Please be patient as the page loads
|
TF3C1_MOUSE
|
||||||
| SwissProt Accessions | Q8K284, Q8CAL9 | Gene names | Gtf3c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TF3C1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.973778 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12789, Q12838, Q9Y4W9 | Gene names | GTF3C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
TF3C1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q8K284, Q8CAL9 | Gene names | Gtf3c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 3) | NC score | 0.075754 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CN155_HUMAN
|
||||||
| θ value | 0.279714 (rank : 4) | NC score | 0.062287 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 5) | NC score | 0.053764 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.47712 (rank : 6) | NC score | 0.046421 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.62314 (rank : 7) | NC score | 0.027222 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RFIP5_MOUSE
|
||||||
| θ value | 0.62314 (rank : 8) | NC score | 0.035538 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R361 | Gene names | Rab11fip5, D6Ertd32e, Rip11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 5 (Rab11-FIP5) (Rab11-interacting protein Rip11). | |||||
|
LRC59_HUMAN
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.025888 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96AG4, Q9P189 | Gene names | LRRC59 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 59. | |||||
|
MAGI1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.027212 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96QZ7, O00309, O43863, O75085, Q96QZ8, Q96QZ9 | Gene names | MAGI1, BAIAP1, BAP1, TNRC19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1) (Atrophin-1-interacting protein 3) (AIP3) (WW domain-containing protein 3) (WWP3) (Trinucleotide repeat-containing gene 19 protein). | |||||
|
MAGI1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 11) | NC score | 0.027483 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6RHR9, O54893, O54894, O54895 | Gene names | Magi1, Baiap1, Bap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1). | |||||
|
CDC37_HUMAN
|
||||||
| θ value | 1.06291 (rank : 12) | NC score | 0.047524 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |
|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
GLU2B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.038483 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P14314, Q96BU9, Q96D06, Q9P0W9 | Gene names | PRKCSH, G19P1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
NKX12_MOUSE
|
||||||
| θ value | 1.06291 (rank : 14) | NC score | 0.011192 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42580, Q9JHF2, Q9JKW5, Q9JKW6, Q9JKW9, Q9JKX0, Q9JKX1 | Gene names | Nkx1-2, Nkx1.1, Sax1 | |||
|
Domain Architecture |

|
|||||
| Description | NK1 transcription factor-related protein 2 (Homeobox protein SAX-1) (NKX-1.1). | |||||
|
SF3B1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 15) | NC score | 0.046084 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75533 | Gene names | SF3B1, SAP155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
SF3B1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.046080 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99NB9, Q9CSK5 | Gene names | Sf3b1, Sap155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
ZBT37_HUMAN
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.005462 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 815 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5TC79, Q5TC80, Q96M87, Q9BQ88 | Gene names | ZBTB37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 37. | |||||
|
T2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 18) | NC score | 0.050139 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q06666 | Gene names | Srst, T2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Octapeptide-repeat protein T2. | |||||
|
EHMT2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.014924 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96KQ7, Q14349, Q6PK06, Q96MH5, Q96QD0, Q9UQL8, Q9Y331 | Gene names | EHMT2, BAT8, G9A, NG36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.016162 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
MFAP1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.048996 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P55081 | Gene names | MFAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Microfibrillar-associated protein 1. | |||||
|
MFAP1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.041815 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CQU1, Q3TU29, Q8CCL1, Q9CSJ5 | Gene names | Mfap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microfibrillar-associated protein 1. | |||||
|
RBM25_HUMAN
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.026670 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
TNIK_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.007896 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
A4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.027827 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P05067, P09000, P78438, Q13764, Q13778, Q13793, Q16011, Q16014, Q16019, Q16020, Q9BT38, Q9UCA9, Q9UCB6, Q9UCC8, Q9UCD1, Q9UQ58 | Gene names | APP, A4, AD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein) (Cerebral vascular amyloid peptide) (CVAP) (Protease nexin-II) (PN-II) (APPI) (PreA4) [Contains: Soluble APP-alpha (S-APP- alpha); Soluble APP-beta (S-APP-beta); C99; Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)); Gamma-CTF(57) (Gamma- secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
A4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.028109 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P12023, P97487, P97942, Q99K32 | Gene names | App | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein homolog) (Amyloidogenic glycoprotein) (AG) [Contains: Soluble APP-alpha (S-APP-alpha); Soluble APP-beta (S-APP-beta); C99 (APP-C99); Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)) (APP-C59); Gamma-CTF(57) (Gamma-secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)) (APP-C57); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
GPTC8_HUMAN
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.047527 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
M4K4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.007462 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1589 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O95819, O75172, Q9NST7 | Gene names | MAP4K4, HGK, KIAA0687, NIK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 4 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 4) (MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase HGK) (Nck-interacting kinase). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.034460 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PPIL2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.037218 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13356, Q13357, Q8TAH2, Q9BWR8 | Gene names | PPIL2 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase-like 2 (EC 5.2.1.8) (PPIase) (Rotamase) (Cyclophilin-60) (Cyclophilin-like protein Cyp-60). | |||||
|
RN168_MOUSE
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.023221 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q80XJ2 | Gene names | Rnf168 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 168. | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.013577 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
WNK1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.007346 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1158 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H4A3, O15052, Q86WL5, Q8N673, Q9P1S9 | Gene names | WNK1, KDP, KIAA0344, PRKWNK1 | |||
|
Domain Architecture |
|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1) (Kinase deficient protein). | |||||
|
CABYR_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.028670 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75952, Q8WXW5, Q9HAY3, Q9HAY4, Q9HAY5, Q9HCY9 | Gene names | CABYR, CBP86, FSP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding tyrosine phosphorylation-regulated protein (Calcium- binding protein 86) (Testis-specific calcium-binding protein CBP86) (Fibrousheathin-2) (FSP-2). | |||||
|
CAF1A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.024305 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QWF0 | Gene names | Chaf1a, Caip150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
CC45L_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.035833 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75419, O60856, Q6UW54, Q9UP68 | Gene names | CDC45L, CDC45L2 | |||
|
Domain Architecture |

|
|||||
| Description | CDC45-related protein (PORC-PI-1) (Cdc45). | |||||
|
CDC37_MOUSE
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.044515 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61081, Q3TGP0 | Gene names | Cdc37 | |||
|
Domain Architecture |
|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
FA98B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.053600 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q52LJ0, Q8N935 | Gene names | FAM98B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98B. | |||||
|
LRC59_MOUSE
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.021796 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q922Q8, Q3TJ35, Q3TLC7, Q3TWT9, Q3TX86 | Gene names | Lrrc59 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 59. | |||||
|
PPIL2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.035333 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D787, Q542A2, Q9CZL1 | Gene names | Ppil2 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase-like 2 (EC 5.2.1.8) (PPIase) (Rotamase) (Cyclophilin-60) (Cyclophilin-like protein Cyp-60). | |||||
|
XE7_HUMAN
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.021129 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q02040, Q02832 | Gene names | CXYorf3, DXYS155E, XE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-lymphocyte antigen precursor (B-lymphocyte surface antigen) (721P) (Protein XE7). | |||||
|
CC45L_MOUSE
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.034028 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1X9, O70547, Q9QUK1, Q9R212 | Gene names | Cdc45l, Cdc45l2 | |||
|
Domain Architecture |

|
|||||
| Description | CDC45-related protein (PORC-PI-1). | |||||
|
CS007_HUMAN
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.023831 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
DDX23_HUMAN
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.008914 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9BUQ8, O43188 | Gene names | DDX23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX23 (EC 3.6.1.-) (DEAD box protein 23) (100 kDa U5 snRNP-specific protein) (U5-100kD) (PRP28 homolog). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.037354 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PPCKC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.016327 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2V4, Q3UEH3 | Gene names | Pck1, Pepck | |||
|
Domain Architecture |

|
|||||
| Description | Phosphoenolpyruvate carboxykinase, cytosolic [GTP] (EC 4.1.1.32) (Phosphoenolpyruvate carboxylase) (PEPCK-C). | |||||
|
SON_MOUSE
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.021840 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
STK10_HUMAN
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.007497 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O94804, Q9UIW4 | Gene names | STK10, LOK | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase 10 (EC 2.7.11.1) (Lymphocyte-oriented kinase). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.031141 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
ZC313_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.037314 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
ACINU_HUMAN
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.035100 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.021178 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
CALD1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.019120 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
FA98B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.025262 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VD1, Q9CS28, Q9CS37 | Gene names | Fam98b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98B. | |||||
|
FANCE_HUMAN
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.028752 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HB96, Q4ZGH2 | Gene names | FANCE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group E protein (Protein FACE). | |||||
|
PAR3L_HUMAN
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.014812 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8TEW8, Q8IUC7, Q8IUC9, Q96DK9, Q96N09, Q96NX6, Q96NX7, Q96Q29 | Gene names | PARD3B, ALS2CR19, PAR3B, PAR3L | |||
|
Domain Architecture |

|
|||||
| Description | Partitioning-defective 3 homolog B (PAR3-beta) (Partitioning-defective 3-like protein) (PAR3-L protein) (Amyotrophic lateral sclerosis 2 chromosome region candidate gene 19 protein). | |||||
|
PELP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.028678 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
SAFB2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.025621 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80YR5, Q8K153 | Gene names | Safb2 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
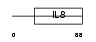
CCL21_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.014989 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00585 | Gene names | CCL21, SCYA21 | |||
|
Domain Architecture |
|
|||||
| Description | Small inducible cytokine A21 precursor (CCL21) (Beta chemokine exodus- 2) (6Ckine) (Secondary lymphoid-tissue chemokine) (SLC). | |||||
|
CGRE1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.034725 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.018035 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
MAP4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.020667 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
MEFV_HUMAN
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.005539 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O15553, Q96PN4, Q96PN5 | Gene names | MEFV, MEF | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
MYO6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.005207 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q64331 | Gene names | Myo6, Sv | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
PGCA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.007951 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
S6A11_MOUSE
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.006558 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P31650 | Gene names | Slc6a11, Gabt4, Gat-4, Gat4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium- and chloride-dependent GABA transporter 4 (GAT4). | |||||
|
SAKS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.016844 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q04323, Q9BV93, Q9BVV5 | Gene names | SAKS1 | |||
|
Domain Architecture |

|
|||||
| Description | SAPK substrate protein 1 (UBA/UBX 33.3 kDa protein). | |||||
|
SAS10_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.018766 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JI13, Q3UL95, Q8R5C6, Q9JJ12 | Gene names | Crlz1, Crl1, Sas10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Something about silencing protein 10 (Disrupter of silencing SAS10) (Charged amino acid-rich leucine zipper 1) (Crl-1). | |||||
|
WWC3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.016227 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
ARI4A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.020769 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
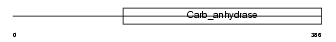
CAH9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.007263 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16790 | Gene names | CA9, G250, MN | |||
|
Domain Architecture |
|
|||||
| Description | Carbonic anhydrase 9 precursor (EC 4.2.1.1) (Carbonic anhydrase IX) (Carbonate dehydratase IX) (CA-IX) (CAIX) (Membrane antigen MN) (P54/58N) (Renal cell carcinoma-associated antigen G250) (RCC- associated antigen G250) (pMW1). | |||||
|
CD008_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.019154 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P78312, O43607, P78311, P78313, Q9UEG8 | Gene names | C4orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf8 (Protein IT14). | |||||
|
CTRO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.005484 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
CTRO_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.007069 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
E41L1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.006835 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2H5 | Gene names | Epb41l1, Epb4, Epb4.1l1 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 1 (Neuronal protein 4.1) (4.1N). | |||||
|
ELF4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.007413 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
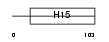
H10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.019662 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P10922 | Gene names | H1f0, H1fv | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1' (H1.0) (H1(0)). | |||||
|
INCE_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.011246 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
ITSN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.010453 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
KBTBA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.006548 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60662 | Gene names | KBTBD10, KRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch repeat and BTB domain-containing protein 10 (Kelch-related protein 1) (Kel-like protein 23) (Sarcosin). | |||||
|
LTB1L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.004701 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14766 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1L precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
LTB1S_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.004333 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P22064, Q8TD95 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1S precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
MASTL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.003743 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96GX5, Q5T8D5, Q5T8D7, Q8NCD6, Q96SJ5 | Gene names | MASTL, THC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase-like (EC 2.7.11.1). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.015153 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MTO1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.013292 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q923Z3, Q9D2Q5 | Gene names | Mto1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MTO1 homolog, mitochondrial precursor. | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.025134 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
RN146_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.014090 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NTX7, Q6FIB2, Q7L8H4, Q96K03, Q9NTX6 | Gene names | RNF146 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 146 (Dactylidin). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.014274 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
SAFB2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.018145 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14151, Q8TB13 | Gene names | SAFB2, KIAA0138 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
TOP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.018422 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q04750 | Gene names | Top1, Top-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 1 (EC 5.99.1.2) (DNA topoisomerase I). | |||||
|
ZFP91_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.003245 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 769 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96JP5, Q86V47, Q96JP4, Q96QA3 | Gene names | ZFP91 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 91 homolog (Zfp-91). | |||||
|
ZFP91_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.002576 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62511, Q8BPY3, Q8C2B4 | Gene names | Zfp91, Pzf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 91 (Zfp-91) (Zinc finger protein PZF) (Penta Zf protein). | |||||
|
TF3C1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q8K284, Q8CAL9 | Gene names | Gtf3c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
TF3C1_HUMAN
|
||||||
| NC score | 0.973778 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12789, Q12838, Q9Y4W9 | Gene names | GTF3C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.075754 (rank : 3) | θ value | 0.00390308 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.062287 (rank : 4) | θ value | 0.279714 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.053764 (rank : 5) | θ value | 0.279714 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
FA98B_HUMAN
|
||||||
| NC score | 0.053600 (rank : 6) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q52LJ0, Q8N935 | Gene names | FAM98B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98B. | |||||
|
T2_MOUSE
|
||||||
| NC score | 0.050139 (rank : 7) | θ value | 1.38821 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q06666 | Gene names | Srst, T2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Octapeptide-repeat protein T2. | |||||
|
MFAP1_HUMAN
|
||||||
| NC score | 0.048996 (rank : 8) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P55081 | Gene names | MFAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Microfibrillar-associated protein 1. | |||||
|
GPTC8_HUMAN
|
||||||
| NC score | 0.047527 (rank : 9) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
CDC37_HUMAN
|
||||||
| NC score | 0.047524 (rank : 10) | θ value | 1.06291 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |

|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.046421 (rank : 11) | θ value | 0.47712 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
SF3B1_HUMAN
|
||||||
| NC score | 0.046084 (rank : 12) | θ value | 1.06291 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75533 | Gene names | SF3B1, SAP155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
SF3B1_MOUSE
|
||||||
| NC score | 0.046080 (rank : 13) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99NB9, Q9CSK5 | Gene names | Sf3b1, Sap155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
CDC37_MOUSE
|
||||||
| NC score | 0.044515 (rank : 14) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61081, Q3TGP0 | Gene names | Cdc37 | |||
|
Domain Architecture |

|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
MFAP1_MOUSE
|
||||||
| NC score | 0.041815 (rank : 15) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CQU1, Q3TU29, Q8CCL1, Q9CSJ5 | Gene names | Mfap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microfibrillar-associated protein 1. | |||||
|
GLU2B_HUMAN
|
||||||
| NC score | 0.038483 (rank : 16) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P14314, Q96BU9, Q96D06, Q9P0W9 | Gene names | PRKCSH, G19P1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.037354 (rank : 17) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
ZC313_HUMAN
|
||||||
| NC score | 0.037314 (rank : 18) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
PPIL2_HUMAN
|
||||||
| NC score | 0.037218 (rank : 19) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13356, Q13357, Q8TAH2, Q9BWR8 | Gene names | PPIL2 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase-like 2 (EC 5.2.1.8) (PPIase) (Rotamase) (Cyclophilin-60) (Cyclophilin-like protein Cyp-60). | |||||
|
CC45L_HUMAN
|
||||||
| NC score | 0.035833 (rank : 20) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75419, O60856, Q6UW54, Q9UP68 | Gene names | CDC45L, CDC45L2 | |||
|
Domain Architecture |

|
|||||
| Description | CDC45-related protein (PORC-PI-1) (Cdc45). | |||||
|
RFIP5_MOUSE
|
||||||
| NC score | 0.035538 (rank : 21) | θ value | 0.62314 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R361 | Gene names | Rab11fip5, D6Ertd32e, Rip11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 5 (Rab11-FIP5) (Rab11-interacting protein Rip11). | |||||
|
PPIL2_MOUSE
|
||||||
| NC score | 0.035333 (rank : 22) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D787, Q542A2, Q9CZL1 | Gene names | Ppil2 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase-like 2 (EC 5.2.1.8) (PPIase) (Rotamase) (Cyclophilin-60) (Cyclophilin-like protein Cyp-60). | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.035100 (rank : 23) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
CGRE1_HUMAN
|
||||||
| NC score | 0.034725 (rank : 24) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.034460 (rank : 25) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
CC45L_MOUSE
|
||||||
| NC score | 0.034028 (rank : 26) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1X9, O70547, Q9QUK1, Q9R212 | Gene names | Cdc45l, Cdc45l2 | |||
|
Domain Architecture |

|
|||||
| Description | CDC45-related protein (PORC-PI-1). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.031141 (rank : 27) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
FANCE_HUMAN
|
||||||
| NC score | 0.028752 (rank : 28) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HB96, Q4ZGH2 | Gene names | FANCE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group E protein (Protein FACE). | |||||
|
PELP1_HUMAN
|
||||||
| NC score | 0.028678 (rank : 29) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
CABYR_HUMAN
|
||||||
| NC score | 0.028670 (rank : 30) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75952, Q8WXW5, Q9HAY3, Q9HAY4, Q9HAY5, Q9HCY9 | Gene names | CABYR, CBP86, FSP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding tyrosine phosphorylation-regulated protein (Calcium- binding protein 86) (Testis-specific calcium-binding protein CBP86) (Fibrousheathin-2) (FSP-2). | |||||
|
A4_MOUSE
|
||||||
| NC score | 0.028109 (rank : 31) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P12023, P97487, P97942, Q99K32 | Gene names | App | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein homolog) (Amyloidogenic glycoprotein) (AG) [Contains: Soluble APP-alpha (S-APP-alpha); Soluble APP-beta (S-APP-beta); C99 (APP-C99); Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)) (APP-C59); Gamma-CTF(57) (Gamma-secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)) (APP-C57); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
A4_HUMAN
|
||||||
| NC score | 0.027827 (rank : 32) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P05067, P09000, P78438, Q13764, Q13778, Q13793, Q16011, Q16014, Q16019, Q16020, Q9BT38, Q9UCA9, Q9UCB6, Q9UCC8, Q9UCD1, Q9UQ58 | Gene names | APP, A4, AD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein) (Cerebral vascular amyloid peptide) (CVAP) (Protease nexin-II) (PN-II) (APPI) (PreA4) [Contains: Soluble APP-alpha (S-APP- alpha); Soluble APP-beta (S-APP-beta); C99; Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)); Gamma-CTF(57) (Gamma- secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
MAGI1_MOUSE
|
||||||
| NC score | 0.027483 (rank : 33) | θ value | 0.813845 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6RHR9, O54893, O54894, O54895 | Gene names | Magi1, Baiap1, Bap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.027222 (rank : 34) | θ value | 0.62314 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
MAGI1_HUMAN
|
||||||
| NC score | 0.027212 (rank : 35) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96QZ7, O00309, O43863, O75085, Q96QZ8, Q96QZ9 | Gene names | MAGI1, BAIAP1, BAP1, TNRC19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1) (Atrophin-1-interacting protein 3) (AIP3) (WW domain-containing protein 3) (WWP3) (Trinucleotide repeat-containing gene 19 protein). | |||||
|
RBM25_HUMAN
|
||||||
| NC score | 0.026670 (rank : 36) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
LRC59_HUMAN
|
||||||
| NC score | 0.025888 (rank : 37) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96AG4, Q9P189 | Gene names | LRRC59 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 59. | |||||
|
SAFB2_MOUSE
|
||||||
| NC score | 0.025621 (rank : 38) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80YR5, Q8K153 | Gene names | Safb2 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
FA98B_MOUSE
|
||||||
| NC score | 0.025262 (rank : 39) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VD1, Q9CS28, Q9CS37 | Gene names | Fam98b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98B. | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.025134 (rank : 40) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
CAF1A_MOUSE
|
||||||
| NC score | 0.024305 (rank : 41) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QWF0 | Gene names | Chaf1a, Caip150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
CS007_HUMAN
|
||||||
| NC score | 0.023831 (rank : 42) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
RN168_MOUSE
|
||||||
| NC score | 0.023221 (rank : 43) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q80XJ2 | Gene names | Rnf168 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 168. | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.021840 (rank : 44) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
LRC59_MOUSE
|
||||||
| NC score | 0.021796 (rank : 45) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q922Q8, Q3TJ35, Q3TLC7, Q3TWT9, Q3TX86 | Gene names | Lrrc59 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 59. | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.021178 (rank : 46) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
XE7_HUMAN
|
||||||
| NC score | 0.021129 (rank : 47) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q02040, Q02832 | Gene names | CXYorf3, DXYS155E, XE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-lymphocyte antigen precursor (B-lymphocyte surface antigen) (721P) (Protein XE7). | |||||
|
ARI4A_HUMAN
|
||||||
| NC score | 0.020769 (rank : 48) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
MAP4_MOUSE
|
||||||
| NC score | 0.020667 (rank : 49) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
H10_MOUSE
|
||||||
| NC score | 0.019662 (rank : 50) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P10922 | Gene names | H1f0, H1fv | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1' (H1.0) (H1(0)). | |||||
|
CD008_HUMAN
|
||||||
| NC score | 0.019154 (rank : 51) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P78312, O43607, P78311, P78313, Q9UEG8 | Gene names | C4orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf8 (Protein IT14). | |||||
|
CALD1_HUMAN
|
||||||
| NC score | 0.019120 (rank : 52) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
SAS10_MOUSE
|
||||||
| NC score | 0.018766 (rank : 53) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JI13, Q3UL95, Q8R5C6, Q9JJ12 | Gene names | Crlz1, Crl1, Sas10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Something about silencing protein 10 (Disrupter of silencing SAS10) (Charged amino acid-rich leucine zipper 1) (Crl-1). | |||||
|
TOP1_MOUSE
|
||||||
| NC score | 0.018422 (rank : 54) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q04750 | Gene names | Top1, Top-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 1 (EC 5.99.1.2) (DNA topoisomerase I). | |||||
|
SAFB2_HUMAN
|
||||||
| NC score | 0.018145 (rank : 55) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14151, Q8TB13 | Gene names | SAFB2, KIAA0138 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.018035 (rank : 56) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
SAKS1_HUMAN
|
||||||
| NC score | 0.016844 (rank : 57) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q04323, Q9BV93, Q9BVV5 | Gene names | SAKS1 | |||
|
Domain Architecture |

|
|||||
| Description | SAPK substrate protein 1 (UBA/UBX 33.3 kDa protein). | |||||
|
PPCKC_MOUSE
|
||||||
| NC score | 0.016327 (rank : 58) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2V4, Q3UEH3 | Gene names | Pck1, Pepck | |||
|
Domain Architecture |

|
|||||
| Description | Phosphoenolpyruvate carboxykinase, cytosolic [GTP] (EC 4.1.1.32) (Phosphoenolpyruvate carboxylase) (PEPCK-C). | |||||
|
WWC3_HUMAN
|
||||||
| NC score | 0.016227 (rank : 59) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
EHMT2_MOUSE
|
||||||
| NC score | 0.016162 (rank : 60) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.015153 (rank : 61) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
CCL21_HUMAN
|
||||||
| NC score | 0.014989 (rank : 62) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00585 | Gene names | CCL21, SCYA21 | |||
|
Domain Architecture |

|
|||||
| Description | Small inducible cytokine A21 precursor (CCL21) (Beta chemokine exodus- 2) (6Ckine) (Secondary lymphoid-tissue chemokine) (SLC). | |||||
|
EHMT2_HUMAN
|
||||||
| NC score | 0.014924 (rank : 63) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96KQ7, Q14349, Q6PK06, Q96MH5, Q96QD0, Q9UQL8, Q9Y331 | Gene names | EHMT2, BAT8, G9A, NG36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
PAR3L_HUMAN
|
||||||
| NC score | 0.014812 (rank : 64) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8TEW8, Q8IUC7, Q8IUC9, Q96DK9, Q96N09, Q96NX6, Q96NX7, Q96Q29 | Gene names | PARD3B, ALS2CR19, PAR3B, PAR3L | |||
|
Domain Architecture |

|
|||||
| Description | Partitioning-defective 3 homolog B (PAR3-beta) (Partitioning-defective 3-like protein) (PAR3-L protein) (Amyotrophic lateral sclerosis 2 chromosome region candidate gene 19 protein). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.014274 (rank : 65) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
RN146_HUMAN
|
||||||
| NC score | 0.014090 (rank : 66) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NTX7, Q6FIB2, Q7L8H4, Q96K03, Q9NTX6 | Gene names | RNF146 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 146 (Dactylidin). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.013577 (rank : 67) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
MTO1_MOUSE
|
||||||
| NC score | 0.013292 (rank : 68) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q923Z3, Q9D2Q5 | Gene names | Mto1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MTO1 homolog, mitochondrial precursor. | |||||
|
INCE_HUMAN
|
||||||
| NC score | 0.011246 (rank : 69) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
NKX12_MOUSE
|
||||||
| NC score | 0.011192 (rank : 70) | θ value | 1.06291 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42580, Q9JHF2, Q9JKW5, Q9JKW6, Q9JKW9, Q9JKX0, Q9JKX1 | Gene names | Nkx1-2, Nkx1.1, Sax1 | |||
|
Domain Architecture |

|
|||||
| Description | NK1 transcription factor-related protein 2 (Homeobox protein SAX-1) (NKX-1.1). | |||||
|
ITSN1_HUMAN
|
||||||
| NC score | 0.010453 (rank : 71) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
DDX23_HUMAN
|
||||||
| NC score | 0.008914 (rank : 72) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9BUQ8, O43188 | Gene names | DDX23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX23 (EC 3.6.1.-) (DEAD box protein 23) (100 kDa U5 snRNP-specific protein) (U5-100kD) (PRP28 homolog). | |||||
|
PGCA_HUMAN
|
||||||
| NC score | 0.007951 (rank : 73) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
TNIK_HUMAN
|
||||||
| NC score | 0.007896 (rank : 74) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
STK10_HUMAN
|
||||||
| NC score | 0.007497 (rank : 75) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O94804, Q9UIW4 | Gene names | STK10, LOK | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase 10 (EC 2.7.11.1) (Lymphocyte-oriented kinase). | |||||
|
M4K4_HUMAN
|
||||||
| NC score | 0.007462 (rank : 76) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1589 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O95819, O75172, Q9NST7 | Gene names | MAP4K4, HGK, KIAA0687, NIK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 4 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 4) (MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase HGK) (Nck-interacting kinase). | |||||
|
ELF4_MOUSE
|
||||||
| NC score | 0.007413 (rank : 77) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
WNK1_HUMAN
|
||||||
| NC score | 0.007346 (rank : 78) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1158 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H4A3, O15052, Q86WL5, Q8N673, Q9P1S9 | Gene names | WNK1, KDP, KIAA0344, PRKWNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1) (Kinase deficient protein). | |||||
|
CAH9_HUMAN
|
||||||
| NC score | 0.007263 (rank : 79) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16790 | Gene names | CA9, G250, MN | |||
|
Domain Architecture |

|
|||||
| Description | Carbonic anhydrase 9 precursor (EC 4.2.1.1) (Carbonic anhydrase IX) (Carbonate dehydratase IX) (CA-IX) (CAIX) (Membrane antigen MN) (P54/58N) (Renal cell carcinoma-associated antigen G250) (RCC- associated antigen G250) (pMW1). | |||||
|
CTRO_MOUSE
|
||||||
| NC score | 0.007069 (rank : 80) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
E41L1_MOUSE
|
||||||
| NC score | 0.006835 (rank : 81) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2H5 | Gene names | Epb41l1, Epb4, Epb4.1l1 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 1 (Neuronal protein 4.1) (4.1N). | |||||
|
S6A11_MOUSE
|
||||||
| NC score | 0.006558 (rank : 82) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P31650 | Gene names | Slc6a11, Gabt4, Gat-4, Gat4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium- and chloride-dependent GABA transporter 4 (GAT4). | |||||
|
KBTBA_HUMAN
|
||||||
| NC score | 0.006548 (rank : 83) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60662 | Gene names | KBTBD10, KRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch repeat and BTB domain-containing protein 10 (Kelch-related protein 1) (Kel-like protein 23) (Sarcosin). | |||||
|
MEFV_HUMAN
|
||||||
| NC score | 0.005539 (rank : 84) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O15553, Q96PN4, Q96PN5 | Gene names | MEFV, MEF | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
CTRO_HUMAN
|
||||||
| NC score | 0.005484 (rank : 85) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
ZBT37_HUMAN
|
||||||
| NC score | 0.005462 (rank : 86) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 815 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5TC79, Q5TC80, Q96M87, Q9BQ88 | Gene names | ZBTB37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 37. | |||||
|
MYO6_MOUSE
|
||||||
| NC score | 0.005207 (rank : 87) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q64331 | Gene names | Myo6, Sv | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
LTB1L_HUMAN
|
||||||
| NC score | 0.004701 (rank : 88) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14766 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1L precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
LTB1S_HUMAN
|
||||||
| NC score | 0.004333 (rank : 89) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P22064, Q8TD95 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1S precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
MASTL_HUMAN
|
||||||
| NC score | 0.003743 (rank : 90) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96GX5, Q5T8D5, Q5T8D7, Q8NCD6, Q96SJ5 | Gene names | MASTL, THC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase-like (EC 2.7.11.1). | |||||
|
ZFP91_HUMAN
|
||||||
| NC score | 0.003245 (rank : 91) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 769 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96JP5, Q86V47, Q96JP4, Q96QA3 | Gene names | ZFP91 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 91 homolog (Zfp-91). | |||||
|
ZFP91_MOUSE
|
||||||
| NC score | 0.002576 (rank : 92) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62511, Q8BPY3, Q8C2B4 | Gene names | Zfp91, Pzf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 91 (Zfp-91) (Zinc finger protein PZF) (Penta Zf protein). | |||||