Please be patient as the page loads
|
PREP_HUMAN
|
||||||
| SwissProt Accessions | Q5JRX3, O95204, Q2M2G6, Q4VBR1, Q5JRW7, Q7L5Z7, Q9BSI6, Q9BVJ5, Q9UPP8 | Gene names | PITRM1, KIAA1104, MP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Presequence protease, mitochondrial precursor (EC 3.4.24.-) (hPreP) (Pitrilysin metalloproteinase 1) (Metalloprotease 1) (hMP1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PREP_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5JRX3, O95204, Q2M2G6, Q4VBR1, Q5JRW7, Q7L5Z7, Q9BSI6, Q9BVJ5, Q9UPP8 | Gene names | PITRM1, KIAA1104, MP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Presequence protease, mitochondrial precursor (EC 3.4.24.-) (hPreP) (Pitrilysin metalloproteinase 1) (Metalloprotease 1) (hMP1). | |||||
|
PREP_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.986161 (rank : 2) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K411, Q3THC4, Q3TJI1, Q3TPP0, Q3URQ8, Q4KUG2, Q4KUG3, Q6ZPX6, Q922N1, Q9CV63 | Gene names | Pitrm1, Kiaa1104, Ntup1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Presequence protease, mitochondrial precursor (EC 3.4.24.-) (Pitrilysin metalloproteinase 1). | |||||
|
FA13A_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 3) | NC score | 0.075131 (rank : 7) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O94988, Q8NBA3 | Gene names | FAM13A1, KIAA0914 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1. | |||||
|
IDE_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 4) | NC score | 0.138061 (rank : 3) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P14735 | Gene names | IDE | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-degrading enzyme (EC 3.4.24.56) (Insulysin) (Insulinase) (Insulin protease). | |||||
|
IDE_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 5) | NC score | 0.135856 (rank : 4) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JHR7 | Gene names | Ide | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-degrading enzyme (EC 3.4.24.56) (Insulysin) (Insulinase) (Insulin protease). | |||||
|
NRDC_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 6) | NC score | 0.134074 (rank : 5) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43847, O15241, O15242, Q5VUL0, Q96HB2, Q9NU57 | Gene names | NRD1 | |||
|
Domain Architecture |

|
|||||
| Description | Nardilysin precursor (EC 3.4.24.61) (N-arginine dibasic convertase) (NRD convertase) (NRD-C). | |||||
|
NRDC_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 7) | NC score | 0.132666 (rank : 6) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BHG1 | Gene names | Nrd1 | |||
|
Domain Architecture |

|
|||||
| Description | Nardilysin precursor (EC 3.4.24.61) (N-arginine dibasic convertase) (NRD convertase) (NRD-C). | |||||
|
SMC1A_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 8) | NC score | 0.030891 (rank : 11) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
FA13A_MOUSE
|
||||||
| θ value | 0.163984 (rank : 9) | NC score | 0.067377 (rank : 9) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGI4, Q8BZ91 | Gene names | Fam13a1, Precm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1 (Precm1 protein). | |||||
|
SMC1A_HUMAN
|
||||||
| θ value | 0.21417 (rank : 10) | NC score | 0.030923 (rank : 10) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
BRWD1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 11) | NC score | 0.027147 (rank : 12) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q921C3, Q921C2 | Gene names | Brwd1, Wdr9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
BTLA_HUMAN
|
||||||
| θ value | 1.06291 (rank : 12) | NC score | 0.069550 (rank : 8) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z6A9, Q6ZNH9 | Gene names | BTLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B- and T-lymphocyte attenuator precursor (B- and T-lymphocyte- associated protein) (CD272 antigen). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.018440 (rank : 15) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 3.0926 (rank : 14) | NC score | 0.007679 (rank : 26) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
TR240_HUMAN
|
||||||
| θ value | 3.0926 (rank : 15) | NC score | 0.022330 (rank : 13) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UHV7, O60334 | Gene names | THRAP1, ARC250, KIAA0593, TRAP240 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid hormone receptor-associated protein complex 240 kDa component (Trap240) (Thyroid hormone receptor-associated protein 1) (Vitamin D3 receptor-interacting protein complex component DRIP250) (DRIP 250) (Activator-recruited cofactor 250 kDa component) (ARC250). | |||||
|
BI2L1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 16) | NC score | 0.021325 (rank : 14) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHR4, Q75L21, Q75L22, Q96CV4, Q9Y2M8 | Gene names | BAIAP2L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1-associated protein 2-like protein 1 (BAI1-associated protein 2-like protein 1). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 17) | NC score | 0.017545 (rank : 17) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.012789 (rank : 20) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
FA50B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 19) | NC score | 0.017569 (rank : 16) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WTJ8, Q80X84, Q9D993 | Gene names | Fam50b, D0H6S2654E, X5l | |||
|
Domain Architecture |

|
|||||
| Description | Protein FAM50B (XAP-5-like protein). | |||||
|
ZN642_HUMAN
|
||||||
| θ value | 5.27518 (rank : 20) | NC score | 0.000984 (rank : 29) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q49AA0, Q5SWM5, Q6ZWK8 | Gene names | ZNF642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 642. | |||||
|
DNJA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 21) | NC score | 0.007117 (rank : 27) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P31689 | Gene names | DNAJA1, DNAJ2, HDJ2, HSJ2, HSPF4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2) (HSDJ). | |||||
|
GOGA3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 22) | NC score | 0.010258 (rank : 22) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q08378, O43241, Q6P9C7, Q86XW3, Q8TDA9, Q8WZA3 | Gene names | GOLGA3 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Golgi complex-associated protein of 170 kDa) (GCP170). | |||||
|
JADE1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 23) | NC score | 0.008764 (rank : 24) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6IE81, Q4W5D5, Q6ZSL7, Q8NC41, Q96JL8, Q96SQ1, Q9H692 | Gene names | PHF17, JADE1, KIAA1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | 0.008373 (rank : 25) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PLXB2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.010702 (rank : 21) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15031, Q7KZU3, Q9BSU7 | Gene names | PLXNB2, KIAA0315 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B2 precursor (MM1). | |||||
|
HS105_HUMAN
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.005464 (rank : 28) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92598, O95739, Q5TBM6, Q5TBM8, Q9UPC4 | Gene names | HSPH1, HSP105, HSP110, KIAA0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock 110 kDa protein) (Antigen NY- CO-25). | |||||
|
MYBB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.009431 (rank : 23) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P48972 | Gene names | Mybl2, Bmyb | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein B (B-Myb). | |||||
|
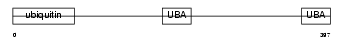
RD23B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.016183 (rank : 18) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54727, Q8WUB0 | Gene names | RAD23B | |||
|
Domain Architecture |
|
|||||
| Description | UV excision repair protein RAD23 homolog B (hHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
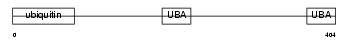
RD23B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.015544 (rank : 19) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54728 | Gene names | Rad23b, Mhr23b | |||
|
Domain Architecture |
|
|||||
| Description | UV excision repair protein RAD23 homolog B (mHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
PREP_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5JRX3, O95204, Q2M2G6, Q4VBR1, Q5JRW7, Q7L5Z7, Q9BSI6, Q9BVJ5, Q9UPP8 | Gene names | PITRM1, KIAA1104, MP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Presequence protease, mitochondrial precursor (EC 3.4.24.-) (hPreP) (Pitrilysin metalloproteinase 1) (Metalloprotease 1) (hMP1). | |||||
|
PREP_MOUSE
|
||||||
| NC score | 0.986161 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K411, Q3THC4, Q3TJI1, Q3TPP0, Q3URQ8, Q4KUG2, Q4KUG3, Q6ZPX6, Q922N1, Q9CV63 | Gene names | Pitrm1, Kiaa1104, Ntup1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Presequence protease, mitochondrial precursor (EC 3.4.24.-) (Pitrilysin metalloproteinase 1). | |||||
|
IDE_HUMAN
|
||||||
| NC score | 0.138061 (rank : 3) | θ value | 0.0330416 (rank : 4) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P14735 | Gene names | IDE | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-degrading enzyme (EC 3.4.24.56) (Insulysin) (Insulinase) (Insulin protease). | |||||
|
IDE_MOUSE
|
||||||
| NC score | 0.135856 (rank : 4) | θ value | 0.0431538 (rank : 5) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JHR7 | Gene names | Ide | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-degrading enzyme (EC 3.4.24.56) (Insulysin) (Insulinase) (Insulin protease). | |||||
|
NRDC_HUMAN
|
||||||
| NC score | 0.134074 (rank : 5) | θ value | 0.0961366 (rank : 6) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43847, O15241, O15242, Q5VUL0, Q96HB2, Q9NU57 | Gene names | NRD1 | |||
|
Domain Architecture |

|
|||||
| Description | Nardilysin precursor (EC 3.4.24.61) (N-arginine dibasic convertase) (NRD convertase) (NRD-C). | |||||
|
NRDC_MOUSE
|
||||||
| NC score | 0.132666 (rank : 6) | θ value | 0.0961366 (rank : 7) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BHG1 | Gene names | Nrd1 | |||
|
Domain Architecture |

|
|||||
| Description | Nardilysin precursor (EC 3.4.24.61) (N-arginine dibasic convertase) (NRD convertase) (NRD-C). | |||||
|
FA13A_HUMAN
|
||||||
| NC score | 0.075131 (rank : 7) | θ value | 0.0330416 (rank : 3) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O94988, Q8NBA3 | Gene names | FAM13A1, KIAA0914 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1. | |||||
|
BTLA_HUMAN
|
||||||
| NC score | 0.069550 (rank : 8) | θ value | 1.06291 (rank : 12) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z6A9, Q6ZNH9 | Gene names | BTLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B- and T-lymphocyte attenuator precursor (B- and T-lymphocyte- associated protein) (CD272 antigen). | |||||
|
FA13A_MOUSE
|
||||||
| NC score | 0.067377 (rank : 9) | θ value | 0.163984 (rank : 9) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGI4, Q8BZ91 | Gene names | Fam13a1, Precm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1 (Precm1 protein). | |||||
|
SMC1A_HUMAN
|
||||||
| NC score | 0.030923 (rank : 10) | θ value | 0.21417 (rank : 10) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
SMC1A_MOUSE
|
||||||
| NC score | 0.030891 (rank : 11) | θ value | 0.0961366 (rank : 8) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
BRWD1_MOUSE
|
||||||
| NC score | 0.027147 (rank : 12) | θ value | 0.813845 (rank : 11) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q921C3, Q921C2 | Gene names | Brwd1, Wdr9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
TR240_HUMAN
|
||||||
| NC score | 0.022330 (rank : 13) | θ value | 3.0926 (rank : 15) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UHV7, O60334 | Gene names | THRAP1, ARC250, KIAA0593, TRAP240 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid hormone receptor-associated protein complex 240 kDa component (Trap240) (Thyroid hormone receptor-associated protein 1) (Vitamin D3 receptor-interacting protein complex component DRIP250) (DRIP 250) (Activator-recruited cofactor 250 kDa component) (ARC250). | |||||
|
BI2L1_HUMAN
|
||||||
| NC score | 0.021325 (rank : 14) | θ value | 4.03905 (rank : 16) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHR4, Q75L21, Q75L22, Q96CV4, Q9Y2M8 | Gene names | BAIAP2L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1-associated protein 2-like protein 1 (BAI1-associated protein 2-like protein 1). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.018440 (rank : 15) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
FA50B_MOUSE
|
||||||
| NC score | 0.017569 (rank : 16) | θ value | 5.27518 (rank : 19) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WTJ8, Q80X84, Q9D993 | Gene names | Fam50b, D0H6S2654E, X5l | |||
|
Domain Architecture |

|
|||||
| Description | Protein FAM50B (XAP-5-like protein). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.017545 (rank : 17) | θ value | 4.03905 (rank : 17) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
RD23B_HUMAN
|
||||||
| NC score | 0.016183 (rank : 18) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54727, Q8WUB0 | Gene names | RAD23B | |||
|
Domain Architecture |

|
|||||
| Description | UV excision repair protein RAD23 homolog B (hHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
RD23B_MOUSE
|
||||||
| NC score | 0.015544 (rank : 19) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54728 | Gene names | Rad23b, Mhr23b | |||
|
Domain Architecture |

|
|||||
| Description | UV excision repair protein RAD23 homolog B (mHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.012789 (rank : 20) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
PLXB2_HUMAN
|
||||||
| NC score | 0.010702 (rank : 21) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15031, Q7KZU3, Q9BSU7 | Gene names | PLXNB2, KIAA0315 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B2 precursor (MM1). | |||||
|
GOGA3_HUMAN
|
||||||
| NC score | 0.010258 (rank : 22) | θ value | 6.88961 (rank : 22) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q08378, O43241, Q6P9C7, Q86XW3, Q8TDA9, Q8WZA3 | Gene names | GOLGA3 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Golgi complex-associated protein of 170 kDa) (GCP170). | |||||
|
MYBB_MOUSE
|
||||||
| NC score | 0.009431 (rank : 23) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P48972 | Gene names | Mybl2, Bmyb | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein B (B-Myb). | |||||
|
JADE1_HUMAN
|
||||||
| NC score | 0.008764 (rank : 24) | θ value | 6.88961 (rank : 23) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6IE81, Q4W5D5, Q6ZSL7, Q8NC41, Q96JL8, Q96SQ1, Q9H692 | Gene names | PHF17, JADE1, KIAA1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.008373 (rank : 25) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.007679 (rank : 26) | θ value | 3.0926 (rank : 14) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
DNJA1_HUMAN
|
||||||
| NC score | 0.007117 (rank : 27) | θ value | 6.88961 (rank : 21) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P31689 | Gene names | DNAJA1, DNAJ2, HDJ2, HSJ2, HSPF4 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily A member 1 (Heat shock 40 kDa protein 4) (DnaJ protein homolog 2) (HSJ-2) (HSDJ). | |||||
|
HS105_HUMAN
|
||||||
| NC score | 0.005464 (rank : 28) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92598, O95739, Q5TBM6, Q5TBM8, Q9UPC4 | Gene names | HSPH1, HSP105, HSP110, KIAA0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock 110 kDa protein) (Antigen NY- CO-25). | |||||
|
ZN642_HUMAN
|
||||||
| NC score | 0.000984 (rank : 29) | θ value | 5.27518 (rank : 20) | |||
| Query Neighborhood Hits | 29 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q49AA0, Q5SWM5, Q6ZWK8 | Gene names | ZNF642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 642. | |||||