Please be patient as the page loads
|
UBAP1_MOUSE
|
||||||
| SwissProt Accessions | Q8BH48, Q8BQ80, Q8BSW6, Q9D749, Q9ERV5 | Gene names | Ubap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP) (Ubiquitin-associated protein NAG20). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
UBAP1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.992585 (rank : 2) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NZ09, Q4V759, Q5T7B3, Q6FI75, Q8NC52, Q8NCG6, Q8NCH9 | Gene names | UBAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP). | |||||
|
UBAP1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BH48, Q8BQ80, Q8BSW6, Q9D749, Q9ERV5 | Gene names | Ubap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP) (Ubiquitin-associated protein NAG20). | |||||
|
UBP5_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 3) | NC score | 0.173293 (rank : 4) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP5_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 4) | NC score | 0.173305 (rank : 3) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP13_HUMAN
|
||||||
| θ value | 0.813845 (rank : 5) | NC score | 0.125930 (rank : 5) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
COA1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 6) | NC score | 0.038226 (rank : 11) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13085 | Gene names | ACACA, ACAC, ACC1, ACCA | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
CYLN2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 7) | NC score | 0.011550 (rank : 25) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
CHD9_HUMAN
|
||||||
| θ value | 2.36792 (rank : 8) | NC score | 0.019104 (rank : 19) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 9) | NC score | 0.020004 (rank : 18) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
K1267_MOUSE
|
||||||
| θ value | 2.36792 (rank : 10) | NC score | 0.046255 (rank : 8) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80TG1, Q3TT88, Q3U5D8, Q3V3N3, Q7TMU3, Q80XP7, Q8R3L6, Q9D9G0 | Gene names | Kiaa1267 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1267. | |||||
|
TRI66_MOUSE
|
||||||
| θ value | 2.36792 (rank : 11) | NC score | 0.026445 (rank : 14) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
V2RX_MOUSE
|
||||||
| θ value | 2.36792 (rank : 12) | NC score | 0.021040 (rank : 17) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70410 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vomeronasal type-2 receptor X precursor (Putative pheromone receptor V2RX). | |||||
|
CT141_HUMAN
|
||||||
| θ value | 3.0926 (rank : 13) | NC score | 0.084302 (rank : 6) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NUB4 | Gene names | C20orf141 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf141. | |||||
|
FGFP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 14) | NC score | 0.035504 (rank : 12) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70514, Q62399 | Gene names | Fgfbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor-binding protein 1 precursor (FGF-binding protein 1) (FGF-BP1) (FGFBP-1) (FGF-BP). | |||||
|
K1267_HUMAN
|
||||||
| θ value | 3.0926 (rank : 15) | NC score | 0.044310 (rank : 10) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z3B3, Q6AW85, Q8IYH1, Q9NTE7, Q9UFT0, Q9ULF3 | Gene names | KIAA1267 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1267. | |||||
|
KNTC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 16) | NC score | 0.019095 (rank : 20) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 673 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14777, Q6PJX2 | Gene names | KNTC2, HEC, HEC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Hec1 (HsHec1) (Kinetochore-associated protein 2) (Highly expressed in cancer protein) (Retinoblastoma-associated protein HEC). | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 17) | NC score | 0.002922 (rank : 30) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
TRI66_HUMAN
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.024643 (rank : 15) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
CCD27_MOUSE
|
||||||
| θ value | 5.27518 (rank : 19) | NC score | 0.015272 (rank : 22) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3V036 | Gene names | Ccdc27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 27. | |||||
|
LGR5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 20) | NC score | 0.012921 (rank : 24) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75473, Q9UP75 | Gene names | LGR5, GPR49, GPR67 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (Orphan G-protein coupled receptor HG38) (G-protein coupled receptor 49) (G-protein coupled receptor 67). | |||||
|
LGR5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 21) | NC score | 0.013017 (rank : 23) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z1P4 | Gene names | Lgr5, Fex, Gpr49 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (G-protein coupled receptor 49) (Orphan G-protein coupled receptor FEX). | |||||
|
CCHL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 22) | NC score | 0.049585 (rank : 7) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53702 | Gene names | Hccs, Cchl | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome c-type heme lyase (EC 4.4.1.17) (CCHL) (Holocytochrome c- type synthase). | |||||
|
FA44A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 23) | NC score | 0.016719 (rank : 21) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NFC6, Q6P0M8 | Gene names | FAM44A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM44A. | |||||
|
GLIS3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | 0.002026 (rank : 32) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 861 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6XP49, Q8BI60 | Gene names | Glis3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLIS3 (GLI-similar 3). | |||||
|
LIPA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.005505 (rank : 29) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
WNK4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.002817 (rank : 31) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 961 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80UE6, Q4VAC1, Q80XB5, Q80XN2, Q8R0N0, Q8R340 | Gene names | Wnk4, Prkwnk4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
CCHL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.045898 (rank : 9) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53701, Q502X8 | Gene names | HCCS, CCHL | |||
|
Domain Architecture |
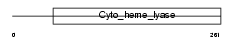
|
|||||
| Description | Cytochrome c-type heme lyase (EC 4.4.1.17) (CCHL) (Holocytochrome c- type synthase). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.009921 (rank : 26) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
MUC13_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.031944 (rank : 13) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3R2, Q6UWD9, Q9NXT5 | Gene names | MUC13, DRCC1, RECC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-13 precursor (Down-regulated in colon cancer 1). | |||||
|
PAPOA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.009436 (rank : 27) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61183, Q61208, Q61209, Q8K4X2 | Gene names | Papola, Pap, Plap | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase alpha (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase). | |||||
|
RD23A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.024543 (rank : 16) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54725 | Gene names | RAD23A | |||
|
Domain Architecture |
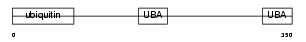
|
|||||
| Description | UV excision repair protein RAD23 homolog A (hHR23A). | |||||
|
SON_HUMAN
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.007248 (rank : 28) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
UBAP1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BH48, Q8BQ80, Q8BSW6, Q9D749, Q9ERV5 | Gene names | Ubap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP) (Ubiquitin-associated protein NAG20). | |||||
|
UBAP1_HUMAN
|
||||||
| NC score | 0.992585 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NZ09, Q4V759, Q5T7B3, Q6FI75, Q8NC52, Q8NCG6, Q8NCH9 | Gene names | UBAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP). | |||||
|
UBP5_MOUSE
|
||||||
| NC score | 0.173305 (rank : 3) | θ value | 0.000461057 (rank : 4) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP5_HUMAN
|
||||||
| NC score | 0.173293 (rank : 4) | θ value | 0.000461057 (rank : 3) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP13_HUMAN
|
||||||
| NC score | 0.125930 (rank : 5) | θ value | 0.813845 (rank : 5) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
CT141_HUMAN
|
||||||
| NC score | 0.084302 (rank : 6) | θ value | 3.0926 (rank : 13) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NUB4 | Gene names | C20orf141 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf141. | |||||
|
CCHL_MOUSE
|
||||||
| NC score | 0.049585 (rank : 7) | θ value | 6.88961 (rank : 22) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53702 | Gene names | Hccs, Cchl | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome c-type heme lyase (EC 4.4.1.17) (CCHL) (Holocytochrome c- type synthase). | |||||
|
K1267_MOUSE
|
||||||
| NC score | 0.046255 (rank : 8) | θ value | 2.36792 (rank : 10) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80TG1, Q3TT88, Q3U5D8, Q3V3N3, Q7TMU3, Q80XP7, Q8R3L6, Q9D9G0 | Gene names | Kiaa1267 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1267. | |||||
|
CCHL_HUMAN
|
||||||
| NC score | 0.045898 (rank : 9) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53701, Q502X8 | Gene names | HCCS, CCHL | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome c-type heme lyase (EC 4.4.1.17) (CCHL) (Holocytochrome c- type synthase). | |||||
|
K1267_HUMAN
|
||||||
| NC score | 0.044310 (rank : 10) | θ value | 3.0926 (rank : 15) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z3B3, Q6AW85, Q8IYH1, Q9NTE7, Q9UFT0, Q9ULF3 | Gene names | KIAA1267 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1267. | |||||
|
COA1_HUMAN
|
||||||
| NC score | 0.038226 (rank : 11) | θ value | 1.81305 (rank : 6) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13085 | Gene names | ACACA, ACAC, ACC1, ACCA | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
FGFP1_MOUSE
|
||||||
| NC score | 0.035504 (rank : 12) | θ value | 3.0926 (rank : 14) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70514, Q62399 | Gene names | Fgfbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor-binding protein 1 precursor (FGF-binding protein 1) (FGF-BP1) (FGFBP-1) (FGF-BP). | |||||
|
MUC13_HUMAN
|
||||||
| NC score | 0.031944 (rank : 13) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3R2, Q6UWD9, Q9NXT5 | Gene names | MUC13, DRCC1, RECC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-13 precursor (Down-regulated in colon cancer 1). | |||||
|
TRI66_MOUSE
|
||||||
| NC score | 0.026445 (rank : 14) | θ value | 2.36792 (rank : 11) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TRI66_HUMAN
|
||||||
| NC score | 0.024643 (rank : 15) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
RD23A_HUMAN
|
||||||
| NC score | 0.024543 (rank : 16) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54725 | Gene names | RAD23A | |||
|
Domain Architecture |

|
|||||
| Description | UV excision repair protein RAD23 homolog A (hHR23A). | |||||
|
V2RX_MOUSE
|
||||||
| NC score | 0.021040 (rank : 17) | θ value | 2.36792 (rank : 12) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70410 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vomeronasal type-2 receptor X precursor (Putative pheromone receptor V2RX). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.020004 (rank : 18) | θ value | 2.36792 (rank : 9) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
CHD9_HUMAN
|
||||||
| NC score | 0.019104 (rank : 19) | θ value | 2.36792 (rank : 8) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
KNTC2_HUMAN
|
||||||
| NC score | 0.019095 (rank : 20) | θ value | 4.03905 (rank : 16) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 673 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14777, Q6PJX2 | Gene names | KNTC2, HEC, HEC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Hec1 (HsHec1) (Kinetochore-associated protein 2) (Highly expressed in cancer protein) (Retinoblastoma-associated protein HEC). | |||||
|
FA44A_HUMAN
|
||||||
| NC score | 0.016719 (rank : 21) | θ value | 6.88961 (rank : 23) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NFC6, Q6P0M8 | Gene names | FAM44A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM44A. | |||||
|
CCD27_MOUSE
|
||||||
| NC score | 0.015272 (rank : 22) | θ value | 5.27518 (rank : 19) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3V036 | Gene names | Ccdc27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 27. | |||||
|
LGR5_MOUSE
|
||||||
| NC score | 0.013017 (rank : 23) | θ value | 5.27518 (rank : 21) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z1P4 | Gene names | Lgr5, Fex, Gpr49 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (G-protein coupled receptor 49) (Orphan G-protein coupled receptor FEX). | |||||
|
LGR5_HUMAN
|
||||||
| NC score | 0.012921 (rank : 24) | θ value | 5.27518 (rank : 20) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75473, Q9UP75 | Gene names | LGR5, GPR49, GPR67 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (Orphan G-protein coupled receptor HG38) (G-protein coupled receptor 49) (G-protein coupled receptor 67). | |||||
|
CYLN2_HUMAN
|
||||||
| NC score | 0.011550 (rank : 25) | θ value | 1.81305 (rank : 7) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.009921 (rank : 26) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
PAPOA_MOUSE
|
||||||
| NC score | 0.009436 (rank : 27) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61183, Q61208, Q61209, Q8K4X2 | Gene names | Papola, Pap, Plap | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase alpha (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.007248 (rank : 28) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
LIPA1_HUMAN
|
||||||
| NC score | 0.005505 (rank : 29) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | 0.002922 (rank : 30) | θ value | 4.03905 (rank : 17) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
WNK4_MOUSE
|
||||||
| NC score | 0.002817 (rank : 31) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 961 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80UE6, Q4VAC1, Q80XB5, Q80XN2, Q8R0N0, Q8R340 | Gene names | Wnk4, Prkwnk4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
GLIS3_MOUSE
|
||||||
| NC score | 0.002026 (rank : 32) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 861 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6XP49, Q8BI60 | Gene names | Glis3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLIS3 (GLI-similar 3). | |||||