Please be patient as the page loads
|
CRBB1_MOUSE
|
||||||
| SwissProt Accessions | Q9WVJ5 | Gene names | Crybb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta crystallin B1 [Contains: Beta crystallin B1B]. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CRBB1_MOUSE
|
||||||
| θ value | 2.52853e-119 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WVJ5 | Gene names | Crybb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta crystallin B1 [Contains: Beta crystallin B1B]. | |||||
|
CRBB1_HUMAN
|
||||||
| θ value | 4.95767e-99 (rank : 2) | NC score | 0.998578 (rank : 2) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53674 | Gene names | CRYBB1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B1. | |||||
|
CRBB3_MOUSE
|
||||||
| θ value | 7.98882e-65 (rank : 3) | NC score | 0.991236 (rank : 4) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJU9 | Gene names | Crybb3 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
CRBB3_HUMAN
|
||||||
| θ value | 2.84138e-62 (rank : 4) | NC score | 0.991270 (rank : 3) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P26998, Q92965, Q9UH09 | Gene names | CRYBB3, CRYB3 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
CRBB2_MOUSE
|
||||||
| θ value | 5.73848e-55 (rank : 5) | NC score | 0.987690 (rank : 5) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P62696, P19942, P26775 | Gene names | Crybb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta crystallin B2 (Beta-crystallin Bp). | |||||
|
CRBB2_HUMAN
|
||||||
| θ value | 3.71957e-54 (rank : 6) | NC score | 0.987601 (rank : 6) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P43320, Q9UCM8 | Gene names | CRYBB2, CRYB2, CRYB2A | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin B2 (Beta-crystallin Bp). | |||||
|
CRBA1_HUMAN
|
||||||
| θ value | 3.37202e-47 (rank : 7) | NC score | 0.974360 (rank : 7) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05813, Q13633 | Gene names | CRYBA1, CRYB1 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin A3 [Contains: Beta crystallin A3, isoform A1, Delta4 form; Beta crystallin A3, isoform A1, Delta7 form; Beta crystallin A3, isoform A1, Delta8 form]. | |||||
|
CRBA1_MOUSE
|
||||||
| θ value | 2.41655e-45 (rank : 8) | NC score | 0.972979 (rank : 8) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P02525 | Gene names | Cryba1 | |||
|
Domain Architecture |
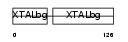
|
|||||
| Description | Beta crystallin A1 (Beta-A1-crystallin). | |||||
|
CRBA4_HUMAN
|
||||||
| θ value | 6.15952e-41 (rank : 9) | NC score | 0.967145 (rank : 9) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53673, Q4VB22, Q6ICE4 | Gene names | CRYBA4 | |||
|
Domain Architecture |
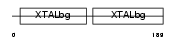
|
|||||
| Description | Beta crystallin A4 (Beta-A4-crystallin). | |||||
|
CRBA4_MOUSE
|
||||||
| θ value | 6.15952e-41 (rank : 10) | NC score | 0.966357 (rank : 10) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJV0 | Gene names | Cryba4 | |||
|
Domain Architecture |
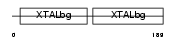
|
|||||
| Description | Beta crystallin A4 (Beta-A4-crystallin). | |||||
|
CRBA2_MOUSE
|
||||||
| θ value | 3.27365e-34 (rank : 11) | NC score | 0.961546 (rank : 11) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJV1 | Gene names | Cryba2 | |||
|
Domain Architecture |
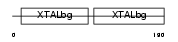
|
|||||
| Description | Beta crystallin A2 (Beta-A2-crystallin). | |||||
|
CRBA2_HUMAN
|
||||||
| θ value | 9.8567e-31 (rank : 12) | NC score | 0.958732 (rank : 12) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53672, Q9Y562 | Gene names | CRYBA2 | |||
|
Domain Architecture |
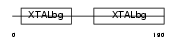
|
|||||
| Description | Beta crystallin A2 (Beta-A2-crystallin). | |||||
|
CRGB_MOUSE
|
||||||
| θ value | 3.62785e-25 (rank : 13) | NC score | 0.894836 (rank : 15) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04344, Q61593 | Gene names | Crygb | |||
|
Domain Architecture |
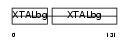
|
|||||
| Description | Gamma crystallin B (Gamma crystallin 3). | |||||
|
CRBS_HUMAN
|
||||||
| θ value | 2.35151e-24 (rank : 14) | NC score | 0.911603 (rank : 13) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P22914 | Gene names | CRYGS, GRYG8 | |||
|
Domain Architecture |
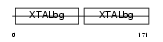
|
|||||
| Description | Beta crystallin S (Gamma crystallin S). | |||||
|
CRBS_MOUSE
|
||||||
| θ value | 2.35151e-24 (rank : 15) | NC score | 0.910515 (rank : 14) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35486 | Gene names | Crygs | |||
|
Domain Architecture |
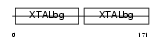
|
|||||
| Description | Beta crystallin S (Gamma crystallin S). | |||||
|
CRGA_MOUSE
|
||||||
| θ value | 6.84181e-24 (rank : 16) | NC score | 0.890704 (rank : 17) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04345 | Gene names | Cryga | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin A (Gamma crystallin 4). | |||||
|
CRGC_HUMAN
|
||||||
| θ value | 2.59989e-23 (rank : 17) | NC score | 0.892709 (rank : 16) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07315 | Gene names | CRYGC, CRYG3 | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin C (Gamma crystallin 2-1) (Gamma crystallin 3). | |||||
|
CRGC_MOUSE
|
||||||
| θ value | 1.6852e-22 (rank : 18) | NC score | 0.885025 (rank : 20) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61597, Q03739 | Gene names | Crygc, Gammab-cry | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin C. | |||||
|
CRGD_MOUSE
|
||||||
| θ value | 1.6852e-22 (rank : 19) | NC score | 0.881921 (rank : 21) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04342, O89027 | Gene names | Crygd | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin D (Gamma crystallin 1). | |||||
|
CRGE_MOUSE
|
||||||
| θ value | 1.6852e-22 (rank : 20) | NC score | 0.881687 (rank : 22) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q03740, O89028, P26999, Q9CXK5 | Gene names | Cryge | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin E. | |||||
|
CRGF_MOUSE
|
||||||
| θ value | 3.75424e-22 (rank : 21) | NC score | 0.880252 (rank : 23) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9CXV3 | Gene names | Crygf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma crystallin F. | |||||
|
CRGB_HUMAN
|
||||||
| θ value | 3.17815e-21 (rank : 22) | NC score | 0.886926 (rank : 18) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07316 | Gene names | CRYGB, CRYG2 | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin B (Gamma crystallin 1-2). | |||||
|
CRGA_HUMAN
|
||||||
| θ value | 9.24701e-21 (rank : 23) | NC score | 0.885271 (rank : 19) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P11844 | Gene names | CRYGA, CRYG1 | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin A (Gamma crystallin 5). | |||||
|
CRGD_HUMAN
|
||||||
| θ value | 5.99374e-20 (rank : 24) | NC score | 0.877074 (rank : 24) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07320, Q53R51, Q99681 | Gene names | CRYGD, CRYG4 | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin D (Gamma crystallin 4). | |||||
|
AIM1_HUMAN
|
||||||
| θ value | 2.13179e-17 (rank : 25) | NC score | 0.798604 (rank : 25) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y4K1, O00296, Q5VWJ2 | Gene names | AIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Absent in melanoma 1 protein. | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 3.0926 (rank : 26) | NC score | 0.006939 (rank : 28) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.010136 (rank : 27) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
LITAF_HUMAN
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.015156 (rank : 26) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99732, Q9C0L6 | Gene names | LITAF, PIG7, SIMPLE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipopolysaccharide-induced tumor necrosis factor-alpha factor (LPS- induced TNF-alpha factor) (p53-induced protein 7) (Small integral membrane protein of lysosome/late endosome). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.001240 (rank : 29) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
CRBB1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 2.52853e-119 (rank : 1) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WVJ5 | Gene names | Crybb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta crystallin B1 [Contains: Beta crystallin B1B]. | |||||
|
CRBB1_HUMAN
|
||||||
| NC score | 0.998578 (rank : 2) | θ value | 4.95767e-99 (rank : 2) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53674 | Gene names | CRYBB1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B1. | |||||
|
CRBB3_HUMAN
|
||||||
| NC score | 0.991270 (rank : 3) | θ value | 2.84138e-62 (rank : 4) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P26998, Q92965, Q9UH09 | Gene names | CRYBB3, CRYB3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
CRBB3_MOUSE
|
||||||
| NC score | 0.991236 (rank : 4) | θ value | 7.98882e-65 (rank : 3) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJU9 | Gene names | Crybb3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
CRBB2_MOUSE
|
||||||
| NC score | 0.987690 (rank : 5) | θ value | 5.73848e-55 (rank : 5) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P62696, P19942, P26775 | Gene names | Crybb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta crystallin B2 (Beta-crystallin Bp). | |||||
|
CRBB2_HUMAN
|
||||||
| NC score | 0.987601 (rank : 6) | θ value | 3.71957e-54 (rank : 6) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P43320, Q9UCM8 | Gene names | CRYBB2, CRYB2, CRYB2A | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B2 (Beta-crystallin Bp). | |||||
|
CRBA1_HUMAN
|
||||||
| NC score | 0.974360 (rank : 7) | θ value | 3.37202e-47 (rank : 7) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05813, Q13633 | Gene names | CRYBA1, CRYB1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A3 [Contains: Beta crystallin A3, isoform A1, Delta4 form; Beta crystallin A3, isoform A1, Delta7 form; Beta crystallin A3, isoform A1, Delta8 form]. | |||||
|
CRBA1_MOUSE
|
||||||
| NC score | 0.972979 (rank : 8) | θ value | 2.41655e-45 (rank : 8) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P02525 | Gene names | Cryba1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A1 (Beta-A1-crystallin). | |||||
|
CRBA4_HUMAN
|
||||||
| NC score | 0.967145 (rank : 9) | θ value | 6.15952e-41 (rank : 9) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53673, Q4VB22, Q6ICE4 | Gene names | CRYBA4 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A4 (Beta-A4-crystallin). | |||||
|
CRBA4_MOUSE
|
||||||
| NC score | 0.966357 (rank : 10) | θ value | 6.15952e-41 (rank : 10) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJV0 | Gene names | Cryba4 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A4 (Beta-A4-crystallin). | |||||
|
CRBA2_MOUSE
|
||||||
| NC score | 0.961546 (rank : 11) | θ value | 3.27365e-34 (rank : 11) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJV1 | Gene names | Cryba2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A2 (Beta-A2-crystallin). | |||||
|
CRBA2_HUMAN
|
||||||
| NC score | 0.958732 (rank : 12) | θ value | 9.8567e-31 (rank : 12) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53672, Q9Y562 | Gene names | CRYBA2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A2 (Beta-A2-crystallin). | |||||
|
CRBS_HUMAN
|
||||||
| NC score | 0.911603 (rank : 13) | θ value | 2.35151e-24 (rank : 14) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P22914 | Gene names | CRYGS, GRYG8 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin S (Gamma crystallin S). | |||||
|
CRBS_MOUSE
|
||||||
| NC score | 0.910515 (rank : 14) | θ value | 2.35151e-24 (rank : 15) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35486 | Gene names | Crygs | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin S (Gamma crystallin S). | |||||
|
CRGB_MOUSE
|
||||||
| NC score | 0.894836 (rank : 15) | θ value | 3.62785e-25 (rank : 13) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04344, Q61593 | Gene names | Crygb | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin B (Gamma crystallin 3). | |||||
|
CRGC_HUMAN
|
||||||
| NC score | 0.892709 (rank : 16) | θ value | 2.59989e-23 (rank : 17) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07315 | Gene names | CRYGC, CRYG3 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin C (Gamma crystallin 2-1) (Gamma crystallin 3). | |||||
|
CRGA_MOUSE
|
||||||
| NC score | 0.890704 (rank : 17) | θ value | 6.84181e-24 (rank : 16) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04345 | Gene names | Cryga | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin A (Gamma crystallin 4). | |||||
|
CRGB_HUMAN
|
||||||
| NC score | 0.886926 (rank : 18) | θ value | 3.17815e-21 (rank : 22) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07316 | Gene names | CRYGB, CRYG2 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin B (Gamma crystallin 1-2). | |||||
|
CRGA_HUMAN
|
||||||
| NC score | 0.885271 (rank : 19) | θ value | 9.24701e-21 (rank : 23) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P11844 | Gene names | CRYGA, CRYG1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin A (Gamma crystallin 5). | |||||
|
CRGC_MOUSE
|
||||||
| NC score | 0.885025 (rank : 20) | θ value | 1.6852e-22 (rank : 18) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61597, Q03739 | Gene names | Crygc, Gammab-cry | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin C. | |||||
|
CRGD_MOUSE
|
||||||
| NC score | 0.881921 (rank : 21) | θ value | 1.6852e-22 (rank : 19) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04342, O89027 | Gene names | Crygd | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin D (Gamma crystallin 1). | |||||
|
CRGE_MOUSE
|
||||||
| NC score | 0.881687 (rank : 22) | θ value | 1.6852e-22 (rank : 20) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q03740, O89028, P26999, Q9CXK5 | Gene names | Cryge | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin E. | |||||
|
CRGF_MOUSE
|
||||||
| NC score | 0.880252 (rank : 23) | θ value | 3.75424e-22 (rank : 21) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9CXV3 | Gene names | Crygf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma crystallin F. | |||||
|
CRGD_HUMAN
|
||||||
| NC score | 0.877074 (rank : 24) | θ value | 5.99374e-20 (rank : 24) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07320, Q53R51, Q99681 | Gene names | CRYGD, CRYG4 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin D (Gamma crystallin 4). | |||||
|
AIM1_HUMAN
|
||||||
| NC score | 0.798604 (rank : 25) | θ value | 2.13179e-17 (rank : 25) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y4K1, O00296, Q5VWJ2 | Gene names | AIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Absent in melanoma 1 protein. | |||||
|
LITAF_HUMAN
|
||||||
| NC score | 0.015156 (rank : 26) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99732, Q9C0L6 | Gene names | LITAF, PIG7, SIMPLE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipopolysaccharide-induced tumor necrosis factor-alpha factor (LPS- induced TNF-alpha factor) (p53-induced protein 7) (Small integral membrane protein of lysosome/late endosome). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.010136 (rank : 27) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.006939 (rank : 28) | θ value | 3.0926 (rank : 26) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.001240 (rank : 29) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||