Please be patient as the page loads
|
PSME1_MOUSE
|
||||||
| SwissProt Accessions | P97371, O35561, Q9D841 | Gene names | Psme1 | |||
|
Domain Architecture |
|
|||||
| Description | Proteasome activator complex subunit 1 (Proteasome activator 28-alpha subunit) (PA28alpha) (PA28a) (Activator of multicatalytic protease subunit 1) (11S regulator complex subunit alpha) (REG-alpha). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PSME1_MOUSE
|
||||||
| θ value | 7.86017e-121 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97371, O35561, Q9D841 | Gene names | Psme1 | |||
|
Domain Architecture |

|
|||||
| Description | Proteasome activator complex subunit 1 (Proteasome activator 28-alpha subunit) (PA28alpha) (PA28a) (Activator of multicatalytic protease subunit 1) (11S regulator complex subunit alpha) (REG-alpha). | |||||
|
PSME1_HUMAN
|
||||||
| θ value | 2.61662e-116 (rank : 2) | NC score | 0.996819 (rank : 2) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q06323, Q9UEF4 | Gene names | PSME1, IFI5111 | |||
|
Domain Architecture |
|
|||||
| Description | Proteasome activator complex subunit 1 (Proteasome activator 28-alpha subunit) (PA28alpha) (PA28a) (Activator of multicatalytic protease subunit 1) (11S regulator complex subunit alpha) (REG-alpha) (Interferon gamma up-regulated I-5111 protein) (IGUP I-5111). | |||||
|
PSME2_MOUSE
|
||||||
| θ value | 5.19021e-56 (rank : 3) | NC score | 0.962779 (rank : 3) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97372, O35562 | Gene names | Psme2, Pa28b1 | |||
|
Domain Architecture |
|
|||||
| Description | Proteasome activator complex subunit 2 (Proteasome activator 28- subunit beta) (PA28beta) (PA28b) (Activator of multicatalytic protease subunit 2) (11S regulator complex subunit beta) (REG-beta). | |||||
|
PSME2_HUMAN
|
||||||
| θ value | 1.15626e-55 (rank : 4) | NC score | 0.962348 (rank : 4) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UL46, Q15129 | Gene names | PSME2 | |||
|
Domain Architecture |
|
|||||
| Description | Proteasome activator complex subunit 2 (Proteasome activator 28- subunit beta) (PA28beta) (PA28b) (Activator of multicatalytic protease subunit 2) (11S regulator complex subunit beta) (REG-beta). | |||||
|
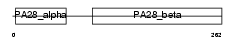
PSME3_HUMAN
|
||||||
| θ value | 1.56274e-52 (rank : 5) | NC score | 0.948527 (rank : 5) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P61289, O35563, P97373, Q12920, Q13172, Q9BQD9 | Gene names | PSME3 | |||
|
Domain Architecture |
|
|||||
| Description | Proteasome activator complex subunit 3 (Proteasome activator 28-gamma subunit) (PA28gamma) (PA28g) (Activator of multicatalytic protease subunit 3) (11S regulator complex subunit gamma) (REG-gamma) (Ki nuclear autoantigen). | |||||
|
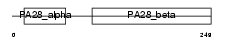
PSME3_MOUSE
|
||||||
| θ value | 1.56274e-52 (rank : 6) | NC score | 0.948527 (rank : 6) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P61290, O35563, P97373, Q12920, Q13172, Q9BQD9 | Gene names | Psme3 | |||
|
Domain Architecture |
|
|||||
| Description | Proteasome activator complex subunit 3 (Proteasome activator 28-gamma subunit) (PA28gamma) (PA28g) (Activator of multicatalytic protease subunit 3) (11S regulator complex subunit gamma) (REG-gamma) (Ki nuclear autoantigen). | |||||
|
SPT16_HUMAN
|
||||||
| θ value | 0.125558 (rank : 7) | NC score | 0.104800 (rank : 7) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y5B9, Q6GMT8, Q6P2F1, Q6PJM1, Q9NRX0 | Gene names | SUPT16H, FACT140, FACTP140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (hSPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin-specific transcription elongation factor 140 kDa subunit). | |||||
|
SPT16_MOUSE
|
||||||
| θ value | 0.125558 (rank : 8) | NC score | 0.103511 (rank : 8) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q920B9, Q3TZ48, Q3UFH0, Q921H4 | Gene names | Supt16h, Fact140, Factp140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin- specific transcription elongation factor 140 kDa subunit). | |||||
|
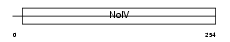
APOA4_MOUSE
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.033451 (rank : 10) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P06728 | Gene names | Apoa4 | |||
|
Domain Architecture |
|
|||||
| Description | Apolipoprotein A-IV precursor (Apo-AIV) (ApoA-IV). | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.017724 (rank : 14) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 1.81305 (rank : 11) | NC score | 0.010850 (rank : 20) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 2.36792 (rank : 12) | NC score | 0.013376 (rank : 17) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
CT077_HUMAN
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.035891 (rank : 9) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NQG5 | Gene names | C20orf77 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf77. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 14) | NC score | 0.010390 (rank : 23) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LYST_HUMAN
|
||||||
| θ value | 4.03905 (rank : 15) | NC score | 0.017303 (rank : 15) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99698, O43274, Q5T2U9, Q96TD7, Q96TD8, Q99709, Q9H133 | Gene names | LYST, CHS, CHS1 | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal-trafficking regulator (Beige homolog). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 4.03905 (rank : 16) | NC score | 0.010478 (rank : 22) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
REST_HUMAN
|
||||||
| θ value | 4.03905 (rank : 17) | NC score | 0.012070 (rank : 19) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
CCHCR_MOUSE
|
||||||
| θ value | 6.88961 (rank : 18) | NC score | 0.019927 (rank : 13) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 932 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K2I2 | Gene names | Cchcr1, Hcr | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil alpha-helical rod protein 1 (Alpha helical coiled-coil rod protein). | |||||
|
CYLN2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 19) | NC score | 0.012807 (rank : 18) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
ROCK2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 20) | NC score | 0.003407 (rank : 25) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
CSPG5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 21) | NC score | 0.010819 (rank : 21) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
EVPL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 22) | NC score | 0.009119 (rank : 24) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
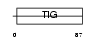
EXOC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 23) | NC score | 0.024146 (rank : 12) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96KP1, Q5JPC8, Q96AN6, Q9NUZ8, Q9UJM7 | Gene names | EXOC2, SEC5, SEC5L1 | |||
|
Domain Architecture |
|
|||||
| Description | Exocyst complex component 2 (Exocyst complex component Sec5). | |||||
|
EXOC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 24) | NC score | 0.024214 (rank : 11) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D4H1 | Gene names | Exoc2, Sec5, Sec5l1 | |||
|
Domain Architecture |
|
|||||
| Description | Exocyst complex component 2 (Exocyst complex component Sec5). | |||||
|
LYST_MOUSE
|
||||||
| θ value | 8.99809 (rank : 25) | NC score | 0.015551 (rank : 16) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97412, Q62403, Q8VBS6 | Gene names | Lyst, Bg, Chs1 | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal-trafficking regulator (Beige protein) (CHS1 homolog). | |||||
|
NKX61_MOUSE
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.002402 (rank : 27) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99MA9, Q9ERQ7 | Gene names | Nkx6-1, Nkx6.1, Nkx6a | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Nkx-6.1. | |||||
|
ROCK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.002873 (rank : 26) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
PSME1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 7.86017e-121 (rank : 1) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97371, O35561, Q9D841 | Gene names | Psme1 | |||
|
Domain Architecture |

|
|||||
| Description | Proteasome activator complex subunit 1 (Proteasome activator 28-alpha subunit) (PA28alpha) (PA28a) (Activator of multicatalytic protease subunit 1) (11S regulator complex subunit alpha) (REG-alpha). | |||||
|
PSME1_HUMAN
|
||||||
| NC score | 0.996819 (rank : 2) | θ value | 2.61662e-116 (rank : 2) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q06323, Q9UEF4 | Gene names | PSME1, IFI5111 | |||
|
Domain Architecture |

|
|||||
| Description | Proteasome activator complex subunit 1 (Proteasome activator 28-alpha subunit) (PA28alpha) (PA28a) (Activator of multicatalytic protease subunit 1) (11S regulator complex subunit alpha) (REG-alpha) (Interferon gamma up-regulated I-5111 protein) (IGUP I-5111). | |||||
|
PSME2_MOUSE
|
||||||
| NC score | 0.962779 (rank : 3) | θ value | 5.19021e-56 (rank : 3) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97372, O35562 | Gene names | Psme2, Pa28b1 | |||
|
Domain Architecture |

|
|||||
| Description | Proteasome activator complex subunit 2 (Proteasome activator 28- subunit beta) (PA28beta) (PA28b) (Activator of multicatalytic protease subunit 2) (11S regulator complex subunit beta) (REG-beta). | |||||
|
PSME2_HUMAN
|
||||||
| NC score | 0.962348 (rank : 4) | θ value | 1.15626e-55 (rank : 4) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UL46, Q15129 | Gene names | PSME2 | |||
|
Domain Architecture |

|
|||||
| Description | Proteasome activator complex subunit 2 (Proteasome activator 28- subunit beta) (PA28beta) (PA28b) (Activator of multicatalytic protease subunit 2) (11S regulator complex subunit beta) (REG-beta). | |||||
|
PSME3_HUMAN
|
||||||
| NC score | 0.948527 (rank : 5) | θ value | 1.56274e-52 (rank : 5) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P61289, O35563, P97373, Q12920, Q13172, Q9BQD9 | Gene names | PSME3 | |||
|
Domain Architecture |

|
|||||
| Description | Proteasome activator complex subunit 3 (Proteasome activator 28-gamma subunit) (PA28gamma) (PA28g) (Activator of multicatalytic protease subunit 3) (11S regulator complex subunit gamma) (REG-gamma) (Ki nuclear autoantigen). | |||||
|
PSME3_MOUSE
|
||||||
| NC score | 0.948527 (rank : 6) | θ value | 1.56274e-52 (rank : 6) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P61290, O35563, P97373, Q12920, Q13172, Q9BQD9 | Gene names | Psme3 | |||
|
Domain Architecture |

|
|||||
| Description | Proteasome activator complex subunit 3 (Proteasome activator 28-gamma subunit) (PA28gamma) (PA28g) (Activator of multicatalytic protease subunit 3) (11S regulator complex subunit gamma) (REG-gamma) (Ki nuclear autoantigen). | |||||
|
SPT16_HUMAN
|
||||||
| NC score | 0.104800 (rank : 7) | θ value | 0.125558 (rank : 7) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y5B9, Q6GMT8, Q6P2F1, Q6PJM1, Q9NRX0 | Gene names | SUPT16H, FACT140, FACTP140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (hSPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin-specific transcription elongation factor 140 kDa subunit). | |||||
|
SPT16_MOUSE
|
||||||
| NC score | 0.103511 (rank : 8) | θ value | 0.125558 (rank : 8) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q920B9, Q3TZ48, Q3UFH0, Q921H4 | Gene names | Supt16h, Fact140, Factp140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin- specific transcription elongation factor 140 kDa subunit). | |||||
|
CT077_HUMAN
|
||||||
| NC score | 0.035891 (rank : 9) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NQG5 | Gene names | C20orf77 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf77. | |||||
|
APOA4_MOUSE
|
||||||
| NC score | 0.033451 (rank : 10) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P06728 | Gene names | Apoa4 | |||
|
Domain Architecture |

|
|||||
| Description | Apolipoprotein A-IV precursor (Apo-AIV) (ApoA-IV). | |||||
|
EXOC2_MOUSE
|
||||||
| NC score | 0.024214 (rank : 11) | θ value | 8.99809 (rank : 24) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D4H1 | Gene names | Exoc2, Sec5, Sec5l1 | |||
|
Domain Architecture |

|
|||||
| Description | Exocyst complex component 2 (Exocyst complex component Sec5). | |||||
|
EXOC2_HUMAN
|
||||||
| NC score | 0.024146 (rank : 12) | θ value | 8.99809 (rank : 23) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96KP1, Q5JPC8, Q96AN6, Q9NUZ8, Q9UJM7 | Gene names | EXOC2, SEC5, SEC5L1 | |||
|
Domain Architecture |

|
|||||
| Description | Exocyst complex component 2 (Exocyst complex component Sec5). | |||||
|
CCHCR_MOUSE
|
||||||
| NC score | 0.019927 (rank : 13) | θ value | 6.88961 (rank : 18) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 932 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K2I2 | Gene names | Cchcr1, Hcr | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil alpha-helical rod protein 1 (Alpha helical coiled-coil rod protein). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.017724 (rank : 14) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
LYST_HUMAN
|
||||||
| NC score | 0.017303 (rank : 15) | θ value | 4.03905 (rank : 15) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99698, O43274, Q5T2U9, Q96TD7, Q96TD8, Q99709, Q9H133 | Gene names | LYST, CHS, CHS1 | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal-trafficking regulator (Beige homolog). | |||||
|
LYST_MOUSE
|
||||||
| NC score | 0.015551 (rank : 16) | θ value | 8.99809 (rank : 25) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97412, Q62403, Q8VBS6 | Gene names | Lyst, Bg, Chs1 | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal-trafficking regulator (Beige protein) (CHS1 homolog). | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.013376 (rank : 17) | θ value | 2.36792 (rank : 12) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
CYLN2_HUMAN
|
||||||
| NC score | 0.012807 (rank : 18) | θ value | 6.88961 (rank : 19) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
REST_HUMAN
|
||||||
| NC score | 0.012070 (rank : 19) | θ value | 4.03905 (rank : 17) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.010850 (rank : 20) | θ value | 1.81305 (rank : 11) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
CSPG5_MOUSE
|
||||||
| NC score | 0.010819 (rank : 21) | θ value | 8.99809 (rank : 21) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.010478 (rank : 22) | θ value | 4.03905 (rank : 16) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.010390 (rank : 23) | θ value | 3.0926 (rank : 14) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
EVPL_MOUSE
|
||||||
| NC score | 0.009119 (rank : 24) | θ value | 8.99809 (rank : 22) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
ROCK2_MOUSE
|
||||||
| NC score | 0.003407 (rank : 25) | θ value | 6.88961 (rank : 20) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
ROCK2_HUMAN
|
||||||
| NC score | 0.002873 (rank : 26) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
NKX61_MOUSE
|
||||||
| NC score | 0.002402 (rank : 27) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99MA9, Q9ERQ7 | Gene names | Nkx6-1, Nkx6.1, Nkx6a | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Nkx-6.1. | |||||