INPP5B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Type II inositol-1,4,5-trisphosphate 5-phosphatase precursor (EC 3.1.3.36) (Phosphoinositide 5-phosphatase) (5PTase) (75 kDa inositol polyphosphate-5-phosphatase). | |||||
|
OCRL_HUMAN
|
||||||
| θ value | 5.59698e-26 (rank : 8) | NC score | 0.734694 (rank : 8) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q01968, O60800, Q15684, Q15774, Q5JQF1, Q5JQF2, Q9UJG5, Q9UMA5 | Gene names | OCRL, INPP5F, OCRL1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol polyphosphate 5-phosphatase OCRL-1 (EC 3.1.3.36) (Lowe oculocerebrorenal syndrome protein). | |||||
|
PI5PA_HUMAN
|
||||||
| θ value | 1.5242e-23 (rank : 9) | NC score | 0.641646 (rank : 12) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
PI5PA_MOUSE
|
||||||
| θ value | 1.6852e-22 (rank : 10) | NC score | 0.704840 (rank : 9) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P59644 | Gene names | Pib5pa | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
SKIP_HUMAN
|
||||||
| θ value | 1.5773e-20 (rank : 11) | NC score | 0.761935 (rank : 5) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BT40, Q15733, Q9NPJ5, Q9P2R5 | Gene names | SKIP, PPS | |||
|
Domain Architecture |

|
|||||
| Description | Skeletal muscle and kidney-enriched inositol phosphatase (EC 3.1.3.56). | |||||
|
SKIP_MOUSE
|
||||||
| θ value | 3.51386e-20 (rank : 12) | NC score | 0.758814 (rank : 6) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8C5L6, O09040 | Gene names | Skip, Pps | |||
|
Domain Architecture |
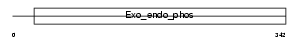
|
|||||
| Description | Skeletal muscle and kidney-enriched inositol phosphatase (EC 3.1.3.56). | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 13) | NC score | 0.110288 (rank : 16) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
BCL9_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 14) | NC score | 0.108554 (rank : 17) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
MKL1_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 15) | NC score | 0.055457 (rank : 34) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
PO121_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 16) | NC score | 0.059007 (rank : 29) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
FOXQ1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 17) | NC score | 0.019025 (rank : 58) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9C009, Q9NS06 | Gene names | FOXQ1, HFH1 | |||
|
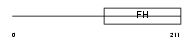
Domain Architecture |
|
|||||
| Description | Forkhead box protein Q1 (Hepatocyte nuclear factor 3 forkhead homolog 1) (HNF-3/forkhead-like protein 1) (HFH-1). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 0.47712 (rank : 18) | NC score | 0.055700 (rank : 33) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 0.47712 (rank : 19) | NC score | 0.066948 (rank : 22) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
GTSE1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.048951 (rank : 38) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8R080, O89015, Q9CSG9 | Gene names | Gtse1, B99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G2 and S phase expressed protein 1 (Gtse-1) (B99 protein). | |||||
|
I5P1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 21) | NC score | 0.206197 (rank : 13) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14642, Q14640 | Gene names | INPP5A | |||
|
Domain Architecture |

|
|||||
| Description | Type I inositol-1,4,5-trisphosphate 5-phosphatase (EC 3.1.3.56) (5PTase). | |||||
|
SYN2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 22) | NC score | 0.031383 (rank : 46) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92777 | Gene names | SYN2 | |||
|
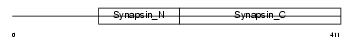
Domain Architecture |
|
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
TAU_HUMAN
|
||||||
| θ value | 0.62314 (rank : 23) | NC score | 0.031198 (rank : 47) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.059464 (rank : 28) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
NFIC_HUMAN
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.018125 (rank : 59) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P08651, P08652, Q14932, Q9UPJ3, Q9UPJ9, Q9UPK0, Q9UPK1 | Gene names | NFIC, NFI | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor 1 C-type (Nuclear factor 1/C) (NF1-C) (NFI-C) (NF-I/C) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.009773 (rank : 70) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
PRDM2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.015401 (rank : 62) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
PYGO2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.036105 (rank : 40) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
BAG3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.031776 (rank : 45) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
BANK1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.023975 (rank : 54) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80VH0, Q3U178, Q8BRV6, Q8BRY8 | Gene names | Bank1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell scaffold protein with ankyrin repeats (Protein AVIEF). | |||||
|
CJ047_MOUSE
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.033082 (rank : 43) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C5R2, Q8R017, Q8R2Z9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf47 homolog. | |||||
|
FA47A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.054598 (rank : 35) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5JRC9, Q8TAA0 | Gene names | FAM47A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47A. | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.013545 (rank : 64) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
PME17_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.026389 (rank : 50) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P40967, Q12763, Q14448, Q14817, Q16565 | Gene names | SILV, D12S53E, PMEL17 | |||
|
Domain Architecture |

|
|||||
| Description | Melanocyte protein Pmel 17 precursor (Melanocyte lineage-specific antigen GP100) (Melanoma-associated ME20 antigen) (ME20M/ME20S) (ME20- M/ME20-S) (95 kDa melanocyte-specific secreted glycoprotein). | |||||
|
M3K12_HUMAN
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | -0.000264 (rank : 75) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12852, Q86VQ5, Q8WY25 | Gene names | MAP3K12, ZPK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 12 (EC 2.7.11.25) (Mixed lineage kinase) (Leucine-zipper protein kinase) (ZPK) (Dual leucine zipper bearing kinase) (DLK) (MAPK-upstream kinase) (MUK). | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.034559 (rank : 41) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
NRG2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.014845 (rank : 63) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14511 | Gene names | NRG2, NTAK | |||
|
Domain Architecture |

|
|||||
| Description | Pro-neuregulin-2, membrane-bound isoform precursor (Pro-NRG2) [Contains: Neuregulin-2 (NRG-2) (Neural- and thymus-derived activator for ERBB kinases) (NTAK) (Divergent of neuregulin-1) (DON-1)]. | |||||
|
PHLB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.017157 (rank : 61) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 678 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6PDH0, Q80TV2 | Gene names | Phldb1, Kiaa0638, Ll5a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.033730 (rank : 42) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.029724 (rank : 48) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
SEM6B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.011353 (rank : 69) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H3T3, Q9NRK9 | Gene names | SEMA6B, SEMAZ | |||
|
Domain Architecture |
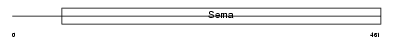
|
|||||
| Description | Semaphorin-6B precursor (Semaphorin Z) (Sema Z). | |||||
|
AMELX_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.029549 (rank : 49) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99217, Q96NW6, Q9UCA7 | Gene names | AMELX, AMG, AMGX | |||
|
Domain Architecture |

|
|||||
| Description | Amelogenin, X isoform precursor. | |||||
|
CIC_HUMAN
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.032413 (rank : 44) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
NRG2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.013404 (rank : 65) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56974 | Gene names | Nrg2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pro-neuregulin-2, membrane-bound isoform precursor (Pro-NRG2) [Contains: Neuregulin-2 (NRG-2) (Divergent of neuregulin 1) (DON-1)]. | |||||
|
RB_MOUSE
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.012919 (rank : 68) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P13405 | Gene names | Rb1, Rb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-associated protein (PP105) (RB). | |||||
|
RTN2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.013288 (rank : 66) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75298, O60509, Q7RTM6, Q7RTN1, Q7RTN2 | Gene names | RTN2, NSPL1 | |||
|
Domain Architecture |
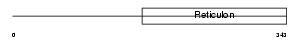
|
|||||
| Description | Reticulon-2 (Neuroendocrine-specific protein-like 1) (NSP-like protein 1) (NSPLI). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.041186 (rank : 39) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
CI066_HUMAN
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.022153 (rank : 56) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5T8R8, Q96NB0 | Gene names | C9orf66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf66. | |||||
|
K1802_HUMAN
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.026279 (rank : 51) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.024957 (rank : 53) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PHLPP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.005018 (rank : 72) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
PO6F1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.003998 (rank : 74) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14863 | Gene names | POU6F1, BRN5, MPOU, TCFB1 | |||
|
Domain Architecture |
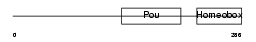
|
|||||
| Description | POU domain, class 6, transcription factor 1 (mPOU homeobox protein) (Brain-specific homeobox/POU domain protein 5) (Brain-5) (Brn-5 protein). | |||||
|
SON_MOUSE
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.026168 (rank : 52) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
SPR2F_HUMAN
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.022586 (rank : 55) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96RM1 | Gene names | SPRR2F | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2F (SPR-2F). | |||||
|
AN13B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.004134 (rank : 73) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5F259, Q6IR40 | Gene names | Ankrd13b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 13B. | |||||
|
CAC1A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.006103 (rank : 71) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
PME17_MOUSE
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.020412 (rank : 57) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60696 | Gene names | Silv, D10H12S53E, Pmel17, Si | |||
|
Domain Architecture |

|
|||||
| Description | Melanocyte protein Pmel 17 precursor (Silver locus protein). | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.017345 (rank : 60) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
TRI66_MOUSE
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.013272 (rank : 67) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
BAT2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.056162 (rank : 30) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CF155_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.062606 (rank : 25) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H8W2, Q5TG13 | Gene names | C6orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf155. | |||||
|
CK024_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.065983 (rank : 23) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96F05, Q9H2K4 | Gene names | C11orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 precursor (Protein DM4E3). | |||||
|
CO039_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.072030 (rank : 20) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.065736 (rank : 24) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.071526 (rank : 21) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
MUC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.060329 (rank : 26) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.074213 (rank : 19) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.060141 (rank : 27) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.056081 (rank : 31) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.052477 (rank : 36) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.079200 (rank : 18) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
RGS3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.055964 (rank : 32) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SAC3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.203257 (rank : 15) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92562, Q53H49, Q5TCS6 | Gene names | SAC3, KIAA0274 | |||
|
Domain Architecture |
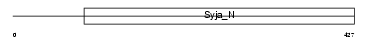
|
|||||
| Description | SAC domain-containing protein 3. | |||||
|
SAC3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.204676 (rank : 14) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91WF7, Q6A092 | Gene names | Sac3, Kiaa0274 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAC domain-containing protein 3. | |||||
|
SON_HUMAN
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.052109 (rank : 37) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
INP5E_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9JII1, Q3TCC9 | Gene names | Inpp5e | |||
|
Domain Architecture |

|
|||||
| Description | 72 kDa inositol polyphosphate 5-phosphatase (EC 3.1.3.36) (Phosphatidylinositol-4,5-bisphosphate 5-phosphatase) (Phosphatidylinositol polyphosphate 5-phosphatase type IV). | |||||
|
INP5E_HUMAN
|
||||||
| NC score | 0.949204 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NRR6, Q15734 | Gene names | INPP5E | |||
|
Domain Architecture |
|
|||||
| Description | 72 kDa inositol polyphosphate 5-phosphatase (EC 3.1.3.36) (Phosphatidylinositol-4,5-bisphosphate 5-phosphatase) (Phosphatidylinositol polyphosphate 5-phosphatase type IV). | |||||
|
SYNJ2_MOUSE
|
||||||
| NC score | 0.768794 (rank : 3) | θ value | 5.0505e-35 (rank : 4) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9D2G5, O35404, O88399, O88400, O88401, O88402, O88403, O88404 | Gene names | Synj2, Kiaa0348 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-2 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 2). | |||||
|
SYNJ2_HUMAN
|
||||||
| NC score | 0.764516 (rank : 4) | θ value | 2.67802e-36 (rank : 3) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15056, Q5TA13, Q5TA16, Q5TA19, Q86XK0, Q8IZA8, Q9H226 | Gene names | SYNJ2, KIAA0348 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptojanin-2 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 2). | |||||
|
SKIP_HUMAN
|
||||||
| NC score | 0.761935 (rank : 5) | θ value | 1.5773e-20 (rank : 11) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BT40, Q15733, Q9NPJ5, Q9P2R5 | Gene names | SKIP, PPS | |||
|
Domain Architecture |

|
|||||
| Description | Skeletal muscle and kidney-enriched inositol phosphatase (EC 3.1.3.56). | |||||
|
SKIP_MOUSE
|
||||||
| NC score | 0.758814 (rank : 6) | θ value | 3.51386e-20 (rank : 12) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8C5L6, O09040 | Gene names | Skip, Pps | |||
|
Domain Architecture |
|
|||||
| Description | Skeletal muscle and kidney-enriched inositol phosphatase (EC 3.1.3.56). | |||||
|
I5P2_HUMAN
|
||||||
| NC score | 0.737969 (rank : 7) | θ value | 5.22648e-32 (rank : 7) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P32019, Q5VSG9, Q5VSH0, Q5VSH1, Q658Q5, Q6P6D4, Q6PD53, Q86YE1 | Gene names | INPP5B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Type II inositol-1,4,5-trisphosphate 5-phosphatase precursor (EC 3.1.3.36) (Phosphoinositide 5-phosphatase) (5PTase) (75 kDa inositol polyphosphate-5-phosphatase). | |||||
|
OCRL_HUMAN
|
||||||
| NC score | 0.734694 (rank : 8) | θ value | 5.59698e-26 (rank : 8) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q01968, O60800, Q15684, Q15774, Q5JQF1, Q5JQF2, Q9UJG5, Q9UMA5 | Gene names | OCRL, INPP5F, OCRL1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol polyphosphate 5-phosphatase OCRL-1 (EC 3.1.3.36) (Lowe oculocerebrorenal syndrome protein). | |||||
|
PI5PA_MOUSE
|
||||||
| NC score | 0.704840 (rank : 9) | θ value | 1.6852e-22 (rank : 10) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P59644 | Gene names | Pib5pa | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
SYNJ1_MOUSE
|
||||||
| NC score | 0.698932 (rank : 10) | θ value | 1.79631e-32 (rank : 6) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
SYNJ1_HUMAN
|
||||||
| NC score | 0.698419 (rank : 11) | θ value | 8.06329e-33 (rank : 5) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
PI5PA_HUMAN
|
||||||
| NC score | 0.641646 (rank : 12) | θ value | 1.5242e-23 (rank : 9) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
I5P1_HUMAN
|
||||||
| NC score | 0.206197 (rank : 13) | θ value | 0.62314 (rank : 21) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14642, Q14640 | Gene names | INPP5A | |||
|
Domain Architecture |

|
|||||
| Description | Type I inositol-1,4,5-trisphosphate 5-phosphatase (EC 3.1.3.56) (5PTase). | |||||
|
SAC3_MOUSE
|
||||||
| NC score | 0.204676 (rank : 14) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91WF7, Q6A092 | Gene names | Sac3, Kiaa0274 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAC domain-containing protein 3. | |||||
|
SAC3_HUMAN
|
||||||
| NC score | 0.203257 (rank : 15) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92562, Q53H49, Q5TCS6 | Gene names | SAC3, KIAA0274 | |||
|
Domain Architecture |

|
|||||
| Description | SAC domain-containing protein 3. | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.110288 (rank : 16) | θ value | 0.00228821 (rank : 13) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
BCL9_MOUSE
|
||||||
| NC score | 0.108554 (rank : 17) | θ value | 0.00298849 (rank : 14) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.079200 (rank : 18) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.074213 (rank : 19) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
CO039_MOUSE
|
||||||
| NC score | 0.072030 (rank : 20) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.071526 (rank : 21) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.066948 (rank : 22) | θ value | 0.47712 (rank : 19) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CK024_HUMAN
|
||||||
| NC score | 0.065983 (rank : 23) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96F05, Q9H2K4 | Gene names | C11orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 precursor (Protein DM4E3). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.065736 (rank : 24) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
CF155_HUMAN
|
||||||
| NC score | 0.062606 (rank : 25) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H8W2, Q5TG13 | Gene names | C6orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf155. | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.060329 (rank : 26) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.060141 (rank : 27) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.059464 (rank : 28) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
PO121_HUMAN
|
||||||
| NC score | 0.059007 (rank : 29) | θ value | 0.0330416 (rank : 16) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.056162 (rank : 30) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.056081 (rank : 31) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.055964 (rank : 32) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.055700 (rank : 33) | θ value | 0.47712 (rank : 18) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
MKL1_MOUSE
|
||||||
| NC score | 0.055457 (rank : 34) | θ value | 0.0252991 (rank : 15) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
FA47A_HUMAN
|
||||||
| NC score | 0.054598 (rank : 35) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5JRC9, Q8TAA0 | Gene names | FAM47A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47A. | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.052477 (rank : 36) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.052109 (rank : 37) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
GTSE1_MOUSE
|
||||||
| NC score | 0.048951 (rank : 38) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8R080, O89015, Q9CSG9 | Gene names | Gtse1, B99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G2 and S phase expressed protein 1 (Gtse-1) (B99 protein). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.041186 (rank : 39) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
PYGO2_HUMAN
|
||||||
| NC score | 0.036105 (rank : 40) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.034559 (rank : 41) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.033730 (rank : 42) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
CJ047_MOUSE
|
||||||
| NC score | 0.033082 (rank : 43) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C5R2, Q8R017, Q8R2Z9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf47 homolog. | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.032413 (rank : 44) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
BAG3_MOUSE
|
||||||
| NC score | 0.031776 (rank : 45) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
SYN2_HUMAN
|
||||||
| NC score | 0.031383 (rank : 46) | θ value | 0.62314 (rank : 22) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92777 | Gene names | SYN2 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
TAU_HUMAN
|
||||||
| NC score | 0.031198 (rank : 47) | θ value | 0.62314 (rank : 23) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.029724 (rank : 48) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
AMELX_HUMAN
|
||||||
| NC score | 0.029549 (rank : 49) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99217, Q96NW6, Q9UCA7 | Gene names | AMELX, AMG, AMGX | |||
|
Domain Architecture |

|
|||||
| Description | Amelogenin, X isoform precursor. | |||||
|
PME17_HUMAN
|
||||||
| NC score | 0.026389 (rank : 50) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P40967, Q12763, Q14448, Q14817, Q16565 | Gene names | SILV, D12S53E, PMEL17 | |||
|
Domain Architecture |

|
|||||
| Description | Melanocyte protein Pmel 17 precursor (Melanocyte lineage-specific antigen GP100) (Melanoma-associated ME20 antigen) (ME20M/ME20S) (ME20- M/ME20-S) (95 kDa melanocyte-specific secreted glycoprotein). | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.026279 (rank : 51) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.026168 (rank : 52) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.024957 (rank : 53) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
BANK1_MOUSE
|
||||||
| NC score | 0.023975 (rank : 54) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80VH0, Q3U178, Q8BRV6, Q8BRY8 | Gene names | Bank1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell scaffold protein with ankyrin repeats (Protein AVIEF). | |||||
|
SPR2F_HUMAN
|
||||||
| NC score | 0.022586 (rank : 55) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96RM1 | Gene names | SPRR2F | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2F (SPR-2F). | |||||
|
CI066_HUMAN
|
||||||
| NC score | 0.022153 (rank : 56) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5T8R8, Q96NB0 | Gene names | C9orf66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf66. | |||||
|
PME17_MOUSE
|
||||||
| NC score | 0.020412 (rank : 57) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60696 | Gene names | Silv, D10H12S53E, Pmel17, Si | |||
|
Domain Architecture |

|
|||||
| Description | Melanocyte protein Pmel 17 precursor (Silver locus protein). | |||||
|
FOXQ1_HUMAN
|
||||||
| NC score | 0.019025 (rank : 58) | θ value | 0.163984 (rank : 17) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9C009, Q9NS06 | Gene names | FOXQ1, HFH1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein Q1 (Hepatocyte nuclear factor 3 forkhead homolog 1) (HNF-3/forkhead-like protein 1) (HFH-1). | |||||
|
NFIC_HUMAN
|
||||||
| NC score | 0.018125 (rank : 59) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P08651, P08652, Q14932, Q9UPJ3, Q9UPJ9, Q9UPK0, Q9UPK1 | Gene names | NFIC, NFI | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor 1 C-type (Nuclear factor 1/C) (NF1-C) (NFI-C) (NF-I/C) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.017345 (rank : 60) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PHLB1_MOUSE
|
||||||
| NC score | 0.017157 (rank : 61) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 678 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6PDH0, Q80TV2 | Gene names | Phldb1, Kiaa0638, Ll5a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
PRDM2_HUMAN
|
||||||
| NC score | 0.015401 (rank : 62) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
NRG2_HUMAN
|
||||||
| NC score | 0.014845 (rank : 63) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14511 | Gene names | NRG2, NTAK | |||
|
Domain Architecture |

|
|||||
| Description | Pro-neuregulin-2, membrane-bound isoform precursor (Pro-NRG2) [Contains: Neuregulin-2 (NRG-2) (Neural- and thymus-derived activator for ERBB kinases) (NTAK) (Divergent of neuregulin-1) (DON-1)]. | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.013545 (rank : 64) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NRG2_MOUSE
|
||||||
| NC score | 0.013404 (rank : 65) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56974 | Gene names | Nrg2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pro-neuregulin-2, membrane-bound isoform precursor (Pro-NRG2) [Contains: Neuregulin-2 (NRG-2) (Divergent of neuregulin 1) (DON-1)]. | |||||
|
RTN2_HUMAN
|
||||||
| NC score | 0.013288 (rank : 66) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75298, O60509, Q7RTM6, Q7RTN1, Q7RTN2 | Gene names | RTN2, NSPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-2 (Neuroendocrine-specific protein-like 1) (NSP-like protein 1) (NSPLI). | |||||
|
TRI66_MOUSE
|
||||||
| NC score | 0.013272 (rank : 67) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
RB_MOUSE
|
||||||
| NC score | 0.012919 (rank : 68) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P13405 | Gene names | Rb1, Rb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-associated protein (PP105) (RB). | |||||
|
SEM6B_HUMAN
|
||||||
| NC score | 0.011353 (rank : 69) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H3T3, Q9NRK9 | Gene names | SEMA6B, SEMAZ | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6B precursor (Semaphorin Z) (Sema Z). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.009773 (rank : 70) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CAC1A_HUMAN
|
||||||
| NC score | 0.006103 (rank : 71) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
PHLPP_HUMAN
|
||||||
| NC score | 0.005018 (rank : 72) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
AN13B_MOUSE
|
||||||
| NC score | 0.004134 (rank : 73) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5F259, Q6IR40 | Gene names | Ankrd13b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 13B. | |||||
|
PO6F1_HUMAN
|
||||||
| NC score | 0.003998 (rank : 74) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14863 | Gene names | POU6F1, BRN5, MPOU, TCFB1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 1 (mPOU homeobox protein) (Brain-specific homeobox/POU domain protein 5) (Brain-5) (Brn-5 protein). | |||||
|
M3K12_HUMAN
|
||||||
| NC score | -0.000264 (rank : 75) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12852, Q86VQ5, Q8WY25 | Gene names | MAP3K12, ZPK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 12 (EC 2.7.11.25) (Mixed lineage kinase) (Leucine-zipper protein kinase) (ZPK) (Dual leucine zipper bearing kinase) (DLK) (MAPK-upstream kinase) (MUK). | |||||