Please be patient as the page loads
|
ERCC3_MOUSE
|
||||||
| SwissProt Accessions | P49135 | Gene names | Ercc3, Xpb, Xpbc | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase XPB subunit (EC 3.6.1.-) (Basic transcription factor 2 89 kDa subunit) (BTF2-p89) (TFIIH 89 kDa subunit) (DNA-repair protein complementing XP-B cells) (Xeroderma pigmentosum group B-complementing protein) (DNA excision repair protein ERCC-3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ERCC3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.995736 (rank : 2) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P19447 | Gene names | ERCC3, XPB, XPBC | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase XPB subunit (EC 3.6.1.-) (Basic transcription factor 2 89 kDa subunit) (BTF2-p89) (TFIIH 89 kDa subunit) (DNA-repair protein complementing XP-B cells) (Xeroderma pigmentosum group B-complementing protein) (DNA excision repair protein ERCC-3). | |||||
|
ERCC3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P49135 | Gene names | Ercc3, Xpb, Xpbc | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase XPB subunit (EC 3.6.1.-) (Basic transcription factor 2 89 kDa subunit) (BTF2-p89) (TFIIH 89 kDa subunit) (DNA-repair protein complementing XP-B cells) (Xeroderma pigmentosum group B-complementing protein) (DNA excision repair protein ERCC-3). | |||||
|
SMRCD_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 3) | NC score | 0.053062 (rank : 3) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
LAMA2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 4) | NC score | 0.023183 (rank : 9) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
RA54B_MOUSE
|
||||||
| θ value | 0.279714 (rank : 5) | NC score | 0.046623 (rank : 5) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PFE3 | Gene names | Rad54b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54B (EC 3.6.1.-) (RAD54 homolog B). | |||||
|
LAMA2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 6) | NC score | 0.020914 (rank : 11) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
BIG1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 7) | NC score | 0.021835 (rank : 10) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6D6, Q9NV46, Q9UFV2, Q9UNL0 | Gene names | ARFGEF1, ARFGEP1, BIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 1 (Brefeldin A-inhibited GEP 1) (p200 ARF-GEP1) (p200 ARF guanine nucleotide exchange factor). | |||||
|
IFIH1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 8) | NC score | 0.037953 (rank : 7) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BYX4, Q6DC96, Q86X56, Q96MX8, Q9H3G6 | Gene names | IFIH1, MDA5, RH116 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-induced helicase C domain-containing protein 1 (EC 3.6.1.-) (Interferon-induced with helicase C domain protein 1) (Helicase with 2 CARD domains) (Helicard) (Melanoma differentiation-associated protein 5) (MDA-5) (RNA helicase-DEAD box protein 116) (Murabutide-down- regulated protein). | |||||
|
TLE2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 9) | NC score | 0.011589 (rank : 17) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q04725, Q8WVY0, Q9Y6S0 | Gene names | TLE2 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 2 (ESG2). | |||||
|
DDX5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 10) | NC score | 0.017489 (rank : 14) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17844, O75681 | Gene names | DDX5, HELR, HLR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX5 (EC 3.6.1.-) (DEAD box protein 5) (RNA helicase p68). | |||||
|
DDX5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 11) | NC score | 0.017512 (rank : 13) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61656 | Gene names | Ddx5, Tnz2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX5 (EC 3.6.1.-) (DEAD box protein 5) (RNA helicase p68) (DEAD box RNA helicase DEAD1) (mDEAD1). | |||||
|
SHAN2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 12) | NC score | 0.016927 (rank : 15) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPX8, Q3Y8G9, Q52LK2, Q9UKP1 | Gene names | SHANK2, KIAA1022 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2). | |||||
|
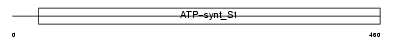
VAS1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 13) | NC score | 0.048804 (rank : 4) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9R1Q9 | Gene names | Atp6ap1, Atp6ip1, Atp6s1 | |||
|
Domain Architecture |
|
|||||
| Description | Vacuolar ATP synthase subunit S1 precursor (EC 3.6.3.14) (V-ATPase S1 subunit) (V-ATPase S1 accessory protein) (V-ATPase Ac45 subunit) (C7-1 protein). | |||||
|
CHD6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 14) | NC score | 0.026246 (rank : 8) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TD26, Q5JYQ0, Q5TGZ9, Q5TH00, Q5TH01, Q8IZR2, Q8WTY0, Q9H4H6, Q9H6D4, Q9NTT7, Q9P2L1 | Gene names | CHD6, CHD5, KIAA1335, RIGB | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 6 (EC 3.6.1.-) (ATP- dependent helicase CHD6) (CHD-6) (Radiation-induced gene B protein). | |||||
|
MYO1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 15) | NC score | 0.004650 (rank : 23) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBC5, Q9UQD7 | Gene names | MYO1A, MYHL | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ia (Brush border myosin I) (BBM-I) (BBMI) (Myosin I heavy chain) (MIHC). | |||||
|
NALDL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 16) | NC score | 0.010646 (rank : 18) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7M758 | Gene names | Naaladl1, Naaladasel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L). | |||||
|
VAS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 17) | NC score | 0.041358 (rank : 6) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15904 | Gene names | ATP6AP1, ATP6IP1, ATP6S1, VATPS1, XAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar ATP synthase subunit S1 precursor (EC 3.6.3.14) (V-ATPase S1 subunit) (V-ATPase S1 accessory protein) (V-ATPase Ac45 subunit) (XAP- 3). | |||||
|
CT077_HUMAN
|
||||||
| θ value | 8.99809 (rank : 18) | NC score | 0.013652 (rank : 16) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQG5 | Gene names | C20orf77 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf77. | |||||
|
IFIH1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.020897 (rank : 12) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8R5F7, Q3U6S2, Q68EM4, Q8BYC9, Q8BZ01, Q8K5C7, Q8R144, Q8VE79, Q99KS4, Q9D2Z5 | Gene names | Ifih1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-induced helicase C domain-containing protein 1 (EC 3.6.1.-) (Interferon induced with helicase C domain protein 1) (Helicase with 2 CARD domains) (Helicard) (Melanoma differentiation-associated protein 5) (MDA-5). | |||||
|
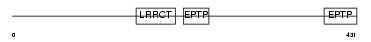
LGI4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 20) | NC score | 0.005265 (rank : 22) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K1S1, Q8K4Y8 | Gene names | Lgi4, Lgil3 | |||
|
Domain Architecture |
|
|||||
| Description | Leucine-rich repeat LGI family member 4 precursor (Leucine-rich glioma-inactivated protein 4) (LGI1-like protein 3). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 21) | NC score | 0.007174 (rank : 21) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
SIA7D_HUMAN
|
||||||
| θ value | 8.99809 (rank : 22) | NC score | 0.009195 (rank : 19) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H4F1, Q9NWU6, Q9UKU1, Q9ULB9, Q9Y3G3, Q9Y3G4 | Gene names | ST6GALNAC4, SIAT3C, SIAT7D | |||
|
Domain Architecture |
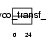
|
|||||
| Description | Alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3-N-acetyl- galactosaminide alpha-2,6-sialyltransferase (EC 2.4.99.7) (NeuAc- alpha-2,3-Gal-beta-1,3-GalNAc-alpha-2,6-sialyltransferase) (ST6GalNAc IV) (Sialyltransferase 7D) (Sialyltransferase 3C). | |||||
|
TRIPB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 23) | NC score | 0.007339 (rank : 20) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
ERCC3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P49135 | Gene names | Ercc3, Xpb, Xpbc | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase XPB subunit (EC 3.6.1.-) (Basic transcription factor 2 89 kDa subunit) (BTF2-p89) (TFIIH 89 kDa subunit) (DNA-repair protein complementing XP-B cells) (Xeroderma pigmentosum group B-complementing protein) (DNA excision repair protein ERCC-3). | |||||
|
ERCC3_HUMAN
|
||||||
| NC score | 0.995736 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P19447 | Gene names | ERCC3, XPB, XPBC | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex helicase XPB subunit (EC 3.6.1.-) (Basic transcription factor 2 89 kDa subunit) (BTF2-p89) (TFIIH 89 kDa subunit) (DNA-repair protein complementing XP-B cells) (Xeroderma pigmentosum group B-complementing protein) (DNA excision repair protein ERCC-3). | |||||
|
SMRCD_MOUSE
|
||||||
| NC score | 0.053062 (rank : 3) | θ value | 0.0431538 (rank : 3) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
VAS1_MOUSE
|
||||||
| NC score | 0.048804 (rank : 4) | θ value | 4.03905 (rank : 13) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9R1Q9 | Gene names | Atp6ap1, Atp6ip1, Atp6s1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar ATP synthase subunit S1 precursor (EC 3.6.3.14) (V-ATPase S1 subunit) (V-ATPase S1 accessory protein) (V-ATPase Ac45 subunit) (C7-1 protein). | |||||
|
RA54B_MOUSE
|
||||||
| NC score | 0.046623 (rank : 5) | θ value | 0.279714 (rank : 5) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PFE3 | Gene names | Rad54b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54B (EC 3.6.1.-) (RAD54 homolog B). | |||||
|
VAS1_HUMAN
|
||||||
| NC score | 0.041358 (rank : 6) | θ value | 6.88961 (rank : 17) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15904 | Gene names | ATP6AP1, ATP6IP1, ATP6S1, VATPS1, XAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar ATP synthase subunit S1 precursor (EC 3.6.3.14) (V-ATPase S1 subunit) (V-ATPase S1 accessory protein) (V-ATPase Ac45 subunit) (XAP- 3). | |||||
|
IFIH1_HUMAN
|
||||||
| NC score | 0.037953 (rank : 7) | θ value | 2.36792 (rank : 8) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BYX4, Q6DC96, Q86X56, Q96MX8, Q9H3G6 | Gene names | IFIH1, MDA5, RH116 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-induced helicase C domain-containing protein 1 (EC 3.6.1.-) (Interferon-induced with helicase C domain protein 1) (Helicase with 2 CARD domains) (Helicard) (Melanoma differentiation-associated protein 5) (MDA-5) (RNA helicase-DEAD box protein 116) (Murabutide-down- regulated protein). | |||||
|
CHD6_HUMAN
|
||||||
| NC score | 0.026246 (rank : 8) | θ value | 6.88961 (rank : 14) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TD26, Q5JYQ0, Q5TGZ9, Q5TH00, Q5TH01, Q8IZR2, Q8WTY0, Q9H4H6, Q9H6D4, Q9NTT7, Q9P2L1 | Gene names | CHD6, CHD5, KIAA1335, RIGB | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 6 (EC 3.6.1.-) (ATP- dependent helicase CHD6) (CHD-6) (Radiation-induced gene B protein). | |||||
|
LAMA2_HUMAN
|
||||||
| NC score | 0.023183 (rank : 9) | θ value | 0.21417 (rank : 4) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
BIG1_HUMAN
|
||||||
| NC score | 0.021835 (rank : 10) | θ value | 1.38821 (rank : 7) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6D6, Q9NV46, Q9UFV2, Q9UNL0 | Gene names | ARFGEF1, ARFGEP1, BIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 1 (Brefeldin A-inhibited GEP 1) (p200 ARF-GEP1) (p200 ARF guanine nucleotide exchange factor). | |||||
|
LAMA2_MOUSE
|
||||||
| NC score | 0.020914 (rank : 11) | θ value | 0.62314 (rank : 6) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
IFIH1_MOUSE
|
||||||
| NC score | 0.020897 (rank : 12) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8R5F7, Q3U6S2, Q68EM4, Q8BYC9, Q8BZ01, Q8K5C7, Q8R144, Q8VE79, Q99KS4, Q9D2Z5 | Gene names | Ifih1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-induced helicase C domain-containing protein 1 (EC 3.6.1.-) (Interferon induced with helicase C domain protein 1) (Helicase with 2 CARD domains) (Helicard) (Melanoma differentiation-associated protein 5) (MDA-5). | |||||
|
DDX5_MOUSE
|
||||||
| NC score | 0.017512 (rank : 13) | θ value | 3.0926 (rank : 11) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61656 | Gene names | Ddx5, Tnz2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX5 (EC 3.6.1.-) (DEAD box protein 5) (RNA helicase p68) (DEAD box RNA helicase DEAD1) (mDEAD1). | |||||
|
DDX5_HUMAN
|
||||||
| NC score | 0.017489 (rank : 14) | θ value | 3.0926 (rank : 10) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17844, O75681 | Gene names | DDX5, HELR, HLR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX5 (EC 3.6.1.-) (DEAD box protein 5) (RNA helicase p68). | |||||
|
SHAN2_HUMAN
|
||||||
| NC score | 0.016927 (rank : 15) | θ value | 4.03905 (rank : 12) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPX8, Q3Y8G9, Q52LK2, Q9UKP1 | Gene names | SHANK2, KIAA1022 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2). | |||||
|
CT077_HUMAN
|
||||||
| NC score | 0.013652 (rank : 16) | θ value | 8.99809 (rank : 18) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQG5 | Gene names | C20orf77 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf77. | |||||
|
TLE2_HUMAN
|
||||||
| NC score | 0.011589 (rank : 17) | θ value | 2.36792 (rank : 9) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q04725, Q8WVY0, Q9Y6S0 | Gene names | TLE2 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 2 (ESG2). | |||||
|
NALDL_MOUSE
|
||||||
| NC score | 0.010646 (rank : 18) | θ value | 6.88961 (rank : 16) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7M758 | Gene names | Naaladl1, Naaladasel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L). | |||||
|
SIA7D_HUMAN
|
||||||
| NC score | 0.009195 (rank : 19) | θ value | 8.99809 (rank : 22) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H4F1, Q9NWU6, Q9UKU1, Q9ULB9, Q9Y3G3, Q9Y3G4 | Gene names | ST6GALNAC4, SIAT3C, SIAT7D | |||
|
Domain Architecture |
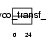
|
|||||
| Description | Alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3-N-acetyl- galactosaminide alpha-2,6-sialyltransferase (EC 2.4.99.7) (NeuAc- alpha-2,3-Gal-beta-1,3-GalNAc-alpha-2,6-sialyltransferase) (ST6GalNAc IV) (Sialyltransferase 7D) (Sialyltransferase 3C). | |||||
|
TRIPB_HUMAN
|
||||||
| NC score | 0.007339 (rank : 20) | θ value | 8.99809 (rank : 23) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.007174 (rank : 21) | θ value | 8.99809 (rank : 21) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
LGI4_MOUSE
|
||||||
| NC score | 0.005265 (rank : 22) | θ value | 8.99809 (rank : 20) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K1S1, Q8K4Y8 | Gene names | Lgi4, Lgil3 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat LGI family member 4 precursor (Leucine-rich glioma-inactivated protein 4) (LGI1-like protein 3). | |||||
|
MYO1A_HUMAN
|
||||||
| NC score | 0.004650 (rank : 23) | θ value | 6.88961 (rank : 15) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBC5, Q9UQD7 | Gene names | MYO1A, MYHL | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ia (Brush border myosin I) (BBM-I) (BBMI) (Myosin I heavy chain) (MIHC). | |||||