Please be patient as the page loads
|
ADO_HUMAN
|
||||||
| SwissProt Accessions | Q06278, O14765, Q9UPG6 | Gene names | AOX1, AO | |||
|
Domain Architecture |

|
|||||
| Description | Aldehyde oxidase (EC 1.2.3.1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ADO_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q06278, O14765, Q9UPG6 | Gene names | AOX1, AO | |||
|
Domain Architecture |

|
|||||
| Description | Aldehyde oxidase (EC 1.2.3.1). | |||||
|
ADO_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.997812 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54754, Q9WU85, Q9Z2Z5 | Gene names | Aox1, Ao, Ro | |||
|
Domain Architecture |

|
|||||
| Description | Aldehyde oxidase (EC 1.2.3.1) (Retinal oxidase). | |||||
|
XDH_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.985584 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P47989, Q16681, Q16712, Q4PJ16 | Gene names | XDH, XDHA | |||
|
Domain Architecture |

|
|||||
| Description | Xanthine dehydrogenase/oxidase [Includes: Xanthine dehydrogenase (EC 1.17.1.4) (XD); Xanthine oxidase (EC 1.17.3.2) (XO) (Xanthine oxidoreductase)]. | |||||
|
XDH_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.985240 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00519 | Gene names | Xdh | |||
|
Domain Architecture |

|
|||||
| Description | Xanthine dehydrogenase/oxidase [Includes: Xanthine dehydrogenase (EC 1.17.1.4) (XD); Xanthine oxidase (EC 1.17.3.2) (XO) (Xanthine oxidoreductase)]. | |||||
|
CR2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 5) | NC score | 0.016714 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P19070 | Gene names | Cr2 | |||
|
Domain Architecture |

|
|||||
| Description | Complement receptor type 2 precursor (Cr2) (Complement C3d receptor) (CD21 antigen). | |||||
|
KRA59_HUMAN
|
||||||
| θ value | 3.0926 (rank : 6) | NC score | 0.010205 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
RHG26_HUMAN
|
||||||
| θ value | 3.0926 (rank : 7) | NC score | 0.009035 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UNA1, O75117, Q9BYS6, Q9BYS7, Q9UJ00 | Gene names | ARHGAP26, GRAF, KIAA0621, OPHN1L | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 26 (Oligophrenin-1-like protein) (GTPase regulator associated with focal adhesion kinase). | |||||
|
S39A5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 8) | NC score | 0.017382 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D856, Q9D909 | Gene names | Slc39a5, Zip5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc transporter ZIP5 precursor (Solute carrier family 39 member 5). | |||||
|
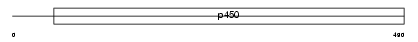
CP2S1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 9) | NC score | 0.004164 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DBX6 | Gene names | Cyp2s1 | |||
|
Domain Architecture |
|
|||||
| Description | Cytochrome P450 2S1 (EC 1.14.14.1) (CYPIIS1). | |||||
|
I12R1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 10) | NC score | 0.024312 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60837 | Gene names | Il12rb1, Il12rb | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-12 receptor beta-1 chain precursor (IL-12R-beta1) (Interleukin-12 receptor beta) (IL-12 receptor beta component) (CD212 antigen). | |||||
|
ATS5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 11) | NC score | 0.006228 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UNA0, Q9UKP2 | Gene names | ADAMTS5, ADAMTS11, ADMP2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-5 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 5) (ADAM-TS 5) (ADAM-TS5) (Aggrecanase-2) (ADMP-2) (ADAM-TS 11). | |||||
|
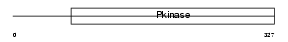
AURKB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 12) | NC score | -0.000036 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96GD4, O14630, O60446, O95083, Q96DV5, Q9UQ46 | Gene names | AURKB, AIK2, AIM1, ARK2, STK12 | |||
|
Domain Architecture |
|
|||||
| Description | Serine/threonine-protein kinase 12 (EC 2.7.11.1) (Aurora-B) (Aurora- and Ipl1-like midbody-associated protein 1) (AIM-1) (Aurora/IPL1- related kinase 2) (Aurora-related kinase 2) (STK-1). | |||||
|
MEGF9_MOUSE
|
||||||
| θ value | 5.27518 (rank : 13) | NC score | 0.010709 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
ATS5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 14) | NC score | 0.005688 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R001 | Gene names | Adamts5 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-5 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 5) (ADAM-TS 5) (ADAM-TS5) (Aggrecanase-2) (ADMP-2) (Implantin). | |||||
|
RGNEF_MOUSE
|
||||||
| θ value | 6.88961 (rank : 15) | NC score | 0.006049 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
STTP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 16) | NC score | 0.006492 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WG4, Q3TIH5, Q3TIT0, Q8CBW6, Q9ESY7 | Gene names | Statip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stat3-interacting protein (StIP1). | |||||
|
ADO_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q06278, O14765, Q9UPG6 | Gene names | AOX1, AO | |||
|
Domain Architecture |

|
|||||
| Description | Aldehyde oxidase (EC 1.2.3.1). | |||||
|
ADO_MOUSE
|
||||||
| NC score | 0.997812 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54754, Q9WU85, Q9Z2Z5 | Gene names | Aox1, Ao, Ro | |||
|
Domain Architecture |

|
|||||
| Description | Aldehyde oxidase (EC 1.2.3.1) (Retinal oxidase). | |||||
|
XDH_HUMAN
|
||||||
| NC score | 0.985584 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P47989, Q16681, Q16712, Q4PJ16 | Gene names | XDH, XDHA | |||
|
Domain Architecture |

|
|||||
| Description | Xanthine dehydrogenase/oxidase [Includes: Xanthine dehydrogenase (EC 1.17.1.4) (XD); Xanthine oxidase (EC 1.17.3.2) (XO) (Xanthine oxidoreductase)]. | |||||
|
XDH_MOUSE
|
||||||
| NC score | 0.985240 (rank : 4) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00519 | Gene names | Xdh | |||
|
Domain Architecture |

|
|||||
| Description | Xanthine dehydrogenase/oxidase [Includes: Xanthine dehydrogenase (EC 1.17.1.4) (XD); Xanthine oxidase (EC 1.17.3.2) (XO) (Xanthine oxidoreductase)]. | |||||
|
I12R1_MOUSE
|
||||||
| NC score | 0.024312 (rank : 5) | θ value | 4.03905 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60837 | Gene names | Il12rb1, Il12rb | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-12 receptor beta-1 chain precursor (IL-12R-beta1) (Interleukin-12 receptor beta) (IL-12 receptor beta component) (CD212 antigen). | |||||
|
S39A5_MOUSE
|
||||||
| NC score | 0.017382 (rank : 6) | θ value | 3.0926 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D856, Q9D909 | Gene names | Slc39a5, Zip5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc transporter ZIP5 precursor (Solute carrier family 39 member 5). | |||||
|
CR2_MOUSE
|
||||||
| NC score | 0.016714 (rank : 7) | θ value | 0.813845 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P19070 | Gene names | Cr2 | |||
|
Domain Architecture |

|
|||||
| Description | Complement receptor type 2 precursor (Cr2) (Complement C3d receptor) (CD21 antigen). | |||||
|
MEGF9_MOUSE
|
||||||
| NC score | 0.010709 (rank : 8) | θ value | 5.27518 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
KRA59_HUMAN
|
||||||
| NC score | 0.010205 (rank : 9) | θ value | 3.0926 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
RHG26_HUMAN
|
||||||
| NC score | 0.009035 (rank : 10) | θ value | 3.0926 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UNA1, O75117, Q9BYS6, Q9BYS7, Q9UJ00 | Gene names | ARHGAP26, GRAF, KIAA0621, OPHN1L | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 26 (Oligophrenin-1-like protein) (GTPase regulator associated with focal adhesion kinase). | |||||
|
STTP1_MOUSE
|
||||||
| NC score | 0.006492 (rank : 11) | θ value | 8.99809 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WG4, Q3TIH5, Q3TIT0, Q8CBW6, Q9ESY7 | Gene names | Statip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stat3-interacting protein (StIP1). | |||||
|
ATS5_HUMAN
|
||||||
| NC score | 0.006228 (rank : 12) | θ value | 5.27518 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UNA0, Q9UKP2 | Gene names | ADAMTS5, ADAMTS11, ADMP2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-5 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 5) (ADAM-TS 5) (ADAM-TS5) (Aggrecanase-2) (ADMP-2) (ADAM-TS 11). | |||||
|
RGNEF_MOUSE
|
||||||
| NC score | 0.006049 (rank : 13) | θ value | 6.88961 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
ATS5_MOUSE
|
||||||
| NC score | 0.005688 (rank : 14) | θ value | 6.88961 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R001 | Gene names | Adamts5 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-5 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 5) (ADAM-TS 5) (ADAM-TS5) (Aggrecanase-2) (ADMP-2) (Implantin). | |||||
|
CP2S1_MOUSE
|
||||||
| NC score | 0.004164 (rank : 15) | θ value | 4.03905 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DBX6 | Gene names | Cyp2s1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome P450 2S1 (EC 1.14.14.1) (CYPIIS1). | |||||
|
AURKB_HUMAN
|
||||||
| NC score | -0.000036 (rank : 16) | θ value | 5.27518 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96GD4, O14630, O60446, O95083, Q96DV5, Q9UQ46 | Gene names | AURKB, AIK2, AIM1, ARK2, STK12 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase 12 (EC 2.7.11.1) (Aurora-B) (Aurora- and Ipl1-like midbody-associated protein 1) (AIM-1) (Aurora/IPL1- related kinase 2) (Aurora-related kinase 2) (STK-1). | |||||