Please be patient as the page loads
|
HMGA2_HUMAN
|
||||||
| SwissProt Accessions | P52926 | Gene names | HMGA2, HMGIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein HMGI-C (High mobility group AT-hook protein 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HMGA2_MOUSE
|
||||||
| θ value | 3.62785e-25 (rank : 1) | NC score | 0.958572 (rank : 2) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P52927, Q3UQW0 | Gene names | Hmga2, Hmgic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein HMGI-C (High mobility group AT-hook protein 2). | |||||
|
HMGA2_HUMAN
|
||||||
| θ value | 1.86321e-21 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P52926 | Gene names | HMGA2, HMGIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein HMGI-C (High mobility group AT-hook protein 2). | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 3) | NC score | 0.040375 (rank : 81) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 4) | NC score | 0.129439 (rank : 6) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 5) | NC score | 0.159601 (rank : 4) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.163984 (rank : 6) | NC score | 0.140400 (rank : 5) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
CJ012_HUMAN
|
||||||
| θ value | 0.279714 (rank : 7) | NC score | 0.160380 (rank : 3) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
PKHA6_HUMAN
|
||||||
| θ value | 0.47712 (rank : 8) | NC score | 0.069678 (rank : 34) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.087144 (rank : 13) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
INVS_MOUSE
|
||||||
| θ value | 1.06291 (rank : 10) | NC score | 0.030957 (rank : 85) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
GON4L_HUMAN
|
||||||
| θ value | 1.38821 (rank : 11) | NC score | 0.065561 (rank : 41) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 12) | NC score | 0.078003 (rank : 19) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 13) | NC score | 0.072179 (rank : 29) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 14) | NC score | 0.084488 (rank : 15) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
ZKSC1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.006125 (rank : 94) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P17029, P52745, Q8TBW5, Q8TEK7 | Gene names | ZKSCAN1, KOX18, ZNF139, ZNF36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1 (Zinc finger protein 36) (Zinc finger protein KOX18). | |||||
|
CCD71_HUMAN
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.092449 (rank : 9) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IV32, Q6IPE2, Q9H8H4, Q9H9F1 | Gene names | CCDC71 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 71. | |||||
|
NOL1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.071208 (rank : 31) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46087 | Gene names | NOL1 | |||
|
Domain Architecture |

|
|||||
| Description | Proliferating-cell nucleolar antigen p120 (Proliferation-associated nucleolar protein p120). | |||||
|
BARX1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.010659 (rank : 93) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HBU1 | Gene names | BARX1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein BarH-like 1. | |||||
|
BAZ2A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 19) | NC score | 0.040627 (rank : 80) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 20) | NC score | 0.059799 (rank : 50) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CT151_HUMAN
|
||||||
| θ value | 4.03905 (rank : 21) | NC score | 0.079936 (rank : 16) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NC74, Q8N4Z9, Q9BR75, Q9H0Y9 | Gene names | C20orf151 | |||
|
Domain Architecture |
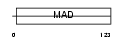
|
|||||
| Description | Uncharacterized protein C20orf151. | |||||
|
JPH2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.026192 (rank : 87) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.086312 (rank : 14) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
CU087_HUMAN
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.087270 (rank : 12) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P59051 | Gene names | C21orf87 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C21orf87. | |||||
|
MINK1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.011839 (rank : 92) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.055099 (rank : 59) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MTF2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.045813 (rank : 78) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
RECQ5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.031208 (rank : 84) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
RIMB1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.025142 (rank : 88) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1479 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O95153, O75111, Q8N5W3 | Gene names | BZRAP1, KIAA0612, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
CI079_HUMAN
|
||||||
| θ value | 6.88961 (rank : 30) | NC score | 0.048466 (rank : 77) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZUB1, Q5SQC9, Q8NA41, Q8ND27 | Gene names | C9orf79 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf79. | |||||
|
EMIL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.022449 (rank : 90) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BXX0, Q8NBH3, Q96JQ4 | Gene names | EMILIN2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Protein FOAP-10). | |||||
|
GIDRP_MOUSE
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.050108 (rank : 76) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BJM3, Q3UGJ8, Q5U5U8 | Gene names | Gidrp88, D19Ertd386e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth inhibition and differentiation-related protein 88 homolog. | |||||
|
MZF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 33) | NC score | 0.002539 (rank : 95) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28698, Q9NRY0, Q9UBW2 | Gene names | MZF1, MZF, ZNF42, ZSCAN6 | |||
|
Domain Architecture |
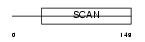
|
|||||
| Description | Myeloid zinc finger 1 (MZF-1) (Zinc finger protein 42) (Zinc finger and SCAN domain-containing protein 6). | |||||
|
NALP5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.024307 (rank : 89) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
NPM_HUMAN
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.035164 (rank : 83) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P06748, P08693, Q12826, Q13440, Q13441, Q14115, Q5EU94, Q5EU95, Q5EU96, Q5EU97, Q5EU98, Q5EU99, Q6V962, Q8WTW5, Q96AT6, Q96DC4, Q96EA5 | Gene names | NPM1, NPM | |||
|
Domain Architecture |

|
|||||
| Description | Nucleophosmin (NPM) (Nucleolar phosphoprotein B23) (Numatrin) (Nucleolar protein NO38). | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.074441 (rank : 23) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
VASH_MOUSE
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.026922 (rank : 86) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C1W1, Q8C394 | Gene names | Vash1, Vash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vasohibin. | |||||
|
CEND1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.016500 (rank : 91) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WZ64, Q4W5D2, Q7Z2L5, Q96L70, Q96P49, Q9Y4E4 | Gene names | CENTD1, ARAP2, KIAA0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 2) (PARX protein). | |||||
|
K1543_MOUSE
|
||||||
| θ value | 8.99809 (rank : 39) | NC score | 0.039907 (rank : 82) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80VC9, Q5DTW9, Q8BUZ0 | Gene names | Kiaa1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
MTF2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 40) | NC score | 0.041538 (rank : 79) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
Domain Architecture |
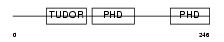
|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
AFF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.066138 (rank : 39) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
AFF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.051002 (rank : 73) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.053380 (rank : 66) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.057323 (rank : 52) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 45) | NC score | 0.076035 (rank : 20) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.072044 (rank : 30) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CIR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.054769 (rank : 62) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
CIR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.057074 (rank : 53) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.062084 (rank : 44) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.074159 (rank : 25) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.063782 (rank : 42) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
DSPP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.051568 (rank : 72) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
ENL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.079168 (rank : 17) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
GNAS3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.071059 (rank : 32) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95467, O95417 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroendocrine secretory protein 55 (NESP55) [Contains: LHAL tetrapeptide; GPIPIRRH peptide]. | |||||
|
GPTC8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.069255 (rank : 36) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
GRIN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.069615 (rank : 35) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
HIRP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.060668 (rank : 46) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
KI67_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.051808 (rank : 70) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.074391 (rank : 24) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MINT_MOUSE
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.054818 (rank : 61) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NEUM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.053588 (rank : 65) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P06837 | Gene names | Gap43, Basp2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuromodulin (Axonal membrane protein GAP-43) (Growth-associated protein 43) (Calmodulin-binding protein P-57). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.093426 (rank : 8) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.056199 (rank : 57) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.059927 (rank : 48) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
PAF49_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.059832 (rank : 49) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15446, Q32N11, Q7Z5U2, Q9UPF6 | Gene names | CD3EAP, ASE1, CAST, PAF49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase I-associated factor PAF49 (Anti-sense to ERCC-1 protein) (ASE-1) (CD3-epsilon-associated protein) (CD3E-associated protein) (CAST). | |||||
|
PININ_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.055386 (rank : 58) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
PKHA6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.057785 (rank : 51) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PPRB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.051694 (rank : 71) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
PPRB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.050366 (rank : 75) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.073846 (rank : 27) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.073907 (rank : 26) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
PRG4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.066896 (rank : 38) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRG4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.074443 (rank : 22) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.072385 (rank : 28) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
PRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.054683 (rank : 63) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRPC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.056991 (rank : 54) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P02810, Q4VBP2, Q53XA2, Q6P2F6 | Gene names | PRH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Salivary acidic proline-rich phosphoprotein 1/2 precursor (PRP-1/PRP- 2) (Parotid proline-rich protein 1/2) (Pr1/Pr2) (Protein C) (Parotid acidic protein) (Pa) (Parotid isoelectric focusing variant protein) (PIF-S) (Parotid double-band protein) (Db-s) [Contains: Salivary acidic proline-rich phosphoprotein 1/2; Salivary acidic proline-rich phosphoprotein 3/4 (PRP-3/PRP-4) (Protein A) (PIF-F) (Db-F); Peptide P-C]. | |||||
|
PRPE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.068159 (rank : 37) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P02811 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide P-E (IB-9). | |||||
|
SFR11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.056936 (rank : 55) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SFR12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.053876 (rank : 64) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
SFR17_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.056620 (rank : 56) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
SFRIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.054957 (rank : 60) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.100890 (rank : 7) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.089020 (rank : 11) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.090448 (rank : 10) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.078423 (rank : 18) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TCOF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.070937 (rank : 33) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
TCOF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.075519 (rank : 21) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
TGON1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.060391 (rank : 47) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
TGON2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.066133 (rank : 40) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
THOC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.062516 (rank : 43) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.060892 (rank : 45) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
VCX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.052548 (rank : 67) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H320, Q9P0H3 | Gene names | VCX, VCX1, VCX10R, VCXB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 1 (VCX-B1 protein) (Variably charged protein X-B1) (Variable charge protein on X with ten repeats) (VCX- 10r). | |||||
|
VCX3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.050630 (rank : 74) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NNX9, Q9P0H4 | Gene names | VCX3A, VCX3, VCX8R, VCXA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 3 (VCX-A protein) (Variably charged protein X-A) (Variable charge protein on X with eight repeats) (VCX- 8r). | |||||
|
VCXC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.051974 (rank : 69) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H321 | Gene names | VCXC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | VCX-C protein (Variably charged protein X-C). | |||||
|
ZC313_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.052480 (rank : 68) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
HMGA2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.86321e-21 (rank : 2) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P52926 | Gene names | HMGA2, HMGIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein HMGI-C (High mobility group AT-hook protein 2). | |||||
|
HMGA2_MOUSE
|
||||||
| NC score | 0.958572 (rank : 2) | θ value | 3.62785e-25 (rank : 1) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P52927, Q3UQW0 | Gene names | Hmga2, Hmgic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein HMGI-C (High mobility group AT-hook protein 2). | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.160380 (rank : 3) | θ value | 0.279714 (rank : 7) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.159601 (rank : 4) | θ value | 0.0563607 (rank : 5) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.140400 (rank : 5) | θ value | 0.163984 (rank : 6) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.129439 (rank : 6) | θ value | 0.0193708 (rank : 4) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.100890 (rank : 7) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.093426 (rank : 8) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
CCD71_HUMAN
|
||||||
| NC score | 0.092449 (rank : 9) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IV32, Q6IPE2, Q9H8H4, Q9H9F1 | Gene names | CCDC71 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 71. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.090448 (rank : 10) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.089020 (rank : 11) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
CU087_HUMAN
|
||||||
| NC score | 0.087270 (rank : 12) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P59051 | Gene names | C21orf87 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C21orf87. | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.087144 (rank : 13) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.086312 (rank : 14) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.084488 (rank : 15) | θ value | 1.81305 (rank : 14) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
CT151_HUMAN
|
||||||
| NC score | 0.079936 (rank : 16) | θ value | 4.03905 (rank : 21) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NC74, Q8N4Z9, Q9BR75, Q9H0Y9 | Gene names | C20orf151 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf151. | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.079168 (rank : 17) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.078423 (rank : 18) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.078003 (rank : 19) | θ value | 1.38821 (rank : 12) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.076035 (rank : 20) | θ value | θ > 10 (rank : 45) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
TCOF_MOUSE
|
||||||
| NC score | 0.075519 (rank : 21) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.074443 (rank : 22) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.074441 (rank : 23) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.074391 (rank : 24) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.074159 (rank : 25) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.073907 (rank : 26) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.073846 (rank : 27) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.072385 (rank : 28) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.072179 (rank : 29) | θ value | 1.81305 (rank : 13) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.072044 (rank : 30) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
NOL1_HUMAN
|
||||||
| NC score | 0.071208 (rank : 31) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46087 | Gene names | NOL1 | |||
|
Domain Architecture |

|
|||||
| Description | Proliferating-cell nucleolar antigen p120 (Proliferation-associated nucleolar protein p120). | |||||
|
GNAS3_HUMAN
|
||||||
| NC score | 0.071059 (rank : 32) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95467, O95417 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroendocrine secretory protein 55 (NESP55) [Contains: LHAL tetrapeptide; GPIPIRRH peptide]. | |||||
|
TCOF_HUMAN
|
||||||
| NC score | 0.070937 (rank : 33) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
PKHA6_HUMAN
|
||||||
| NC score | 0.069678 (rank : 34) | θ value | 0.47712 (rank : 8) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
GRIN1_MOUSE
|
||||||
| NC score | 0.069615 (rank : 35) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
GPTC8_HUMAN
|
||||||
| NC score | 0.069255 (rank : 36) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
PRPE_HUMAN
|
||||||
| NC score | 0.068159 (rank : 37) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P02811 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide P-E (IB-9). | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.066896 (rank : 38) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.066138 (rank : 39) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
TGON2_MOUSE
|
||||||
| NC score | 0.066133 (rank : 40) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
GON4L_HUMAN
|
||||||
| NC score | 0.065561 (rank : 41) | θ value | 1.38821 (rank : 11) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.063782 (rank : 42) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
THOC2_HUMAN
|
||||||
| NC score | 0.062516 (rank : 43) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.062084 (rank : 44) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.060892 (rank : 45) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
HIRP3_HUMAN
|
||||||
| NC score | 0.060668 (rank : 46) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
TGON1_MOUSE
|
||||||
| NC score | 0.060391 (rank : 47) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.059927 (rank : 48) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
PAF49_HUMAN
|
||||||
| NC score | 0.059832 (rank : 49) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15446, Q32N11, Q7Z5U2, Q9UPF6 | Gene names | CD3EAP, ASE1, CAST, PAF49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase I-associated factor PAF49 (Anti-sense to ERCC-1 protein) (ASE-1) (CD3-epsilon-associated protein) (CD3E-associated protein) (CAST). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.059799 (rank : 50) | θ value | 4.03905 (rank : 20) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
PKHA6_MOUSE
|
||||||
| NC score | 0.057785 (rank : 51) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.057323 (rank : 52) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
CIR_MOUSE
|
||||||
| NC score | 0.057074 (rank : 53) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
PRPC_HUMAN
|
||||||
| NC score | 0.056991 (rank : 54) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P02810, Q4VBP2, Q53XA2, Q6P2F6 | Gene names | PRH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Salivary acidic proline-rich phosphoprotein 1/2 precursor (PRP-1/PRP- 2) (Parotid proline-rich protein 1/2) (Pr1/Pr2) (Protein C) (Parotid acidic protein) (Pa) (Parotid isoelectric focusing variant protein) (PIF-S) (Parotid double-band protein) (Db-s) [Contains: Salivary acidic proline-rich phosphoprotein 1/2; Salivary acidic proline-rich phosphoprotein 3/4 (PRP-3/PRP-4) (Protein A) (PIF-F) (Db-F); Peptide P-C]. | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.056936 (rank : 55) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.056620 (rank : 56) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.056199 (rank : 57) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
PININ_MOUSE
|
||||||
| NC score | 0.055386 (rank : 58) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.055099 (rank : 59) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SFRIP_HUMAN
|
||||||
| NC score | 0.054957 (rank : 60) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.054818 (rank : 61) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
CIR_HUMAN
|
||||||
| NC score | 0.054769 (rank : 62) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.054683 (rank : 63) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.053876 (rank : 64) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
NEUM_MOUSE
|
||||||
| NC score | 0.053588 (rank : 65) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P06837 | Gene names | Gap43, Basp2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuromodulin (Axonal membrane protein GAP-43) (Growth-associated protein 43) (Calmodulin-binding protein P-57). | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.053380 (rank : 66) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
VCX1_HUMAN
|
||||||
| NC score | 0.052548 (rank : 67) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H320, Q9P0H3 | Gene names | VCX, VCX1, VCX10R, VCXB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 1 (VCX-B1 protein) (Variably charged protein X-B1) (Variable charge protein on X with ten repeats) (VCX- 10r). | |||||
|
ZC313_HUMAN
|
||||||
| NC score | 0.052480 (rank : 68) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
VCXC_HUMAN
|
||||||
| NC score | 0.051974 (rank : 69) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H321 | Gene names | VCXC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | VCX-C protein (Variably charged protein X-C). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.051808 (rank : 70) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
PPRB_HUMAN
|
||||||
| NC score | 0.051694 (rank : 71) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.051568 (rank : 72) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
AFF1_MOUSE
|
||||||
| NC score | 0.051002 (rank : 73) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
VCX3_HUMAN
|
||||||
| NC score | 0.050630 (rank : 74) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NNX9, Q9P0H4 | Gene names | VCX3A, VCX3, VCX8R, VCXA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 3 (VCX-A protein) (Variably charged protein X-A) (Variable charge protein on X with eight repeats) (VCX- 8r). | |||||
|
PPRB_MOUSE
|
||||||
| NC score | 0.050366 (rank : 75) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
GIDRP_MOUSE
|
||||||
| NC score | 0.050108 (rank : 76) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BJM3, Q3UGJ8, Q5U5U8 | Gene names | Gidrp88, D19Ertd386e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth inhibition and differentiation-related protein 88 homolog. | |||||
|
CI079_HUMAN
|
||||||
| NC score | 0.048466 (rank : 77) | θ value | 6.88961 (rank : 30) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZUB1, Q5SQC9, Q8NA41, Q8ND27 | Gene names | C9orf79 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf79. | |||||
|
MTF2_MOUSE
|
||||||
| NC score | 0.045813 (rank : 78) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
MTF2_HUMAN
|
||||||
| NC score | 0.041538 (rank : 79) | θ value | 8.99809 (rank : 40) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
BAZ2A_HUMAN
|
||||||
| NC score | 0.040627 (rank : 80) | θ value | 4.03905 (rank : 19) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | 0.040375 (rank : 81) | θ value | 0.0113563 (rank : 3) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
K1543_MOUSE
|
||||||
| NC score | 0.039907 (rank : 82) | θ value | 8.99809 (rank : 39) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80VC9, Q5DTW9, Q8BUZ0 | Gene names | Kiaa1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
NPM_HUMAN
|
||||||
| NC score | 0.035164 (rank : 83) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P06748, P08693, Q12826, Q13440, Q13441, Q14115, Q5EU94, Q5EU95, Q5EU96, Q5EU97, Q5EU98, Q5EU99, Q6V962, Q8WTW5, Q96AT6, Q96DC4, Q96EA5 | Gene names | NPM1, NPM | |||
|
Domain Architecture |

|
|||||
| Description | Nucleophosmin (NPM) (Nucleolar phosphoprotein B23) (Numatrin) (Nucleolar protein NO38). | |||||
|
RECQ5_HUMAN
|
||||||
| NC score | 0.031208 (rank : 84) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
INVS_MOUSE
|
||||||
| NC score | 0.030957 (rank : 85) | θ value | 1.06291 (rank : 10) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
VASH_MOUSE
|
||||||
| NC score | 0.026922 (rank : 86) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C1W1, Q8C394 | Gene names | Vash1, Vash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vasohibin. | |||||
|
JPH2_HUMAN
|
||||||
| NC score | 0.026192 (rank : 87) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
RIMB1_HUMAN
|
||||||
| NC score | 0.025142 (rank : 88) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1479 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O95153, O75111, Q8N5W3 | Gene names | BZRAP1, KIAA0612, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
NALP5_MOUSE
|
||||||
| NC score | 0.024307 (rank : 89) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
EMIL2_HUMAN
|
||||||
| NC score | 0.022449 (rank : 90) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BXX0, Q8NBH3, Q96JQ4 | Gene names | EMILIN2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Protein FOAP-10). | |||||
|
CEND1_HUMAN
|
||||||
| NC score | 0.016500 (rank : 91) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WZ64, Q4W5D2, Q7Z2L5, Q96L70, Q96P49, Q9Y4E4 | Gene names | CENTD1, ARAP2, KIAA0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 2) (PARX protein). | |||||
|
MINK1_HUMAN
|
||||||
| NC score | 0.011839 (rank : 92) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
BARX1_HUMAN
|
||||||
| NC score | 0.010659 (rank : 93) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HBU1 | Gene names | BARX1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein BarH-like 1. | |||||
|
ZKSC1_HUMAN
|
||||||
| NC score | 0.006125 (rank : 94) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P17029, P52745, Q8TBW5, Q8TEK7 | Gene names | ZKSCAN1, KOX18, ZNF139, ZNF36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1 (Zinc finger protein 36) (Zinc finger protein KOX18). | |||||
|
MZF1_HUMAN
|
||||||
| NC score | 0.002539 (rank : 95) | θ value | 6.88961 (rank : 33) | |||
| Query Neighborhood Hits | 40 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28698, Q9NRY0, Q9UBW2 | Gene names | MZF1, MZF, ZNF42, ZSCAN6 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid zinc finger 1 (MZF-1) (Zinc finger protein 42) (Zinc finger and SCAN domain-containing protein 6). | |||||