Please be patient as the page loads
|
PERM_HUMAN
|
||||||
| SwissProt Accessions | P05164, Q14862, Q4PJH5 | Gene names | MPO | |||
|
Domain Architecture |

|
|||||
| Description | Myeloperoxidase precursor (EC 1.11.1.7) (MPO) [Contains: 89 kDa myeloperoxidase; 84 kDa myeloperoxidase; Myeloperoxidase light chain; Myeloperoxidase heavy chain]. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PERE_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.996789 (rank : 3) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P11678, Q4TVP3 | Gene names | EPX, EPER, EPO, EPP | |||
|
Domain Architecture |

|
|||||
| Description | Eosinophil peroxidase precursor (EC 1.11.1.7) (EPO) [Contains: Eosinophil peroxidase light chain; Eosinophil peroxidase heavy chain]. | |||||
|
PERE_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.996155 (rank : 4) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49290, Q61798 | Gene names | Epx, Eper | |||
|
Domain Architecture |

|
|||||
| Description | Eosinophil peroxidase precursor (EC 1.11.1.7) (EPO) [Contains: Eosinophil peroxidase light chain; Eosinophil peroxidase heavy chain]. | |||||
|
PERL_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.991810 (rank : 5) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22079, Q13408 | Gene names | LPO, SAPX | |||
|
Domain Architecture |

|
|||||
| Description | Lactoperoxidase precursor (EC 1.11.1.7) (LPO) (Salivary peroxidase) (SPO). | |||||
|
PERM_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P05164, Q14862, Q4PJH5 | Gene names | MPO | |||
|
Domain Architecture |

|
|||||
| Description | Myeloperoxidase precursor (EC 1.11.1.7) (MPO) [Contains: 89 kDa myeloperoxidase; 84 kDa myeloperoxidase; Myeloperoxidase light chain; Myeloperoxidase heavy chain]. | |||||
|
PERM_MOUSE
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.999247 (rank : 2) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P11247 | Gene names | Mpo | |||
|
Domain Architecture |

|
|||||
| Description | Myeloperoxidase precursor (EC 1.11.1.7) (MPO) [Contains: Myeloperoxidase light chain; Myeloperoxidase heavy chain]. | |||||
|
PERT_HUMAN
|
||||||
| θ value | 4.39142e-172 (rank : 6) | NC score | 0.851230 (rank : 7) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P07202, P09934, P09935, Q8IUL0, Q8NF94, Q8NF95, Q8NF96, Q8NF97, Q8TCI9 | Gene names | TPO | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid peroxidase precursor (EC 1.11.1.8) (TPO). | |||||
|
PERT_MOUSE
|
||||||
| θ value | 2.03989e-169 (rank : 7) | NC score | 0.858518 (rank : 6) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35419, Q8C8B1 | Gene names | Tpo | |||
|
Domain Architecture |
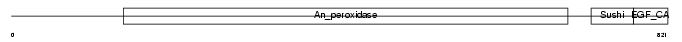
|
|||||
| Description | Thyroid peroxidase precursor (EC 1.11.1.8) (TPO). | |||||
|
DUOX1_HUMAN
|
||||||
| θ value | 2.68423e-28 (rank : 8) | NC score | 0.507442 (rank : 8) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NRD9, Q6ZMB3, Q6ZR09, Q9NZC1 | Gene names | DUOX1, DUOX, LNOX1, THOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 1 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH thyroid oxidase 1) (Thyroid oxidase 1) (Large NOX 1) (Long NOX 1). | |||||
|
DUOX2_HUMAN
|
||||||
| θ value | 1.62847e-25 (rank : 9) | NC score | 0.498406 (rank : 9) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NRD8, Q9NR02, Q9UHF9 | Gene names | DUOX2, LNOX2, THOX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 2 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH oxidase/peroxidase DUOX2) (NADPH thyroid oxidase 2) (Thyroid oxidase 2) (NADH/NADPH thyroid oxidase p138-tox) (p138 thyroid oxidase) (Large NOX 2) (Long NOX 2). | |||||
|
PGH1_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 10) | NC score | 0.285866 (rank : 10) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P22437 | Gene names | Ptgs1, Cox-1, Cox1 | |||
|
Domain Architecture |

|
|||||
| Description | Prostaglandin G/H synthase 1 precursor (EC 1.14.99.1) (Cyclooxygenase- 1) (COX-1) (Prostaglandin-endoperoxide synthase 1) (Prostaglandin H2 synthase 1) (PGH synthase 1) (PGHS-1) (PHS 1). | |||||
|
PGH2_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 11) | NC score | 0.271906 (rank : 12) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P35354, Q16876 | Gene names | PTGS2, COX2 | |||
|
Domain Architecture |

|
|||||
| Description | Prostaglandin G/H synthase 2 precursor (EC 1.14.99.1) (Cyclooxygenase- 2) (COX-2) (Prostaglandin-endoperoxide synthase 2) (Prostaglandin H2 synthase 2) (PGH synthase 2) (PGHS-2) (PHS II). | |||||
|
PGH1_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 12) | NC score | 0.278098 (rank : 11) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P23219, Q15122 | Gene names | PTGS1, COX1 | |||
|
Domain Architecture |

|
|||||
| Description | Prostaglandin G/H synthase 1 precursor (EC 1.14.99.1) (Cyclooxygenase- 1) (COX-1) (Prostaglandin-endoperoxide synthase 1) (Prostaglandin H2 synthase 1) (PGH synthase 1) (PGHS-1) (PHS 1). | |||||
|
PGH2_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 13) | NC score | 0.258958 (rank : 13) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q05769 | Gene names | Ptgs2, Cox-2, Cox2, Pghs-b, Tis10 | |||
|
Domain Architecture |

|
|||||
| Description | Prostaglandin G/H synthase 2 precursor (EC 1.14.99.1) (Cyclooxygenase- 2) (COX-2) (Prostaglandin-endoperoxide synthase 2) (Prostaglandin H2 synthase 2) (PGH synthase 2) (PGHS-2) (PHS II) (Glucocorticoid- regulated inflammatory cyclooxygenase) (Gripghs) (TIS10 protein) (Macrophage activation-associated marker protein P71/73) (PES-2). | |||||
|
ADAM9_MOUSE
|
||||||
| θ value | 3.0926 (rank : 14) | NC score | 0.005629 (rank : 24) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61072, Q60618, Q61853, Q80U94 | Gene names | Adam9, Kiaa0021, Mdc9, Mltng | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma). | |||||
|
ADAM9_HUMAN
|
||||||
| θ value | 4.03905 (rank : 15) | NC score | 0.005463 (rank : 25) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13443, Q10718, Q8NFM6 | Gene names | ADAM9, KIAA0021, MCMP, MDC9, MLTNG | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma) (Cellular disintegrin-related protein). | |||||
|
CNTP4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 16) | NC score | 0.006011 (rank : 23) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9C0A0 | Gene names | CNTNAP4, CASPR4, KIAA1763 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 4 precursor (Cell recognition molecule Caspr4). | |||||
|
NAT6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 17) | NC score | 0.010267 (rank : 22) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R123, Q8QZY5, Q8R014, Q9ERM0 | Gene names | Nat6, Fus2 | |||
|
Domain Architecture |

|
|||||
| Description | N-acetyltransferase 6 (EC 2.3.1.-) (Fus-2 protein) (Fusion 2 protein). | |||||
|
CY24B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 18) | NC score | 0.065653 (rank : 18) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04839, Q2PP16 | Gene names | CYBB, NOX2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome b-245 heavy chain (p22 phagocyte B-cytochrome) (Neutrophil cytochrome b 91 kDa polypeptide) (CGD91-phox) (gp91-phox) (gp91-1) (Heme-binding membrane glycoprotein gp91phox) (Cytochrome b(558) subunit beta) (Cytochrome b558 subunit beta) (Superoxide-generating NADPH oxidase heavy chain subunit) (NADPH oxidase 2). | |||||
|
CY24B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 19) | NC score | 0.065591 (rank : 19) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61093 | Gene names | Cybb, Cgd | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome b-245 heavy chain (p22 phagocyte B-cytochrome) (Neutrophil cytochrome b 91 kDa polypeptide) (CGD91-phox) (gp91-phox) (gp91-1) (Heme-binding membrane glycoprotein gp91phox) (Cytochrome b(558) subunit beta) (Cytochrome b558 subunit beta). | |||||
|
NOX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 20) | NC score | 0.069879 (rank : 16) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5S8, O95691, Q2PP02 | Gene names | NOX1, NOH1 | |||
|
Domain Architecture |
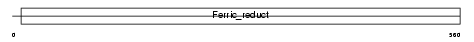
|
|||||
| Description | NADPH oxidase homolog 1 (NOX-1) (NOH-1) (NADH/NADPH mitogenic oxidase subunit P65-MOX) (Mitogenic oxidase 1) (MOX1). | |||||
|
NOX3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 21) | NC score | 0.071220 (rank : 15) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HBY0, Q9HBJ9 | Gene names | NOX3, MOX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 3 (EC 1.6.3.-) (gp91phox homolog 3) (GP91-3) (Mitogenic oxidase 2). | |||||
|
NOX3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 22) | NC score | 0.069633 (rank : 17) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q672J9, Q6Y4Q8 | Gene names | Nox3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 3 (EC 1.6.3.-). | |||||
|
NOX4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 23) | NC score | 0.061176 (rank : 21) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NPH5, Q5K3R4, Q5K3R5, Q5K3R6, Q5K3R8, Q7Z7G3, Q86V92 | Gene names | NOX4, RENOX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 4 (EC 1.6.3.-) (Kidney superoxide-producing NADPH oxidase) (KOX-1) (Renal NAD(P)H-oxidase). | |||||
|
NOX4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 24) | NC score | 0.062492 (rank : 20) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHI8, Q3TF39, Q8C3M1, Q8VCA3 | Gene names | Nox4, Renox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 4 (EC 1.6.3.-) (Kidney superoxide-producing NADPH oxidase) (Kox-1) (Renal NAD(P)H-oxidase) (Superoxide-generating NADPH oxidase 4). | |||||
|
NOX5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 25) | NC score | 0.081803 (rank : 14) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96PH1, Q8TEQ1, Q8TER4, Q96PH2, Q96PJ8, Q96PJ9, Q9H6E0, Q9HAM8 | Gene names | NOX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 5 (EC 1.6.3.-). | |||||
|
PERM_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P05164, Q14862, Q4PJH5 | Gene names | MPO | |||
|
Domain Architecture |

|
|||||
| Description | Myeloperoxidase precursor (EC 1.11.1.7) (MPO) [Contains: 89 kDa myeloperoxidase; 84 kDa myeloperoxidase; Myeloperoxidase light chain; Myeloperoxidase heavy chain]. | |||||
|
PERM_MOUSE
|
||||||
| NC score | 0.999247 (rank : 2) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P11247 | Gene names | Mpo | |||
|
Domain Architecture |

|
|||||
| Description | Myeloperoxidase precursor (EC 1.11.1.7) (MPO) [Contains: Myeloperoxidase light chain; Myeloperoxidase heavy chain]. | |||||
|
PERE_HUMAN
|
||||||
| NC score | 0.996789 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P11678, Q4TVP3 | Gene names | EPX, EPER, EPO, EPP | |||
|
Domain Architecture |

|
|||||
| Description | Eosinophil peroxidase precursor (EC 1.11.1.7) (EPO) [Contains: Eosinophil peroxidase light chain; Eosinophil peroxidase heavy chain]. | |||||
|
PERE_MOUSE
|
||||||
| NC score | 0.996155 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49290, Q61798 | Gene names | Epx, Eper | |||
|
Domain Architecture |

|
|||||
| Description | Eosinophil peroxidase precursor (EC 1.11.1.7) (EPO) [Contains: Eosinophil peroxidase light chain; Eosinophil peroxidase heavy chain]. | |||||
|
PERL_HUMAN
|
||||||
| NC score | 0.991810 (rank : 5) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22079, Q13408 | Gene names | LPO, SAPX | |||
|
Domain Architecture |

|
|||||
| Description | Lactoperoxidase precursor (EC 1.11.1.7) (LPO) (Salivary peroxidase) (SPO). | |||||
|
PERT_MOUSE
|
||||||
| NC score | 0.858518 (rank : 6) | θ value | 2.03989e-169 (rank : 7) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35419, Q8C8B1 | Gene names | Tpo | |||
|
Domain Architecture |
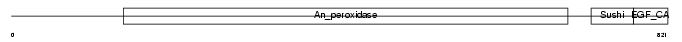
|
|||||
| Description | Thyroid peroxidase precursor (EC 1.11.1.8) (TPO). | |||||
|
PERT_HUMAN
|
||||||
| NC score | 0.851230 (rank : 7) | θ value | 4.39142e-172 (rank : 6) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P07202, P09934, P09935, Q8IUL0, Q8NF94, Q8NF95, Q8NF96, Q8NF97, Q8TCI9 | Gene names | TPO | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid peroxidase precursor (EC 1.11.1.8) (TPO). | |||||
|
DUOX1_HUMAN
|
||||||
| NC score | 0.507442 (rank : 8) | θ value | 2.68423e-28 (rank : 8) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NRD9, Q6ZMB3, Q6ZR09, Q9NZC1 | Gene names | DUOX1, DUOX, LNOX1, THOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 1 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH thyroid oxidase 1) (Thyroid oxidase 1) (Large NOX 1) (Long NOX 1). | |||||
|
DUOX2_HUMAN
|
||||||
| NC score | 0.498406 (rank : 9) | θ value | 1.62847e-25 (rank : 9) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NRD8, Q9NR02, Q9UHF9 | Gene names | DUOX2, LNOX2, THOX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 2 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH oxidase/peroxidase DUOX2) (NADPH thyroid oxidase 2) (Thyroid oxidase 2) (NADH/NADPH thyroid oxidase p138-tox) (p138 thyroid oxidase) (Large NOX 2) (Long NOX 2). | |||||
|
PGH1_MOUSE
|
||||||
| NC score | 0.285866 (rank : 10) | θ value | 2.44474e-05 (rank : 10) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P22437 | Gene names | Ptgs1, Cox-1, Cox1 | |||
|
Domain Architecture |

|
|||||
| Description | Prostaglandin G/H synthase 1 precursor (EC 1.14.99.1) (Cyclooxygenase- 1) (COX-1) (Prostaglandin-endoperoxide synthase 1) (Prostaglandin H2 synthase 1) (PGH synthase 1) (PGHS-1) (PHS 1). | |||||
|
PGH1_HUMAN
|
||||||
| NC score | 0.278098 (rank : 11) | θ value | 0.00035302 (rank : 12) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P23219, Q15122 | Gene names | PTGS1, COX1 | |||
|
Domain Architecture |

|
|||||
| Description | Prostaglandin G/H synthase 1 precursor (EC 1.14.99.1) (Cyclooxygenase- 1) (COX-1) (Prostaglandin-endoperoxide synthase 1) (Prostaglandin H2 synthase 1) (PGH synthase 1) (PGHS-1) (PHS 1). | |||||
|
PGH2_HUMAN
|
||||||
| NC score | 0.271906 (rank : 12) | θ value | 0.00020696 (rank : 11) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P35354, Q16876 | Gene names | PTGS2, COX2 | |||
|
Domain Architecture |

|
|||||
| Description | Prostaglandin G/H synthase 2 precursor (EC 1.14.99.1) (Cyclooxygenase- 2) (COX-2) (Prostaglandin-endoperoxide synthase 2) (Prostaglandin H2 synthase 2) (PGH synthase 2) (PGHS-2) (PHS II). | |||||
|
PGH2_MOUSE
|
||||||
| NC score | 0.258958 (rank : 13) | θ value | 0.00035302 (rank : 13) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q05769 | Gene names | Ptgs2, Cox-2, Cox2, Pghs-b, Tis10 | |||
|
Domain Architecture |

|
|||||
| Description | Prostaglandin G/H synthase 2 precursor (EC 1.14.99.1) (Cyclooxygenase- 2) (COX-2) (Prostaglandin-endoperoxide synthase 2) (Prostaglandin H2 synthase 2) (PGH synthase 2) (PGHS-2) (PHS II) (Glucocorticoid- regulated inflammatory cyclooxygenase) (Gripghs) (TIS10 protein) (Macrophage activation-associated marker protein P71/73) (PES-2). | |||||
|
NOX5_HUMAN
|
||||||
| NC score | 0.081803 (rank : 14) | θ value | θ > 10 (rank : 25) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96PH1, Q8TEQ1, Q8TER4, Q96PH2, Q96PJ8, Q96PJ9, Q9H6E0, Q9HAM8 | Gene names | NOX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 5 (EC 1.6.3.-). | |||||
|
NOX3_HUMAN
|
||||||
| NC score | 0.071220 (rank : 15) | θ value | θ > 10 (rank : 21) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HBY0, Q9HBJ9 | Gene names | NOX3, MOX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 3 (EC 1.6.3.-) (gp91phox homolog 3) (GP91-3) (Mitogenic oxidase 2). | |||||
|
NOX1_HUMAN
|
||||||
| NC score | 0.069879 (rank : 16) | θ value | θ > 10 (rank : 20) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5S8, O95691, Q2PP02 | Gene names | NOX1, NOH1 | |||
|
Domain Architecture |
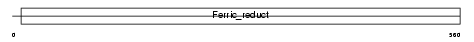
|
|||||
| Description | NADPH oxidase homolog 1 (NOX-1) (NOH-1) (NADH/NADPH mitogenic oxidase subunit P65-MOX) (Mitogenic oxidase 1) (MOX1). | |||||
|
NOX3_MOUSE
|
||||||
| NC score | 0.069633 (rank : 17) | θ value | θ > 10 (rank : 22) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q672J9, Q6Y4Q8 | Gene names | Nox3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 3 (EC 1.6.3.-). | |||||
|
CY24B_HUMAN
|
||||||
| NC score | 0.065653 (rank : 18) | θ value | θ > 10 (rank : 18) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04839, Q2PP16 | Gene names | CYBB, NOX2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome b-245 heavy chain (p22 phagocyte B-cytochrome) (Neutrophil cytochrome b 91 kDa polypeptide) (CGD91-phox) (gp91-phox) (gp91-1) (Heme-binding membrane glycoprotein gp91phox) (Cytochrome b(558) subunit beta) (Cytochrome b558 subunit beta) (Superoxide-generating NADPH oxidase heavy chain subunit) (NADPH oxidase 2). | |||||
|
CY24B_MOUSE
|
||||||
| NC score | 0.065591 (rank : 19) | θ value | θ > 10 (rank : 19) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61093 | Gene names | Cybb, Cgd | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome b-245 heavy chain (p22 phagocyte B-cytochrome) (Neutrophil cytochrome b 91 kDa polypeptide) (CGD91-phox) (gp91-phox) (gp91-1) (Heme-binding membrane glycoprotein gp91phox) (Cytochrome b(558) subunit beta) (Cytochrome b558 subunit beta). | |||||
|
NOX4_MOUSE
|
||||||
| NC score | 0.062492 (rank : 20) | θ value | θ > 10 (rank : 24) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHI8, Q3TF39, Q8C3M1, Q8VCA3 | Gene names | Nox4, Renox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 4 (EC 1.6.3.-) (Kidney superoxide-producing NADPH oxidase) (Kox-1) (Renal NAD(P)H-oxidase) (Superoxide-generating NADPH oxidase 4). | |||||
|
NOX4_HUMAN
|
||||||
| NC score | 0.061176 (rank : 21) | θ value | θ > 10 (rank : 23) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NPH5, Q5K3R4, Q5K3R5, Q5K3R6, Q5K3R8, Q7Z7G3, Q86V92 | Gene names | NOX4, RENOX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 4 (EC 1.6.3.-) (Kidney superoxide-producing NADPH oxidase) (KOX-1) (Renal NAD(P)H-oxidase). | |||||
|
NAT6_MOUSE
|
||||||
| NC score | 0.010267 (rank : 22) | θ value | 8.99809 (rank : 17) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R123, Q8QZY5, Q8R014, Q9ERM0 | Gene names | Nat6, Fus2 | |||
|
Domain Architecture |

|
|||||
| Description | N-acetyltransferase 6 (EC 2.3.1.-) (Fus-2 protein) (Fusion 2 protein). | |||||
|
CNTP4_HUMAN
|
||||||
| NC score | 0.006011 (rank : 23) | θ value | 6.88961 (rank : 16) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9C0A0 | Gene names | CNTNAP4, CASPR4, KIAA1763 | |||
|
Domain Architecture |

|
|||||
| Description | Contactin-associated protein-like 4 precursor (Cell recognition molecule Caspr4). | |||||
|
ADAM9_MOUSE
|
||||||
| NC score | 0.005629 (rank : 24) | θ value | 3.0926 (rank : 14) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61072, Q60618, Q61853, Q80U94 | Gene names | Adam9, Kiaa0021, Mdc9, Mltng | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma). | |||||
|
ADAM9_HUMAN
|
||||||
| NC score | 0.005463 (rank : 25) | θ value | 4.03905 (rank : 15) | |||
| Query Neighborhood Hits | 17 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13443, Q10718, Q8NFM6 | Gene names | ADAM9, KIAA0021, MCMP, MDC9, MLTNG | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma) (Cellular disintegrin-related protein). | |||||