Please be patient as the page loads
|
PLCC_HUMAN
|
||||||
| SwissProt Accessions | Q9NRZ7, Q3ZCU2, Q6UWP6, Q6ZUC6, Q8N3Q7, Q9NRZ6 | Gene names | AGPAT3 | |||
|
Domain Architecture |
|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase gamma (EC 2.3.1.51) (1- AGP acyltransferase 3) (1-AGPAT 3) (Lysophosphatidic acid acyltransferase-gamma) (LPAAT-gamma) (1-acylglycerol-3-phosphate O- acyltransferase 3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PLCC_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NRZ7, Q3ZCU2, Q6UWP6, Q6ZUC6, Q8N3Q7, Q9NRZ6 | Gene names | AGPAT3 | |||
|
Domain Architecture |

|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase gamma (EC 2.3.1.51) (1- AGP acyltransferase 3) (1-AGPAT 3) (Lysophosphatidic acid acyltransferase-gamma) (LPAAT-gamma) (1-acylglycerol-3-phosphate O- acyltransferase 3). | |||||
|
PLCC_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.998582 (rank : 2) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D517, Q7TT39, Q8BST2 | Gene names | Agpat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase gamma (EC 2.3.1.51) (1- AGP acyltransferase 3) (1-AGPAT 3) (Lysophosphatidic acid acyltransferase-gamma) (LPAAT-gamma) (1-acylglycerol-3-phosphate O- acyltransferase 3). | |||||
|
PLCD_MOUSE
|
||||||
| θ value | 8.93083e-141 (rank : 3) | NC score | 0.990062 (rank : 4) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K4X7, Q3TKN0, Q9DB84 | Gene names | Agpat4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase delta (EC 2.3.1.51) (1- AGP acyltransferase 4) (1-AGPAT 4) (Lysophosphatidic acid acyltransferase-delta) (LPAAT-delta) (1-acylglycerol-3-phosphate O- acyltransferase 4). | |||||
|
PLCD_HUMAN
|
||||||
| θ value | 1.92707e-135 (rank : 4) | NC score | 0.990500 (rank : 3) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NRZ5, Q5TEF0 | Gene names | AGPAT4 | |||
|
Domain Architecture |
|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase delta (EC 2.3.1.51) (1- AGP acyltransferase 4) (1-AGPAT 4) (Lysophosphatidic acid acyltransferase-delta) (LPAAT-delta) (1-acylglycerol-3-phosphate O- acyltransferase 4). | |||||
|
PLCE_HUMAN
|
||||||
| θ value | 7.32683e-18 (rank : 5) | NC score | 0.675792 (rank : 5) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NUQ2, Q8IZ47, Q9BQG4 | Gene names | AGPAT5 | |||
|
Domain Architecture |
|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase epsilon (EC 2.3.1.51) (1-AGP acyltransferase 5) (1-AGPAT 5) (Lysophosphatidic acid acyltransferase-epsilon) (LPAAT-epsilon) (1-acylglycerol-3-phosphate O-acyltransferase 5). | |||||
|
PLCE_MOUSE
|
||||||
| θ value | 2.13179e-17 (rank : 6) | NC score | 0.673172 (rank : 6) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D1E8, Q3U702, Q8BG61, Q8CGN6 | Gene names | Agpat5, D8Ertd319e | |||
|
Domain Architecture |
|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase epsilon (EC 2.3.1.51) (1-AGP acyltransferase 5) (1-AGPAT 5) (Lysophosphatidic acid acyltransferase-epsilon) (LPAAT-epsilon) (1-acylglycerol-3-phosphate O-acyltransferase 5). | |||||
|
LGAT1_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 7) | NC score | 0.349569 (rank : 7) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91YX5 | Gene names | Lpgat1, Fam34a | |||
|
Domain Architecture |
|
|||||
| Description | Acyl-CoA:lysophosphatidylglycerol acyltransferase 1 (EC 2.3.1.-). | |||||
|
LGAT1_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 8) | NC score | 0.319443 (rank : 8) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92604 | Gene names | LPGAT1, FAM34A, KIAA0205 | |||
|
Domain Architecture |
|
|||||
| Description | Acyl-CoA:lysophosphatidylglycerol acyltransferase 1 (EC 2.3.1.-). | |||||
|
ENH2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.039576 (rank : 10) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9N2J9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_3q26 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p60) (HERV-H/env60) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.040370 (rank : 9) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9N2J8 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_2q24.1 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p59) (HERV-H/env59) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 11) | NC score | 0.035679 (rank : 11) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9N2K0, O00354, Q96L63, Q9UNM3 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | HERV-H_2q24.3 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p62) (Env protein HERV-H19) (HERV- H/env62) (Env protein HERV-Hcl.3) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
CRLS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 12) | NC score | 0.019982 (rank : 12) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJA2, Q27RP0, Q69YQ5 | Gene names | CRLS1, C20orf155, CLS1 | |||
|
Domain Architecture |
|
|||||
| Description | Cardiolipin synthetase (EC 2.7.8.-) (Cardiolipin synthase) (CLS) (Protein GCD10 homolog). | |||||
|
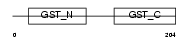
GSTO1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 13) | NC score | 0.019001 (rank : 13) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O09131, Q3TH87 | Gene names | Gsto1, Gstx, Gtsttl | |||
|
Domain Architecture |
|
|||||
| Description | Glutathione transferase omega-1 (EC 2.5.1.18) (GSTO 1-1) (p28). | |||||
|
RGS14_MOUSE
|
||||||
| θ value | 8.99809 (rank : 14) | NC score | 0.006158 (rank : 14) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||
|
PLCC_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NRZ7, Q3ZCU2, Q6UWP6, Q6ZUC6, Q8N3Q7, Q9NRZ6 | Gene names | AGPAT3 | |||
|
Domain Architecture |

|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase gamma (EC 2.3.1.51) (1- AGP acyltransferase 3) (1-AGPAT 3) (Lysophosphatidic acid acyltransferase-gamma) (LPAAT-gamma) (1-acylglycerol-3-phosphate O- acyltransferase 3). | |||||
|
PLCC_MOUSE
|
||||||
| NC score | 0.998582 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D517, Q7TT39, Q8BST2 | Gene names | Agpat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase gamma (EC 2.3.1.51) (1- AGP acyltransferase 3) (1-AGPAT 3) (Lysophosphatidic acid acyltransferase-gamma) (LPAAT-gamma) (1-acylglycerol-3-phosphate O- acyltransferase 3). | |||||
|
PLCD_HUMAN
|
||||||
| NC score | 0.990500 (rank : 3) | θ value | 1.92707e-135 (rank : 4) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NRZ5, Q5TEF0 | Gene names | AGPAT4 | |||
|
Domain Architecture |

|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase delta (EC 2.3.1.51) (1- AGP acyltransferase 4) (1-AGPAT 4) (Lysophosphatidic acid acyltransferase-delta) (LPAAT-delta) (1-acylglycerol-3-phosphate O- acyltransferase 4). | |||||
|
PLCD_MOUSE
|
||||||
| NC score | 0.990062 (rank : 4) | θ value | 8.93083e-141 (rank : 3) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K4X7, Q3TKN0, Q9DB84 | Gene names | Agpat4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase delta (EC 2.3.1.51) (1- AGP acyltransferase 4) (1-AGPAT 4) (Lysophosphatidic acid acyltransferase-delta) (LPAAT-delta) (1-acylglycerol-3-phosphate O- acyltransferase 4). | |||||
|
PLCE_HUMAN
|
||||||
| NC score | 0.675792 (rank : 5) | θ value | 7.32683e-18 (rank : 5) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NUQ2, Q8IZ47, Q9BQG4 | Gene names | AGPAT5 | |||
|
Domain Architecture |

|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase epsilon (EC 2.3.1.51) (1-AGP acyltransferase 5) (1-AGPAT 5) (Lysophosphatidic acid acyltransferase-epsilon) (LPAAT-epsilon) (1-acylglycerol-3-phosphate O-acyltransferase 5). | |||||
|
PLCE_MOUSE
|
||||||
| NC score | 0.673172 (rank : 6) | θ value | 2.13179e-17 (rank : 6) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D1E8, Q3U702, Q8BG61, Q8CGN6 | Gene names | Agpat5, D8Ertd319e | |||
|
Domain Architecture |

|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase epsilon (EC 2.3.1.51) (1-AGP acyltransferase 5) (1-AGPAT 5) (Lysophosphatidic acid acyltransferase-epsilon) (LPAAT-epsilon) (1-acylglycerol-3-phosphate O-acyltransferase 5). | |||||
|
LGAT1_MOUSE
|
||||||
| NC score | 0.349569 (rank : 7) | θ value | 4.1701e-05 (rank : 7) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91YX5 | Gene names | Lpgat1, Fam34a | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-CoA:lysophosphatidylglycerol acyltransferase 1 (EC 2.3.1.-). | |||||
|
LGAT1_HUMAN
|
||||||
| NC score | 0.319443 (rank : 8) | θ value | 0.00228821 (rank : 8) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92604 | Gene names | LPGAT1, FAM34A, KIAA0205 | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-CoA:lysophosphatidylglycerol acyltransferase 1 (EC 2.3.1.-). | |||||
|
ENH3_HUMAN
|
||||||
| NC score | 0.040370 (rank : 9) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9N2J8 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_2q24.1 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p59) (HERV-H/env59) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH2_HUMAN
|
||||||
| NC score | 0.039576 (rank : 10) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9N2J9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_3q26 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p60) (HERV-H/env60) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH1_HUMAN
|
||||||
| NC score | 0.035679 (rank : 11) | θ value | 2.36792 (rank : 11) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9N2K0, O00354, Q96L63, Q9UNM3 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_2q24.3 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p62) (Env protein HERV-H19) (HERV- H/env62) (Env protein HERV-Hcl.3) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
CRLS1_HUMAN
|
||||||
| NC score | 0.019982 (rank : 12) | θ value | 5.27518 (rank : 12) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJA2, Q27RP0, Q69YQ5 | Gene names | CRLS1, C20orf155, CLS1 | |||
|
Domain Architecture |

|
|||||
| Description | Cardiolipin synthetase (EC 2.7.8.-) (Cardiolipin synthase) (CLS) (Protein GCD10 homolog). | |||||
|
GSTO1_MOUSE
|
||||||
| NC score | 0.019001 (rank : 13) | θ value | 8.99809 (rank : 13) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O09131, Q3TH87 | Gene names | Gsto1, Gstx, Gtsttl | |||
|
Domain Architecture |

|
|||||
| Description | Glutathione transferase omega-1 (EC 2.5.1.18) (GSTO 1-1) (p28). | |||||
|
RGS14_MOUSE
|
||||||
| NC score | 0.006158 (rank : 14) | θ value | 8.99809 (rank : 14) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||