Please be patient as the page loads
|
SPAG8_HUMAN
|
||||||
| SwissProt Accessions | Q99932, Q12937, Q5TCV8, Q8WWB4 | Gene names | SPAG8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 8 (Sperm membrane protein 1) (SMP-1) (HSD-1) (Sperm membrane protein BS-84). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SPAG8_HUMAN
|
||||||
| θ value | 1.86219e-138 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q99932, Q12937, Q5TCV8, Q8WWB4 | Gene names | SPAG8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 8 (Sperm membrane protein 1) (SMP-1) (HSD-1) (Sperm membrane protein BS-84). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 2) | NC score | 0.095442 (rank : 12) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 3) | NC score | 0.086611 (rank : 19) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 4) | NC score | 0.114212 (rank : 5) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
TNC6B_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 5) | NC score | 0.131767 (rank : 4) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
GP112_HUMAN
|
||||||
| θ value | 0.125558 (rank : 6) | NC score | 0.053077 (rank : 111) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IZF6, Q86SM6 | Gene names | GPR112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 112. | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.125558 (rank : 7) | NC score | 0.054989 (rank : 102) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 8) | NC score | 0.151349 (rank : 3) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
LPP_MOUSE
|
||||||
| θ value | 0.21417 (rank : 9) | NC score | 0.057075 (rank : 84) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BFW7, Q5U407, Q8BHI1, Q8BKI0, Q8BKN2, Q8BLF4, Q8BLG3, Q8C101 | Gene names | Lpp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner homolog. | |||||
|
SPRR3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 10) | NC score | 0.106000 (rank : 9) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UBC9, O75597, Q4ZGI7, Q5T525, Q8NET7, Q9UDG3 | Gene names | SPRR3, SPRC | |||
|
Domain Architecture |
|
|||||
| Description | Small proline-rich protein 3 (Cornifin beta) (Esophagin) (22 kDa pancornulin). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 11) | NC score | 0.152280 (rank : 2) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 0.279714 (rank : 12) | NC score | 0.095167 (rank : 13) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 0.279714 (rank : 13) | NC score | 0.088558 (rank : 18) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 0.365318 (rank : 14) | NC score | 0.063309 (rank : 61) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
B4GT1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 15) | NC score | 0.041532 (rank : 130) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
EMIL2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 16) | NC score | 0.053484 (rank : 109) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
U383_MOUSE
|
||||||
| θ value | 0.47712 (rank : 17) | NC score | 0.081372 (rank : 23) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D2Q2, Q3V1K3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative methyltransferase UPF0383 (EC 2.1.1.-). | |||||
|
ZN385_HUMAN
|
||||||
| θ value | 0.47712 (rank : 18) | NC score | 0.070885 (rank : 39) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
RGS12_HUMAN
|
||||||
| θ value | 0.62314 (rank : 19) | NC score | 0.057060 (rank : 85) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
SF3B1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 20) | NC score | 0.104689 (rank : 11) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75533 | Gene names | SF3B1, SAP155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
SF3B1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 21) | NC score | 0.105893 (rank : 10) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99NB9, Q9CSK5 | Gene names | Sf3b1, Sap155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
ZIMP7_MOUSE
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.056013 (rank : 94) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8CIE2, Q6P558, Q6P9Q5, Q8BJ00 | Gene names | Zimp7, D11Bwg0280e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
ZN646_HUMAN
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.010902 (rank : 142) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15015 | Gene names | ZNF646, KIAA0296 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 646. | |||||
|
ARMX2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.068208 (rank : 47) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6A058, Q9CXI9 | Gene names | Armcx2, Kiaa0512 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 2. | |||||
|
CBX2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.059068 (rank : 76) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14781, Q9BTB1 | Gene names | CBX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromobox protein homolog 2. | |||||
|
CO4A2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.046489 (rank : 127) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P08122, Q61375 | Gene names | Col4a2 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
JIP2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.060697 (rank : 68) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13387, Q96G62, Q99771, Q9NZ59, Q9UKQ4 | Gene names | MAPK8IP2, IB2, JIP2 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 2 (JNK-interacting protein 2) (JIP-2) (JNK MAP kinase scaffold protein 2) (Islet-brain-2) (IB-2) (Mitogen-activated protein kinase 8-interacting protein 2). | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.013367 (rank : 140) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
GR65_MOUSE
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.049291 (rank : 125) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91X51, Q9D3L9 | Gene names | Gorasp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65). | |||||
|
TAF1C_MOUSE
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.048629 (rank : 126) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
CO5A2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.058889 (rank : 77) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
MAML1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.058740 (rank : 79) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
COBA2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 33) | NC score | 0.054315 (rank : 104) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
PGCA_MOUSE
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.045822 (rank : 128) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
SNPH_HUMAN
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.034124 (rank : 133) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15079 | Gene names | SNPH, KIAA0374 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaphilin. | |||||
|
ZN358_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.007082 (rank : 145) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NW07, Q9BTM7 | Gene names | ZNF358 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 358. | |||||
|
CRSP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.055919 (rank : 95) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
CSPG3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.027330 (rank : 137) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P55066, Q6P1E3, Q8C4F8 | Gene names | Cspg3, Ncan | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
FILA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.029873 (rank : 136) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
JHD2B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.021526 (rank : 138) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZPY7, Q2VPQ5, Q5U5V7, Q6P9K3, Q8CCE2, Q8K2A5, Q9CU57 | Gene names | Jmjd1b, Jhdm2b, Kiaa1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B). | |||||
|
NPL4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.032150 (rank : 135) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TAT6, Q8N3J1, Q9H8V2, Q9H964, Q9NWR5, Q9P229 | Gene names | NPLOC4, KIAA1499, NPL4 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear protein localization protein 4 homolog (Protein NPL4). | |||||
|
ABL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.005360 (rank : 146) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P00519, Q13869, Q13870, Q16133, Q45F09 | Gene names | ABL1, ABL, JTK7 | |||
|
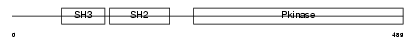
Domain Architecture |
|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||
|
BAT3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.074362 (rank : 32) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46379, O95874, Q5SQ35, Q5SQ36, Q5SQ37, Q5SRP8, Q96SA6, Q9BCN4 | Gene names | BAT3, G3 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3) (Protein G3). | |||||
|
BAT3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.079092 (rank : 25) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
CCD71_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.038766 (rank : 131) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IV32, Q6IPE2, Q9H8H4, Q9H9F1 | Gene names | CCDC71 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 71. | |||||
|
COFA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.066844 (rank : 51) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P39059, Q5T6J4, Q9Y4W4 | Gene names | COL15A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV) (Restin) (Related to endostatin)]. | |||||
|
CSKI2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.036914 (rank : 132) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WXE0, Q7LG69, Q9ULT1 | Gene names | CASKIN2, KIAA1139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-2. | |||||
|
ENAM_HUMAN
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.045529 (rank : 129) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NRM1, Q9H3D1 | Gene names | ENAM | |||
|
Domain Architecture |

|
|||||
| Description | Enamelin precursor. | |||||
|
NU214_HUMAN
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.085084 (rank : 22) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
PGCA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.053281 (rank : 110) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
TNR6A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.078392 (rank : 27) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
CRK_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.012405 (rank : 141) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64010 | Gene names | Crk, Crko | |||
|
Domain Architecture |
|
|||||
| Description | Proto-oncogene C-crk (P38) (Adapter molecule crk). | |||||
|
JIP3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.017191 (rank : 139) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
RX_MOUSE
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.010440 (rank : 143) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35602, O08748 | Gene names | Rax, Rx | |||
|
Domain Architecture |

|
|||||
| Description | Retinal homeobox protein Rx (Retina and anterior neural fold homeobox protein). | |||||
|
SYNPO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.032367 (rank : 134) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N3V7, O15271, Q9UPX1 | Gene names | SYNPO, KIAA1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
TRIM8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.010376 (rank : 144) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BZR9, Q9C028 | Gene names | TRIM8, GERP, RNF27 | |||
|
Domain Architecture |
|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
AP180_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.052880 (rank : 112) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61548, Q61547 | Gene names | Snap91 | |||
|
Domain Architecture |
|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein) (Phosphoprotein F1- 20). | |||||
|
ARI1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.066256 (rank : 53) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.067555 (rank : 49) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.068768 (rank : 46) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.079664 (rank : 24) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.067594 (rank : 48) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BSN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.051614 (rank : 119) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CDN1C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.069452 (rank : 41) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
CIC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.054145 (rank : 106) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
CK024_HUMAN
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.073028 (rank : 33) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96F05, Q9H2K4 | Gene names | C11orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 precursor (Protein DM4E3). | |||||
|
CN032_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.074595 (rank : 31) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
CN032_MOUSE
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.085331 (rank : 21) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
CN155_HUMAN
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.077077 (rank : 30) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
CO002_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.051252 (rank : 120) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
CO1A1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.065101 (rank : 56) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P02452, P78441, Q13896, Q13902, Q13903, Q14037, Q14992, Q15176, Q15201, Q16050, Q7KZ30, Q7KZ34, Q8IVI5, Q9UML6, Q9UMM7 | Gene names | COL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
CO1A1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.066156 (rank : 54) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P11087, Q53WT0, Q60635, Q61367, Q61427, Q63919, Q6PCL3, Q810J9 | Gene names | Col1a1, Cola1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
CO1A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.055613 (rank : 97) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q01149, Q8CGA5 | Gene names | Col1a2, Cola2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
CO3A1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.069056 (rank : 43) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
CO3A1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.069194 (rank : 42) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P08121, Q61429, Q9CRN7 | Gene names | Col3a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
CO4A1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.052042 (rank : 116) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CO4A4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.053743 (rank : 108) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |
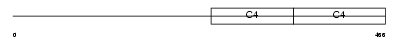
|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
CO4A5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.071160 (rank : 38) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P29400, Q16006, Q16126, Q6LD84 | Gene names | COL4A5 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-5(IV) chain precursor. | |||||
|
CO8A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.056576 (rank : 87) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P25067, Q8TEJ5 | Gene names | COL8A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
CO8A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.054540 (rank : 103) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P25318, Q68ED0, Q6KAQ4, Q6P1C4 | Gene names | Col8a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
COAA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.058875 (rank : 78) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
COBA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.051031 (rank : 121) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
COFA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.055304 (rank : 98) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35206, Q9EQD9 | Gene names | Col15a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV)]. | |||||
|
COGA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.059324 (rank : 74) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q07092 | Gene names | COL16A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVI) chain precursor. | |||||
|
COHA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.059902 (rank : 72) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UMD9, Q02802, Q99018, Q9NQK9, Q9UC14 | Gene names | COL17A1, BP180, BPAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
COHA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.057237 (rank : 83) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q07563, Q99LK8 | Gene names | Col17a1, Bp180, Bpag2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
COIA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.066848 (rank : 50) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P39060, Q9UK38, Q9Y6Q7, Q9Y6Q8 | Gene names | COL18A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
COIA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.056290 (rank : 93) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
CPSF6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.062812 (rank : 64) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
CPSF6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.063954 (rank : 59) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
ELN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.111945 (rank : 7) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P15502, O15336, O15337, Q14233, Q14234, Q14235, Q14238, Q6P0L4, Q6ZWJ6, Q75MU5, Q7Z316, Q7Z3F5, Q9UMF5 | Gene names | ELN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elastin precursor (Tropoelastin). | |||||
|
ELN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.056499 (rank : 92) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P54320 | Gene names | Eln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elastin precursor (Tropoelastin). | |||||
|
EMID2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.056507 (rank : 90) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96A83 | Gene names | EMID2, COL26A1, EMU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
ENAH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.052810 (rank : 113) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
FMN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.055859 (rank : 96) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
GGN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.072441 (rank : 36) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86UU5, Q7RTU6, Q86UU4, Q8NAA1 | Gene names | GGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
GGN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.062538 (rank : 65) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.057368 (rank : 82) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
MARCS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.077754 (rank : 28) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
MARCS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.066626 (rank : 52) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
MBD6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.072729 (rank : 35) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
MN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.056563 (rank : 88) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
MUC13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.061753 (rank : 66) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.090001 (rank : 17) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
MUC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.066015 (rank : 55) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.090152 (rank : 15) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
MUCDL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.054166 (rank : 105) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HBB8, Q9H746, Q9HAU3, Q9HBB5, Q9HBB6, Q9HBB7, Q9NX86, Q9NXI9 | Gene names | MUCDHL | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.057635 (rank : 81) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
NUP62_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.052016 (rank : 117) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63850, Q99JN7 | Gene names | Nup62 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore glycoprotein p62 (62 kDa nucleoporin). | |||||
|
PELP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.050196 (rank : 124) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
PO121_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.065015 (rank : 57) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
PO121_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.063655 (rank : 60) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
PP1RA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.050394 (rank : 123) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96QC0, O00405 | Gene names | PPP1R10, CAT53, FB19, PNUTS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein phosphatase 1 regulatory subunit 10 (Phosphatase 1 nuclear targeting subunit) (MHC class I region proline- rich protein CAT53) (FB19 protein) (PP1-binding protein of 114 kDa) (p99). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.108083 (rank : 8) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.095015 (rank : 14) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
PRG4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.068947 (rank : 44) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRG4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.052400 (rank : 114) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.112802 (rank : 6) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
PRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.090121 (rank : 16) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRP5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.064453 (rank : 58) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P04281 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide IB-1. | |||||
|
PRPC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.055023 (rank : 101) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P02810, Q4VBP2, Q53XA2, Q6P2F6 | Gene names | PRH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Salivary acidic proline-rich phosphoprotein 1/2 precursor (PRP-1/PRP- 2) (Parotid proline-rich protein 1/2) (Pr1/Pr2) (Protein C) (Parotid acidic protein) (Pa) (Parotid isoelectric focusing variant protein) (PIF-S) (Parotid double-band protein) (Db-s) [Contains: Salivary acidic proline-rich phosphoprotein 1/2; Salivary acidic proline-rich phosphoprotein 3/4 (PRP-3/PRP-4) (Protein A) (PIF-F) (Db-F); Peptide P-C]. | |||||
|
PRPE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.068939 (rank : 45) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P02811 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide P-E (IB-9). | |||||
|
PRR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.063087 (rank : 62) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.052103 (rank : 115) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.055182 (rank : 99) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SELV_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.069549 (rank : 40) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P59797 | Gene names | SELV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Selenoprotein V. | |||||
|
SF3A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.072884 (rank : 34) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |
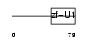
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SF3A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.078560 (rank : 26) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |
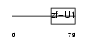
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SFPQ_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.056851 (rank : 86) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
SFPQ_MOUSE
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.060420 (rank : 71) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
SPRR3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.060648 (rank : 69) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O09116 | Gene names | Sprr3 | |||
|
Domain Architecture |

|
|||||
| Description | Small proline-rich protein 3 (Cornifin beta). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.057661 (rank : 80) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.059645 (rank : 73) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.054021 (rank : 107) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SYN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.056502 (rank : 91) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
SYN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.052003 (rank : 118) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |
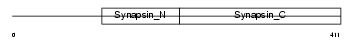
|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
T22D2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.055075 (rank : 100) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.085475 (rank : 20) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
TCOF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.062826 (rank : 63) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.077649 (rank : 29) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
WASIP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.071796 (rank : 37) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
WBP11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.060504 (rank : 70) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
WBP11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.056536 (rank : 89) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q923D5, O88539, Q3UMD8, Q8VDI0 | Gene names | Wbp11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11). | |||||
|
WIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.059100 (rank : 75) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
WIRE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.051028 (rank : 122) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
ZN207_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.061170 (rank : 67) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
SPAG8_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.86219e-138 (rank : 1) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q99932, Q12937, Q5TCV8, Q8WWB4 | Gene names | SPAG8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 8 (Sperm membrane protein 1) (SMP-1) (HSD-1) (Sperm membrane protein BS-84). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.152280 (rank : 2) | θ value | 0.21417 (rank : 11) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.151349 (rank : 3) | θ value | 0.163984 (rank : 8) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
TNC6B_HUMAN
|
||||||
| NC score | 0.131767 (rank : 4) | θ value | 0.0563607 (rank : 5) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.114212 (rank : 5) | θ value | 0.0193708 (rank : 4) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.112802 (rank : 6) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
ELN_HUMAN
|
||||||
| NC score | 0.111945 (rank : 7) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P15502, O15336, O15337, Q14233, Q14234, Q14235, Q14238, Q6P0L4, Q6ZWJ6, Q75MU5, Q7Z316, Q7Z3F5, Q9UMF5 | Gene names | ELN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elastin precursor (Tropoelastin). | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.108083 (rank : 8) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
SPRR3_HUMAN
|
||||||
| NC score | 0.106000 (rank : 9) | θ value | 0.21417 (rank : 10) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UBC9, O75597, Q4ZGI7, Q5T525, Q8NET7, Q9UDG3 | Gene names | SPRR3, SPRC | |||
|
Domain Architecture |

|
|||||
| Description | Small proline-rich protein 3 (Cornifin beta) (Esophagin) (22 kDa pancornulin). | |||||
|
SF3B1_MOUSE
|
||||||
| NC score | 0.105893 (rank : 10) | θ value | 1.06291 (rank : 21) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99NB9, Q9CSK5 | Gene names | Sf3b1, Sap155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
SF3B1_HUMAN
|
||||||
| NC score | 0.104689 (rank : 11) | θ value | 1.06291 (rank : 20) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75533 | Gene names | SF3B1, SAP155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.095442 (rank : 12) | θ value | 0.00228821 (rank : 2) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.095167 (rank : 13) | θ value | 0.279714 (rank : 12) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.095015 (rank : 14) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.090152 (rank : 15) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.090121 (rank : 16) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.090001 (rank : 17) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.088558 (rank : 18) | θ value | 0.279714 (rank : 13) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.086611 (rank : 19) | θ value | 0.0113563 (rank : 3) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.085475 (rank : 20) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
CN032_MOUSE
|
||||||
| NC score | 0.085331 (rank : 21) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
NU214_HUMAN
|
||||||
| NC score | 0.085084 (rank : 22) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
U383_MOUSE
|
||||||
| NC score | 0.081372 (rank : 23) | θ value | 0.47712 (rank : 17) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D2Q2, Q3V1K3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative methyltransferase UPF0383 (EC 2.1.1.-). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.079664 (rank : 24) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BAT3_MOUSE
|
||||||
| NC score | 0.079092 (rank : 25) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
SF3A2_MOUSE
|
||||||
| NC score | 0.078560 (rank : 26) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
TNR6A_HUMAN
|
||||||
| NC score | 0.078392 (rank : 27) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
MARCS_HUMAN
|
||||||
| NC score | 0.077754 (rank : 28) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.077649 (rank : 29) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.077077 (rank : 30) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
CN032_HUMAN
|
||||||
| NC score | 0.074595 (rank : 31) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
BAT3_HUMAN
|
||||||
| NC score | 0.074362 (rank : 32) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46379, O95874, Q5SQ35, Q5SQ36, Q5SQ37, Q5SRP8, Q96SA6, Q9BCN4 | Gene names | BAT3, G3 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3) (Protein G3). | |||||
|
CK024_HUMAN
|
||||||
| NC score | 0.073028 (rank : 33) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96F05, Q9H2K4 | Gene names | C11orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 precursor (Protein DM4E3). | |||||
|
SF3A2_HUMAN
|
||||||
| NC score | 0.072884 (rank : 34) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
MBD6_HUMAN
|
||||||
| NC score | 0.072729 (rank : 35) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
GGN_HUMAN
|
||||||
| NC score | 0.072441 (rank : 36) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86UU5, Q7RTU6, Q86UU4, Q8NAA1 | Gene names | GGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.071796 (rank : 37) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
CO4A5_HUMAN
|
||||||
| NC score | 0.071160 (rank : 38) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P29400, Q16006, Q16126, Q6LD84 | Gene names | COL4A5 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-5(IV) chain precursor. | |||||
|
ZN385_HUMAN
|
||||||
| NC score | 0.070885 (rank : 39) | θ value | 0.47712 (rank : 18) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
SELV_HUMAN
|
||||||
| NC score | 0.069549 (rank : 40) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P59797 | Gene names | SELV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Selenoprotein V. | |||||
|
CDN1C_HUMAN
|
||||||
| NC score | 0.069452 (rank : 41) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
CO3A1_MOUSE
|
||||||
| NC score | 0.069194 (rank : 42) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P08121, Q61429, Q9CRN7 | Gene names | Col3a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
CO3A1_HUMAN
|
||||||
| NC score | 0.069056 (rank : 43) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.068947 (rank : 44) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRPE_HUMAN
|
||||||
| NC score | 0.068939 (rank : 45) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P02811 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide P-E (IB-9). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.068768 (rank : 46) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
ARMX2_MOUSE
|
||||||
| NC score | 0.068208 (rank : 47) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6A058, Q9CXI9 | Gene names | Armcx2, Kiaa0512 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 2. | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.067594 (rank : 48) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.067555 (rank : 49) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
COIA1_HUMAN
|
||||||
| NC score | 0.066848 (rank : 50) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P39060, Q9UK38, Q9Y6Q7, Q9Y6Q8 | Gene names | COL18A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
COFA1_HUMAN
|
||||||
| NC score | 0.066844 (rank : 51) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P39059, Q5T6J4, Q9Y4W4 | Gene names | COL15A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV) (Restin) (Related to endostatin)]. | |||||
|
MARCS_MOUSE
|
||||||
| NC score | 0.066626 (rank : 52) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
ARI1B_HUMAN
|
||||||
| NC score | 0.066256 (rank : 53) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
CO1A1_MOUSE
|
||||||
| NC score | 0.066156 (rank : 54) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P11087, Q53WT0, Q60635, Q61367, Q61427, Q63919, Q6PCL3, Q810J9 | Gene names | Col1a1, Cola1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
MUC2_HUMAN
|
||||||
| NC score | 0.066015 (rank : 55) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
CO1A1_HUMAN
|
||||||
| NC score | 0.065101 (rank : 56) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P02452, P78441, Q13896, Q13902, Q13903, Q14037, Q14992, Q15176, Q15201, Q16050, Q7KZ30, Q7KZ34, Q8IVI5, Q9UML6, Q9UMM7 | Gene names | COL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
PO121_HUMAN
|
||||||
| NC score | 0.065015 (rank : 57) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
PRP5_HUMAN
|
||||||
| NC score | 0.064453 (rank : 58) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P04281 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide IB-1. | |||||
|
CPSF6_MOUSE
|
||||||
| NC score | 0.063954 (rank : 59) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
PO121_MOUSE
|
||||||
| NC score | 0.063655 (rank : 60) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.063309 (rank : 61) | θ value | 0.365318 (rank : 14) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.063087 (rank : 62) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
TCOF_MOUSE
|
||||||
| NC score | 0.062826 (rank : 63) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
CPSF6_HUMAN
|
||||||
| NC score | 0.062812 (rank : 64) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
GGN_MOUSE
|
||||||
| NC score | 0.062538 (rank : 65) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
MUC13_MOUSE
|
||||||
| NC score | 0.061753 (rank : 66) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
ZN207_HUMAN
|
||||||
| NC score | 0.061170 (rank : 67) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
JIP2_HUMAN
|
||||||
| NC score | 0.060697 (rank : 68) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13387, Q96G62, Q99771, Q9NZ59, Q9UKQ4 | Gene names | MAPK8IP2, IB2, JIP2 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 2 (JNK-interacting protein 2) (JIP-2) (JNK MAP kinase scaffold protein 2) (Islet-brain-2) (IB-2) (Mitogen-activated protein kinase 8-interacting protein 2). | |||||
|
SPRR3_MOUSE
|
||||||
| NC score | 0.060648 (rank : 69) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O09116 | Gene names | Sprr3 | |||
|
Domain Architecture |

|
|||||
| Description | Small proline-rich protein 3 (Cornifin beta). | |||||
|
WBP11_HUMAN
|
||||||
| NC score | 0.060504 (rank : 70) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
SFPQ_MOUSE
|
||||||
| NC score | 0.060420 (rank : 71) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
COHA1_HUMAN
|
||||||
| NC score | 0.059902 (rank : 72) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UMD9, Q02802, Q99018, Q9NQK9, Q9UC14 | Gene names | COL17A1, BP180, BPAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.059645 (rank : 73) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
COGA1_HUMAN
|
||||||
| NC score | 0.059324 (rank : 74) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q07092 | Gene names | COL16A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVI) chain precursor. | |||||
|
WIRE_HUMAN
|
||||||
| NC score | 0.059100 (rank : 75) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
CBX2_HUMAN
|
||||||
| NC score | 0.059068 (rank : 76) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14781, Q9BTB1 | Gene names | CBX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromobox protein homolog 2. | |||||
|
CO5A2_HUMAN
|
||||||
| NC score | 0.058889 (rank : 77) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
COAA1_HUMAN
|
||||||
| NC score | 0.058875 (rank : 78) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
MAML1_MOUSE
|
||||||
| NC score | 0.058740 (rank : 79) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.057661 (rank : 80) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.057635 (rank : 81) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.057368 (rank : 82) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
COHA1_MOUSE
|
||||||
| NC score | 0.057237 (rank : 83) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q07563, Q99LK8 | Gene names | Col17a1, Bp180, Bpag2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
LPP_MOUSE
|
||||||
| NC score | 0.057075 (rank : 84) | θ value | 0.21417 (rank : 9) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BFW7, Q5U407, Q8BHI1, Q8BKI0, Q8BKN2, Q8BLF4, Q8BLG3, Q8C101 | Gene names | Lpp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner homolog. | |||||
|
RGS12_HUMAN
|
||||||
| NC score | 0.057060 (rank : 85) | θ value | 0.62314 (rank : 19) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
SFPQ_HUMAN
|
||||||
| NC score | 0.056851 (rank : 86) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
CO8A2_HUMAN
|
||||||
| NC score | 0.056576 (rank : 87) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P25067, Q8TEJ5 | Gene names | COL8A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
MN1_HUMAN
|
||||||
| NC score | 0.056563 (rank : 88) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
WBP11_MOUSE
|
||||||
| NC score | 0.056536 (rank : 89) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q923D5, O88539, Q3UMD8, Q8VDI0 | Gene names | Wbp11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11). | |||||
|
EMID2_HUMAN
|
||||||
| NC score | 0.056507 (rank : 90) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96A83 | Gene names | EMID2, COL26A1, EMU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
SYN1_HUMAN
|
||||||
| NC score | 0.056502 (rank : 91) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
ELN_MOUSE
|
||||||
| NC score | 0.056499 (rank : 92) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P54320 | Gene names | Eln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elastin precursor (Tropoelastin). | |||||
|
COIA1_MOUSE
|
||||||
| NC score | 0.056290 (rank : 93) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
ZIMP7_MOUSE
|
||||||
| NC score | 0.056013 (rank : 94) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8CIE2, Q6P558, Q6P9Q5, Q8BJ00 | Gene names | Zimp7, D11Bwg0280e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
CRSP2_HUMAN
|
||||||
| NC score | 0.055919 (rank : 95) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.055859 (rank : 96) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
CO1A2_MOUSE
|
||||||
| NC score | 0.055613 (rank : 97) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q01149, Q8CGA5 | Gene names | Col1a2, Cola2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
COFA1_MOUSE
|
||||||
| NC score | 0.055304 (rank : 98) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35206, Q9EQD9 | Gene names | Col15a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XV) chain precursor [Contains: Endostatin (Endostatin-XV)]. | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.055182 (rank : 99) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
T22D2_HUMAN
|
||||||
| NC score | 0.055075 (rank : 100) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
PRPC_HUMAN
|
||||||
| NC score | 0.055023 (rank : 101) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P02810, Q4VBP2, Q53XA2, Q6P2F6 | Gene names | PRH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Salivary acidic proline-rich phosphoprotein 1/2 precursor (PRP-1/PRP- 2) (Parotid proline-rich protein 1/2) (Pr1/Pr2) (Protein C) (Parotid acidic protein) (Pa) (Parotid isoelectric focusing variant protein) (PIF-S) (Parotid double-band protein) (Db-s) [Contains: Salivary acidic proline-rich phosphoprotein 1/2; Salivary acidic proline-rich phosphoprotein 3/4 (PRP-3/PRP-4) (Protein A) (PIF-F) (Db-F); Peptide P-C]. | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.054989 (rank : 102) | θ value | 0.125558 (rank : 7) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
CO8A2_MOUSE
|
||||||
| NC score | 0.054540 (rank : 103) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P25318, Q68ED0, Q6KAQ4, Q6P1C4 | Gene names | Col8a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
COBA2_MOUSE
|
||||||
| NC score | 0.054315 (rank : 104) | θ value | 4.03905 (rank : 33) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
MUCDL_HUMAN
|
||||||
| NC score | 0.054166 (rank : 105) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HBB8, Q9H746, Q9HAU3, Q9HBB5, Q9HBB6, Q9HBB7, Q9NX86, Q9NXI9 | Gene names | MUCDHL | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
CIC_MOUSE
|
||||||
| NC score | 0.054145 (rank : 106) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.054021 (rank : 107) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
CO4A4_HUMAN
|
||||||
| NC score | 0.053743 (rank : 108) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
EMIL2_MOUSE
|
||||||
| NC score | 0.053484 (rank : 109) | θ value | 0.47712 (rank : 16) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
PGCA_HUMAN
|
||||||
| NC score | 0.053281 (rank : 110) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
GP112_HUMAN
|
||||||
| NC score | 0.053077 (rank : 111) | θ value | 0.125558 (rank : 6) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IZF6, Q86SM6 | Gene names | GPR112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 112. | |||||
|
AP180_MOUSE
|
||||||
| NC score | 0.052880 (rank : 112) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61548, Q61547 | Gene names | Snap91 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin coat assembly protein AP180 (Clathrin coat-associated protein AP180) (91 kDa synaptosomal-associated protein) (Phosphoprotein F1- 20). | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.052810 (rank : 113) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.052400 (rank : 114) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.052103 (rank : 115) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
CO4A1_MOUSE
|
||||||
| NC score | 0.052042 (rank : 116) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
NUP62_MOUSE
|
||||||
| NC score | 0.052016 (rank : 117) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63850, Q99JN7 | Gene names | Nup62 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore glycoprotein p62 (62 kDa nucleoporin). | |||||
|
SYN1_MOUSE
|
||||||
| NC score | 0.052003 (rank : 118) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.051614 (rank : 119) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CO002_HUMAN
|
||||||
| NC score | 0.051252 (rank : 120) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
COBA2_HUMAN
|
||||||
| NC score | 0.051031 (rank : 121) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
WIRE_MOUSE
|
||||||
| NC score | 0.051028 (rank : 122) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
PP1RA_HUMAN
|
||||||
| NC score | 0.050394 (rank : 123) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96QC0, O00405 | Gene names | PPP1R10, CAT53, FB19, PNUTS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein phosphatase 1 regulatory subunit 10 (Phosphatase 1 nuclear targeting subunit) (MHC class I region proline- rich protein CAT53) (FB19 protein) (PP1-binding protein of 114 kDa) (p99). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.050196 (rank : 124) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
GR65_MOUSE
|
||||||
| NC score | 0.049291 (rank : 125) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91X51, Q9D3L9 | Gene names | Gorasp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65). | |||||
|
TAF1C_MOUSE
|
||||||
| NC score | 0.048629 (rank : 126) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
CO4A2_MOUSE
|
||||||
| NC score | 0.046489 (rank : 127) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P08122, Q61375 | Gene names | Col4a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
PGCA_MOUSE
|
||||||
| NC score | 0.045822 (rank : 128) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
ENAM_HUMAN
|
||||||
| NC score | 0.045529 (rank : 129) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NRM1, Q9H3D1 | Gene names | ENAM | |||
|
Domain Architecture |

|
|||||
| Description | Enamelin precursor. | |||||
|
B4GT1_HUMAN
|
||||||
| NC score | 0.041532 (rank : 130) | θ value | 0.47712 (rank : 15) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
CCD71_HUMAN
|
||||||
| NC score | 0.038766 (rank : 131) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IV32, Q6IPE2, Q9H8H4, Q9H9F1 | Gene names | CCDC71 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 71. | |||||
|
CSKI2_HUMAN
|
||||||
| NC score | 0.036914 (rank : 132) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WXE0, Q7LG69, Q9ULT1 | Gene names | CASKIN2, KIAA1139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-2. | |||||
|
SNPH_HUMAN
|
||||||
| NC score | 0.034124 (rank : 133) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15079 | Gene names | SNPH, KIAA0374 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaphilin. | |||||
|
SYNPO_HUMAN
|
||||||
| NC score | 0.032367 (rank : 134) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N3V7, O15271, Q9UPX1 | Gene names | SYNPO, KIAA1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
NPL4_HUMAN
|
||||||
| NC score | 0.032150 (rank : 135) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TAT6, Q8N3J1, Q9H8V2, Q9H964, Q9NWR5, Q9P229 | Gene names | NPLOC4, KIAA1499, NPL4 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear protein localization protein 4 homolog (Protein NPL4). | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.029873 (rank : 136) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
CSPG3_MOUSE
|
||||||
| NC score | 0.027330 (rank : 137) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P55066, Q6P1E3, Q8C4F8 | Gene names | Cspg3, Ncan | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
JHD2B_MOUSE
|
||||||
| NC score | 0.021526 (rank : 138) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZPY7, Q2VPQ5, Q5U5V7, Q6P9K3, Q8CCE2, Q8K2A5, Q9CU57 | Gene names | Jmjd1b, Jhdm2b, Kiaa1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B). | |||||
|
JIP3_HUMAN
|
||||||
| NC score | 0.017191 (rank : 139) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.013367 (rank : 140) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
CRK_MOUSE
|
||||||
| NC score | 0.012405 (rank : 141) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64010 | Gene names | Crk, Crko | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene C-crk (P38) (Adapter molecule crk). | |||||
|
ZN646_HUMAN
|
||||||
| NC score | 0.010902 (rank : 142) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15015 | Gene names | ZNF646, KIAA0296 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 646. | |||||
|
RX_MOUSE
|
||||||
| NC score | 0.010440 (rank : 143) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35602, O08748 | Gene names | Rax, Rx | |||
|
Domain Architecture |

|
|||||
| Description | Retinal homeobox protein Rx (Retina and anterior neural fold homeobox protein). | |||||
|
TRIM8_HUMAN
|
||||||
| NC score | 0.010376 (rank : 144) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BZR9, Q9C028 | Gene names | TRIM8, GERP, RNF27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
ZN358_HUMAN
|
||||||
| NC score | 0.007082 (rank : 145) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NW07, Q9BTM7 | Gene names | ZNF358 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 358. | |||||
|
ABL1_HUMAN
|
||||||
| NC score | 0.005360 (rank : 146) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P00519, Q13869, Q13870, Q16133, Q45F09 | Gene names | ABL1, ABL, JTK7 | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||