Please be patient as the page loads
|
CLIC5_HUMAN
|
||||||
| SwissProt Accessions | Q9NZA1, Q5T4Z0, Q96JT5, Q9BWZ0 | Gene names | CLIC5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 5. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CLIC5_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NZA1, Q5T4Z0, Q96JT5, Q9BWZ0 | Gene names | CLIC5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 5. | |||||
|
CLIC5_MOUSE
|
||||||
| θ value | 1.16911e-132 (rank : 2) | NC score | 0.996377 (rank : 2) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BXK9, Q3U0H8 | Gene names | Clic5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 5. | |||||
|
CLIC4_HUMAN
|
||||||
| θ value | 7.63087e-108 (rank : 3) | NC score | 0.992414 (rank : 4) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y696, Q9UFW9, Q9UQJ6 | Gene names | CLIC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 4 (Intracellular chloride ion channel protein p64H1). | |||||
|
CLIC4_MOUSE
|
||||||
| θ value | 2.89973e-107 (rank : 4) | NC score | 0.992503 (rank : 3) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QYB1, Q8BMG5 | Gene names | Clic4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 4 (mc3s5/mtCLIC). | |||||
|
CLIC6_MOUSE
|
||||||
| θ value | 2.15048e-102 (rank : 5) | NC score | 0.953452 (rank : 10) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BHB9 | Gene names | Clic6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
CLIC6_HUMAN
|
||||||
| θ value | 2.01279e-100 (rank : 6) | NC score | 0.867294 (rank : 11) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
CLIC2_HUMAN
|
||||||
| θ value | 1.88829e-90 (rank : 7) | NC score | 0.990631 (rank : 5) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15247, O15174, Q5JT80, Q8TCE3 | Gene names | CLIC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 2 (XAP121). | |||||
|
CLIC1_HUMAN
|
||||||
| θ value | 3.44927e-84 (rank : 8) | NC score | 0.986513 (rank : 6) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O00299, Q15089 | Gene names | CLIC1, NCC27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 1 (Nuclear chloride ion channel 27) (NCC27) (Chloride channel ABP) (Regulatory nuclear chloride ion channel protein) (hRNCC). | |||||
|
CLIC1_MOUSE
|
||||||
| θ value | 2.23575e-83 (rank : 9) | NC score | 0.986354 (rank : 7) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Z1Q5 | Gene names | Clic1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 1 (Nuclear chloride ion channel 27) (NCC27). | |||||
|
CLIC3_HUMAN
|
||||||
| θ value | 5.92465e-60 (rank : 10) | NC score | 0.971859 (rank : 8) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95833 | Gene names | CLIC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 3. | |||||
|
CLIC3_MOUSE
|
||||||
| θ value | 1.45929e-58 (rank : 11) | NC score | 0.969939 (rank : 9) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D7P7 | Gene names | Clic3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 3. | |||||
|
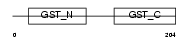
GSTO1_MOUSE
|
||||||
| θ value | 5.44631e-05 (rank : 12) | NC score | 0.315660 (rank : 12) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O09131, Q3TH87 | Gene names | Gsto1, Gstx, Gtsttl | |||
|
Domain Architecture |
|
|||||
| Description | Glutathione transferase omega-1 (EC 2.5.1.18) (GSTO 1-1) (p28). | |||||
|
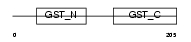
GSTO1_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 13) | NC score | 0.208673 (rank : 13) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P78417, Q5TA03, Q7Z3T2 | Gene names | GSTO1, GSTTLP28 | |||
|
Domain Architecture |
|
|||||
| Description | Glutathione transferase omega-1 (EC 2.5.1.18) (GSTO 1-1). | |||||
|
ZN148_HUMAN
|
||||||
| θ value | 0.125558 (rank : 14) | NC score | 0.005705 (rank : 32) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UQR1, O00389, O43591 | Gene names | ZNF148, ZBP89 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 148 (Zinc finger DNA-binding protein 89) (Transcription factor ZBP-89). | |||||
|
GDAP1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.085928 (rank : 18) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8TB36 | Gene names | GDAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ganglioside-induced differentiation-associated protein 1 (GDAP1). | |||||
|
GDAP1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.085969 (rank : 17) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O88741, Q8C7Q5, Q9CTN2 | Gene names | Gdap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ganglioside-induced differentiation-associated protein 1 (GDAP1). | |||||
|
ZN148_MOUSE
|
||||||
| θ value | 0.62314 (rank : 17) | NC score | 0.004860 (rank : 33) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61624, P97475 | Gene names | Znf148, Zbp89, Zfp148 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 148 (Zinc finger DNA-binding protein 89) (Transcription factor ZBP-89) (G-rich box-binding protein) (Beta enolase repressor factor 1) (Transcription factor BFCOL1). | |||||
|
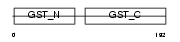
MAAI_MOUSE
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.102811 (rank : 14) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WVL0 | Gene names | Gstz1, Maai | |||
|
Domain Architecture |
|
|||||
| Description | Maleylacetoacetate isomerase (EC 5.2.1.2) (MAAI) (Glutathione S- transferase zeta 1) (EC 2.5.1.18) (GSTZ1-1). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 1.06291 (rank : 19) | NC score | 0.010786 (rank : 28) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
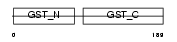
MAAI_HUMAN
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.097868 (rank : 16) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O43708, O15308, O75430, Q7Z610, Q9BV63 | Gene names | GSTZ1, MAAI | |||
|
Domain Architecture |
|
|||||
| Description | Maleylacetoacetate isomerase (EC 5.2.1.2) (MAAI) (Glutathione S- transferase zeta 1) (EC 2.5.1.18) (GSTZ1-1). | |||||
|
TM7S3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.019246 (rank : 22) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NS93, Q9NUS4 | Gene names | TM7SF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane 7 superfamily member 3 precursor (Seven span transmembrane protein). | |||||
|
EVPL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.003221 (rank : 34) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
RFIP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.010901 (rank : 27) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D620, Q3UBC2, Q8BN24 | Gene names | Rab11fip1, Rcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
CCD1A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.015428 (rank : 23) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P1N0, Q7Z435, Q86XV0, Q8NF89, Q9H603, Q9NXI1 | Gene names | CC2D1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1) (Putative NF-kappa-B- activating protein 023N). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.020021 (rank : 21) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
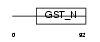
GSTO2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.069143 (rank : 19) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H4Y5, Q49TW5, Q5GM70, Q86WP3 | Gene names | GSTO2 | |||
|
Domain Architecture |
|
|||||
| Description | Glutathione transferase omega-2 (EC 2.5.1.18) (GSTO-2). | |||||
|
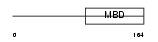
MECP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.012019 (rank : 26) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |
|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
MUG1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.006497 (rank : 31) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28665 | Gene names | Mug1, Mug-1 | |||
|
Domain Architecture |

|
|||||
| Description | Murinoglobulin-1 precursor (MuG1). | |||||
|
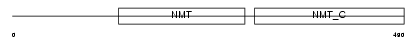
NMT1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.013658 (rank : 24) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70310 | Gene names | Nmt1 | |||
|
Domain Architecture |
|
|||||
| Description | Glycylpeptide N-tetradecanoyltransferase 1 (EC 2.3.1.97) (Peptide N- myristoyltransferase 1) (Myristoyl-CoA:protein N-myristoyltransferase 1) (NMT 1) (Type I N-myristoyltransferase). | |||||
|
ELL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 30) | NC score | 0.009234 (rank : 29) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00472 | Gene names | ELL2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL2. | |||||
|
GSTO2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.100718 (rank : 15) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K2Q2 | Gene names | Gsto2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutathione transferase omega-2 (EC 2.5.1.18). | |||||
|
NMT1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.012899 (rank : 25) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30419, Q9UE09 | Gene names | NMT1, NMT | |||
|
Domain Architecture |

|
|||||
| Description | Glycylpeptide N-tetradecanoyltransferase 1 (EC 2.3.1.97) (Peptide N- myristoyltransferase 1) (Myristoyl-CoA:protein N-myristoyltransferase 1) (NMT 1) (Type I N-myristoyltransferase). | |||||
|
PSN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.006907 (rank : 30) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49768, O95465, Q14762, Q15719, Q15720, Q96P33, Q9UIF0 | Gene names | PSEN1, AD3, PS1, PSNL1 | |||
|
Domain Architecture |

|
|||||
| Description | Presenilin-1 (EC 3.4.23.-) (PS-1) (Protein S182) [Contains: Presenilin-1 NTF subunit; Presenilin-1 CTF subunit; Presenilin-1 CTF12 (PS1-CTF12)]. | |||||
|
ASPX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 34) | NC score | 0.055025 (rank : 20) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26436 | Gene names | ACRV1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1). | |||||
|
CLIC5_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NZA1, Q5T4Z0, Q96JT5, Q9BWZ0 | Gene names | CLIC5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 5. | |||||
|
CLIC5_MOUSE
|
||||||
| NC score | 0.996377 (rank : 2) | θ value | 1.16911e-132 (rank : 2) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BXK9, Q3U0H8 | Gene names | Clic5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 5. | |||||
|
CLIC4_MOUSE
|
||||||
| NC score | 0.992503 (rank : 3) | θ value | 2.89973e-107 (rank : 4) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QYB1, Q8BMG5 | Gene names | Clic4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 4 (mc3s5/mtCLIC). | |||||
|
CLIC4_HUMAN
|
||||||
| NC score | 0.992414 (rank : 4) | θ value | 7.63087e-108 (rank : 3) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y696, Q9UFW9, Q9UQJ6 | Gene names | CLIC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 4 (Intracellular chloride ion channel protein p64H1). | |||||
|
CLIC2_HUMAN
|
||||||
| NC score | 0.990631 (rank : 5) | θ value | 1.88829e-90 (rank : 7) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15247, O15174, Q5JT80, Q8TCE3 | Gene names | CLIC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 2 (XAP121). | |||||
|
CLIC1_HUMAN
|
||||||
| NC score | 0.986513 (rank : 6) | θ value | 3.44927e-84 (rank : 8) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O00299, Q15089 | Gene names | CLIC1, NCC27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 1 (Nuclear chloride ion channel 27) (NCC27) (Chloride channel ABP) (Regulatory nuclear chloride ion channel protein) (hRNCC). | |||||
|
CLIC1_MOUSE
|
||||||
| NC score | 0.986354 (rank : 7) | θ value | 2.23575e-83 (rank : 9) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Z1Q5 | Gene names | Clic1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 1 (Nuclear chloride ion channel 27) (NCC27). | |||||
|
CLIC3_HUMAN
|
||||||
| NC score | 0.971859 (rank : 8) | θ value | 5.92465e-60 (rank : 10) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95833 | Gene names | CLIC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 3. | |||||
|
CLIC3_MOUSE
|
||||||
| NC score | 0.969939 (rank : 9) | θ value | 1.45929e-58 (rank : 11) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D7P7 | Gene names | Clic3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel protein 3. | |||||
|
CLIC6_MOUSE
|
||||||
| NC score | 0.953452 (rank : 10) | θ value | 2.15048e-102 (rank : 5) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BHB9 | Gene names | Clic6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
CLIC6_HUMAN
|
||||||
| NC score | 0.867294 (rank : 11) | θ value | 2.01279e-100 (rank : 6) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
GSTO1_MOUSE
|
||||||
| NC score | 0.315660 (rank : 12) | θ value | 5.44631e-05 (rank : 12) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O09131, Q3TH87 | Gene names | Gsto1, Gstx, Gtsttl | |||
|
Domain Architecture |

|
|||||
| Description | Glutathione transferase omega-1 (EC 2.5.1.18) (GSTO 1-1) (p28). | |||||
|
GSTO1_HUMAN
|
||||||
| NC score | 0.208673 (rank : 13) | θ value | 0.00869519 (rank : 13) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P78417, Q5TA03, Q7Z3T2 | Gene names | GSTO1, GSTTLP28 | |||
|
Domain Architecture |

|
|||||
| Description | Glutathione transferase omega-1 (EC 2.5.1.18) (GSTO 1-1). | |||||
|
MAAI_MOUSE
|
||||||
| NC score | 0.102811 (rank : 14) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WVL0 | Gene names | Gstz1, Maai | |||
|
Domain Architecture |

|
|||||
| Description | Maleylacetoacetate isomerase (EC 5.2.1.2) (MAAI) (Glutathione S- transferase zeta 1) (EC 2.5.1.18) (GSTZ1-1). | |||||
|
GSTO2_MOUSE
|
||||||
| NC score | 0.100718 (rank : 15) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K2Q2 | Gene names | Gsto2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutathione transferase omega-2 (EC 2.5.1.18). | |||||
|
MAAI_HUMAN
|
||||||
| NC score | 0.097868 (rank : 16) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O43708, O15308, O75430, Q7Z610, Q9BV63 | Gene names | GSTZ1, MAAI | |||
|
Domain Architecture |

|
|||||
| Description | Maleylacetoacetate isomerase (EC 5.2.1.2) (MAAI) (Glutathione S- transferase zeta 1) (EC 2.5.1.18) (GSTZ1-1). | |||||
|
GDAP1_MOUSE
|
||||||
| NC score | 0.085969 (rank : 17) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O88741, Q8C7Q5, Q9CTN2 | Gene names | Gdap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ganglioside-induced differentiation-associated protein 1 (GDAP1). | |||||
|
GDAP1_HUMAN
|
||||||
| NC score | 0.085928 (rank : 18) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8TB36 | Gene names | GDAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ganglioside-induced differentiation-associated protein 1 (GDAP1). | |||||
|
GSTO2_HUMAN
|
||||||
| NC score | 0.069143 (rank : 19) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H4Y5, Q49TW5, Q5GM70, Q86WP3 | Gene names | GSTO2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutathione transferase omega-2 (EC 2.5.1.18) (GSTO-2). | |||||
|
ASPX_HUMAN
|
||||||
| NC score | 0.055025 (rank : 20) | θ value | θ > 10 (rank : 34) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26436 | Gene names | ACRV1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.020021 (rank : 21) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
TM7S3_HUMAN
|
||||||
| NC score | 0.019246 (rank : 22) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NS93, Q9NUS4 | Gene names | TM7SF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane 7 superfamily member 3 precursor (Seven span transmembrane protein). | |||||
|
CCD1A_HUMAN
|
||||||
| NC score | 0.015428 (rank : 23) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P1N0, Q7Z435, Q86XV0, Q8NF89, Q9H603, Q9NXI1 | Gene names | CC2D1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1) (Putative NF-kappa-B- activating protein 023N). | |||||
|
NMT1_MOUSE
|
||||||
| NC score | 0.013658 (rank : 24) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70310 | Gene names | Nmt1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycylpeptide N-tetradecanoyltransferase 1 (EC 2.3.1.97) (Peptide N- myristoyltransferase 1) (Myristoyl-CoA:protein N-myristoyltransferase 1) (NMT 1) (Type I N-myristoyltransferase). | |||||
|
NMT1_HUMAN
|
||||||
| NC score | 0.012899 (rank : 25) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30419, Q9UE09 | Gene names | NMT1, NMT | |||
|
Domain Architecture |

|
|||||
| Description | Glycylpeptide N-tetradecanoyltransferase 1 (EC 2.3.1.97) (Peptide N- myristoyltransferase 1) (Myristoyl-CoA:protein N-myristoyltransferase 1) (NMT 1) (Type I N-myristoyltransferase). | |||||
|
MECP2_MOUSE
|
||||||
| NC score | 0.012019 (rank : 26) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |

|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
RFIP1_MOUSE
|
||||||
| NC score | 0.010901 (rank : 27) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D620, Q3UBC2, Q8BN24 | Gene names | Rab11fip1, Rcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.010786 (rank : 28) | θ value | 1.06291 (rank : 19) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
ELL2_HUMAN
|
||||||
| NC score | 0.009234 (rank : 29) | θ value | 6.88961 (rank : 30) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00472 | Gene names | ELL2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL2. | |||||
|
PSN1_HUMAN
|
||||||
| NC score | 0.006907 (rank : 30) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49768, O95465, Q14762, Q15719, Q15720, Q96P33, Q9UIF0 | Gene names | PSEN1, AD3, PS1, PSNL1 | |||
|
Domain Architecture |

|
|||||
| Description | Presenilin-1 (EC 3.4.23.-) (PS-1) (Protein S182) [Contains: Presenilin-1 NTF subunit; Presenilin-1 CTF subunit; Presenilin-1 CTF12 (PS1-CTF12)]. | |||||
|
MUG1_MOUSE
|
||||||
| NC score | 0.006497 (rank : 31) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28665 | Gene names | Mug1, Mug-1 | |||
|
Domain Architecture |

|
|||||
| Description | Murinoglobulin-1 precursor (MuG1). | |||||
|
ZN148_HUMAN
|
||||||
| NC score | 0.005705 (rank : 32) | θ value | 0.125558 (rank : 14) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UQR1, O00389, O43591 | Gene names | ZNF148, ZBP89 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 148 (Zinc finger DNA-binding protein 89) (Transcription factor ZBP-89). | |||||
|
ZN148_MOUSE
|
||||||
| NC score | 0.004860 (rank : 33) | θ value | 0.62314 (rank : 17) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61624, P97475 | Gene names | Znf148, Zbp89, Zfp148 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 148 (Zinc finger DNA-binding protein 89) (Transcription factor ZBP-89) (G-rich box-binding protein) (Beta enolase repressor factor 1) (Transcription factor BFCOL1). | |||||
|
EVPL_MOUSE
|
||||||
| NC score | 0.003221 (rank : 34) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||