Please be patient as the page loads
|
SNPC1_MOUSE
|
||||||
| SwissProt Accessions | Q8K0S9 | Gene names | Snapc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 1 (SNAPc subunit 1) (snRNA- activating protein complex 43 kDa subunit) (SNAPc 43 kDa subunit) (Small nuclear RNA-activating complex polypeptide 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SNPC1_MOUSE
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K0S9 | Gene names | Snapc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 1 (SNAPc subunit 1) (snRNA- activating protein complex 43 kDa subunit) (SNAPc 43 kDa subunit) (Small nuclear RNA-activating complex polypeptide 1). | |||||
|
SNPC1_HUMAN
|
||||||
| θ value | 4.71363e-158 (rank : 2) | NC score | 0.985756 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q16533 | Gene names | SNAPC1, SNAP43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 1 (SNAPc subunit 1) (snRNA- activating protein complex 43 kDa subunit) (SNAPc 43 kDa subunit) (Small nuclear RNA-activating complex polypeptide 1) (Proximal sequence element-binding transcription factor subunit gamma) (PSE- binding factor subunit gamma) (PTF subunit gamma). | |||||
|
LIPA2_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 3) | NC score | 0.023993 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1282 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BSS9, Q8BN73 | Gene names | Ppfia2 | |||
|
Domain Architecture |
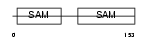
|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
CAC1F_MOUSE
|
||||||
| θ value | 0.62314 (rank : 4) | NC score | 0.018873 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JIS7 | Gene names | Cacna1f | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.813845 (rank : 5) | NC score | 0.041012 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.813845 (rank : 6) | NC score | 0.045035 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
SFRS2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 7) | NC score | 0.032844 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01130 | Gene names | SFRS2 | |||
|
Domain Architecture |
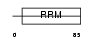
|
|||||
| Description | Splicing factor, arginine/serine-rich 2 (Splicing factor SC35) (SC-35) (Splicing component, 35 kDa) (Protein PR264). | |||||
|
SFRS2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 8) | NC score | 0.032844 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62093, Q542L3, Q60701 | Gene names | Sfrs2, Pr264, Sfrs10 | |||
|
Domain Architecture |
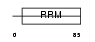
|
|||||
| Description | Splicing factor, arginine/serine-rich 2 (Splicing factor SC35) (SC-35) (Splicing component, 35 kDa) (Protein PR264) (Putative myelin regulatory factor 1) (MRF-1). | |||||
|
ATRX_MOUSE
|
||||||
| θ value | 4.03905 (rank : 9) | NC score | 0.019028 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
ATAD2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 10) | NC score | 0.011508 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 322 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PL18, Q658P2, Q68CQ0, Q6PJV6, Q8N890, Q9UHS5 | Gene names | ATAD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATPase family AAA domain-containing protein 2. | |||||
|
FILA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 11) | NC score | 0.021133 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
COMD9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 12) | NC score | 0.024797 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P000, Q96FI2, Q9H0R0 | Gene names | COMMD9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | COMM domain-containing protein 9. | |||||
|
PELO_HUMAN
|
||||||
| θ value | 6.88961 (rank : 13) | NC score | 0.021986 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BRX2, Q9GZS6, Q9Y306 | Gene names | PELO | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein pelota homolog. | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 14) | NC score | 0.017789 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
GLI3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 15) | NC score | 0.001811 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61602 | Gene names | Gli3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI3. | |||||
|
TRI44_HUMAN
|
||||||
| θ value | 8.99809 (rank : 16) | NC score | 0.011316 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
SNPC1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K0S9 | Gene names | Snapc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 1 (SNAPc subunit 1) (snRNA- activating protein complex 43 kDa subunit) (SNAPc 43 kDa subunit) (Small nuclear RNA-activating complex polypeptide 1). | |||||
|
SNPC1_HUMAN
|
||||||
| NC score | 0.985756 (rank : 2) | θ value | 4.71363e-158 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q16533 | Gene names | SNAPC1, SNAP43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 1 (SNAPc subunit 1) (snRNA- activating protein complex 43 kDa subunit) (SNAPc 43 kDa subunit) (Small nuclear RNA-activating complex polypeptide 1) (Proximal sequence element-binding transcription factor subunit gamma) (PSE- binding factor subunit gamma) (PTF subunit gamma). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.045035 (rank : 3) | θ value | 0.813845 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.041012 (rank : 4) | θ value | 0.813845 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
SFRS2_HUMAN
|
||||||
| NC score | 0.032844 (rank : 5) | θ value | 1.38821 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01130 | Gene names | SFRS2 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 2 (Splicing factor SC35) (SC-35) (Splicing component, 35 kDa) (Protein PR264). | |||||
|
SFRS2_MOUSE
|
||||||
| NC score | 0.032844 (rank : 6) | θ value | 1.38821 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62093, Q542L3, Q60701 | Gene names | Sfrs2, Pr264, Sfrs10 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 2 (Splicing factor SC35) (SC-35) (Splicing component, 35 kDa) (Protein PR264) (Putative myelin regulatory factor 1) (MRF-1). | |||||
|
COMD9_HUMAN
|
||||||
| NC score | 0.024797 (rank : 7) | θ value | 6.88961 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P000, Q96FI2, Q9H0R0 | Gene names | COMMD9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | COMM domain-containing protein 9. | |||||
|
LIPA2_MOUSE
|
||||||
| NC score | 0.023993 (rank : 8) | θ value | 0.0736092 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1282 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BSS9, Q8BN73 | Gene names | Ppfia2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
PELO_HUMAN
|
||||||
| NC score | 0.021986 (rank : 9) | θ value | 6.88961 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BRX2, Q9GZS6, Q9Y306 | Gene names | PELO | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein pelota homolog. | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.021133 (rank : 10) | θ value | 5.27518 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
ATRX_MOUSE
|
||||||
| NC score | 0.019028 (rank : 11) | θ value | 4.03905 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
CAC1F_MOUSE
|
||||||
| NC score | 0.018873 (rank : 12) | θ value | 0.62314 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JIS7 | Gene names | Cacna1f | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.017789 (rank : 13) | θ value | 8.99809 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
ATAD2_HUMAN
|
||||||
| NC score | 0.011508 (rank : 14) | θ value | 5.27518 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 322 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PL18, Q658P2, Q68CQ0, Q6PJV6, Q8N890, Q9UHS5 | Gene names | ATAD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATPase family AAA domain-containing protein 2. | |||||
|
TRI44_HUMAN
|
||||||
| NC score | 0.011316 (rank : 15) | θ value | 8.99809 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
GLI3_MOUSE
|
||||||
| NC score | 0.001811 (rank : 16) | θ value | 8.99809 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61602 | Gene names | Gli3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI3. | |||||