Please be patient as the page loads
|
IGBP1_MOUSE
|
||||||
| SwissProt Accessions | Q61249 | Gene names | Igbp1, Pc52 | |||
|
Domain Architecture |
|
|||||
| Description | Immunoglobulin-binding protein 1 (CD79a-binding protein 1) (Alpha4 phosphoprotein) (Lymphocyte signal transduction molecule alpha 4) (p52). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IGBP1_MOUSE
|
||||||
| θ value | 6.35588e-163 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61249 | Gene names | Igbp1, Pc52 | |||
|
Domain Architecture |

|
|||||
| Description | Immunoglobulin-binding protein 1 (CD79a-binding protein 1) (Alpha4 phosphoprotein) (Lymphocyte signal transduction molecule alpha 4) (p52). | |||||
|
IGBP1_HUMAN
|
||||||
| θ value | 1.57644e-137 (rank : 2) | NC score | 0.986902 (rank : 2) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78318, Q8TAB2 | Gene names | IGBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Immunoglobulin-binding protein 1 (CD79a-binding protein 1) (B cell signal transduction molecule alpha 4) (Alpha 4 protein) (NY-REN-16 antigen). | |||||
|
IGB1B_MOUSE
|
||||||
| θ value | 6.44497e-115 (rank : 3) | NC score | 0.976425 (rank : 3) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9QZ29 | Gene names | Igbp1b | |||
|
Domain Architecture |
|
|||||
| Description | Immunoglobulin-binding protein 1b (CD79a-binding protein 1b) (Alpha 4- b protein) (Alpha4-b). | |||||
|
LIPA3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 4) | NC score | 0.058669 (rank : 5) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
TLK2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 5) | NC score | 0.012056 (rank : 32) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1044 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86UE8, Q9UKI7, Q9Y4F7 | Gene names | TLK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 2 (EC 2.7.11.1) (Tousled- like kinase 2) (PKU-alpha). | |||||
|
LIPA2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 6) | NC score | 0.051737 (rank : 8) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1044 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75334 | Gene names | PPFIA2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
LIPA2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 7) | NC score | 0.047802 (rank : 13) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1282 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BSS9, Q8BN73 | Gene names | Ppfia2 | |||
|
Domain Architecture |
|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
LIPA1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 8) | NC score | 0.054933 (rank : 6) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
BUB1B_MOUSE
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.037443 (rank : 17) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1S0 | Gene names | Bub1b, Mad3l | |||
|
Domain Architecture |

|
|||||
| Description | Mitotic checkpoint serine/threonine-protein kinase BUB1 beta (EC 2.7.11.1) (MAD3/BUB1-related protein kinase) (Mitotic checkpoint kinase MAD3L). | |||||
|
SCN9A_HUMAN
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.014312 (rank : 30) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15858, Q6B4R9, Q6B4S0, Q6B4S1, Q70HX1, Q70HX2, Q8WTU1, Q8WWN4 | Gene names | SCN9A, NENA, PN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 9 subunit alpha (Sodium channel protein type IX subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.7) (Neuroendocrine sodium channel) (hNE-Na) (Peripheral sodium channel 1). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 11) | NC score | 0.019060 (rank : 26) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
API5_MOUSE
|
||||||
| θ value | 1.81305 (rank : 12) | NC score | 0.051367 (rank : 9) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35841, Q922L2 | Gene names | Api5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis inhibitor 5 (API-5) (AAC-11). | |||||
|
API5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.049833 (rank : 10) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BZZ5, O15441, Q9Y4J7 | Gene names | API5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis inhibitor 5 (API-5) (Fibroblast growth factor 2-interacting factor) (FIF) (Protein XAGL) (Antiapoptosis clone 11 protein) (AAC- 11). | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 14) | NC score | 0.037834 (rank : 16) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 15) | NC score | 0.038982 (rank : 15) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
BRE1A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.022959 (rank : 23) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1025 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5DTM8, Q3UT10, Q3V350, Q7TT11, Q8BKA8, Q8BKN8, Q8BUF7, Q8BVU4 | Gene names | Rnf20, Bre1a, Kiaa4116 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (RING finger protein 20). | |||||
|
TRAF3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.049001 (rank : 11) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13114, Q12990, Q13076, Q13947, Q6AZX1, Q9UNL1 | Gene names | TRAF3, CAP1, CRAF1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 3 (CD40 receptor-associated factor 1) (CRAF1) (CD40-binding protein) (CD40BP) (LMP1-associated protein) (LAP1) (CAP-1). | |||||
|
TRAF3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 18) | NC score | 0.048647 (rank : 12) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 476 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60803, Q62380 | Gene names | Traf3, Craf1, Trafamn | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 3 (CD40 receptor-associated factor 1) (CRAF1) (TRAFAMN). | |||||
|
TLK1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 19) | NC score | 0.008987 (rank : 34) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UKI8, Q14150, Q8N591, Q9NYH2, Q9Y4F6 | Gene names | TLK1, KIAA0137 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 1 (EC 2.7.11.1) (Tousled- like kinase 1) (PKU-beta). | |||||
|
ZHX2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 20) | NC score | 0.014284 (rank : 31) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C0C0 | Gene names | Zhx2, Afr1, Raf | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
BRE1A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 21) | NC score | 0.021893 (rank : 24) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1071 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5VTR2, Q69YL5, Q6P527, Q8N3J4, Q96JD3, Q9H9Y7, Q9HA51, Q9NUR4, Q9NWQ3, Q9NX83 | Gene names | RNF20, BRE1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (hBRE1) (RING finger protein 20). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 5.27518 (rank : 22) | NC score | 0.011584 (rank : 33) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
RBP24_HUMAN
|
||||||
| θ value | 5.27518 (rank : 23) | NC score | 0.014998 (rank : 29) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
DOS_HUMAN
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | 0.040101 (rank : 14) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N350, O43385 | Gene names | DOS, C19orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Dos. | |||||
|
GSH0_HUMAN
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.029580 (rank : 21) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48507, Q6FHC1, Q9NPX9, Q9NU74 | Gene names | GCLM, GLCLR | |||
|
Domain Architecture |
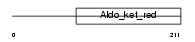
|
|||||
| Description | Glutamate--cysteine ligase regulatory subunit (EC 6.3.2.2) (Gamma- glutamylcysteine synthetase) (Gamma-ECS) (GCS light chain) (Glutamate--cysteine ligase modifier subunit). | |||||
|
GSH0_MOUSE
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.029546 (rank : 22) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O09172 | Gene names | Gclm, Glclr | |||
|
Domain Architecture |
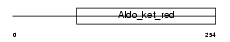
|
|||||
| Description | Glutamate--cysteine ligase regulatory subunit (EC 6.3.2.2) (Gamma- glutamylcysteine synthetase) (Gamma-ECS) (GCS light chain) (Glutamate--cysteine ligase modifier subunit). | |||||
|
LIPB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.032735 (rank : 18) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86W92, O75336, Q86X70, Q9NY03, Q9ULJ0 | Gene names | PPFIBP1, KIAA1230 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 1) (PTPRF-interacting protein-binding protein 1) (hSGT2). | |||||
|
CABIN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.015668 (rank : 27) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
NCOR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.031455 (rank : 19) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
NCOR1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.030812 (rank : 20) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
PDCD7_MOUSE
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.015288 (rank : 28) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WTY1, Q8R5D9 | Gene names | Pdcd7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Programmed cell death protein 7 (ES18). | |||||
|
SYT11_MOUSE
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.004769 (rank : 36) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R0N3 | Gene names | Syt11 | |||
|
Domain Architecture |
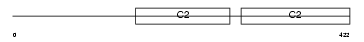
|
|||||
| Description | Synaptotagmin-11 (Synaptotagmin XI) (SytXI). | |||||
|
TRAF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.020963 (rank : 25) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13077, Q8NF13 | Gene names | TRAF1, EBI6 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 1 (Epstein-Barr virus-induced protein 6). | |||||
|
TRI13_MOUSE
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.007138 (rank : 35) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CYB0, Q8CEV0, Q923J0, Q925P1, Q99PQ0 | Gene names | Trim13, Rfp2 | |||
|
Domain Architecture |
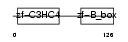
|
|||||
| Description | Tripartite motif-containing protein 13 (Ret finger protein 2) (Putative tumor suppressor RFP2). | |||||
|
LIPA3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 35) | NC score | 0.052328 (rank : 7) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P60469 | Gene names | Ppfia3 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
LIPA4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 36) | NC score | 0.062633 (rank : 4) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75335, O94971 | Gene names | PPFIA4, KIAA0897 | |||
|
Domain Architecture |
|
|||||
| Description | Liprin-alpha-4 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-4) (PTPRF-interacting protein alpha-4). | |||||
|
IGBP1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 6.35588e-163 (rank : 1) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61249 | Gene names | Igbp1, Pc52 | |||
|
Domain Architecture |

|
|||||
| Description | Immunoglobulin-binding protein 1 (CD79a-binding protein 1) (Alpha4 phosphoprotein) (Lymphocyte signal transduction molecule alpha 4) (p52). | |||||
|
IGBP1_HUMAN
|
||||||
| NC score | 0.986902 (rank : 2) | θ value | 1.57644e-137 (rank : 2) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78318, Q8TAB2 | Gene names | IGBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Immunoglobulin-binding protein 1 (CD79a-binding protein 1) (B cell signal transduction molecule alpha 4) (Alpha 4 protein) (NY-REN-16 antigen). | |||||
|
IGB1B_MOUSE
|
||||||
| NC score | 0.976425 (rank : 3) | θ value | 6.44497e-115 (rank : 3) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9QZ29 | Gene names | Igbp1b | |||
|
Domain Architecture |

|
|||||
| Description | Immunoglobulin-binding protein 1b (CD79a-binding protein 1b) (Alpha 4- b protein) (Alpha4-b). | |||||
|
LIPA4_HUMAN
|
||||||
| NC score | 0.062633 (rank : 4) | θ value | θ > 10 (rank : 36) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75335, O94971 | Gene names | PPFIA4, KIAA0897 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-4 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-4) (PTPRF-interacting protein alpha-4). | |||||
|
LIPA3_HUMAN
|
||||||
| NC score | 0.058669 (rank : 5) | θ value | 0.163984 (rank : 4) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
LIPA1_HUMAN
|
||||||
| NC score | 0.054933 (rank : 6) | θ value | 0.62314 (rank : 8) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
LIPA3_MOUSE
|
||||||
| NC score | 0.052328 (rank : 7) | θ value | θ > 10 (rank : 35) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P60469 | Gene names | Ppfia3 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
LIPA2_HUMAN
|
||||||
| NC score | 0.051737 (rank : 8) | θ value | 0.365318 (rank : 6) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1044 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75334 | Gene names | PPFIA2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
API5_MOUSE
|
||||||
| NC score | 0.051367 (rank : 9) | θ value | 1.81305 (rank : 12) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35841, Q922L2 | Gene names | Api5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis inhibitor 5 (API-5) (AAC-11). | |||||
|
API5_HUMAN
|
||||||
| NC score | 0.049833 (rank : 10) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BZZ5, O15441, Q9Y4J7 | Gene names | API5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis inhibitor 5 (API-5) (Fibroblast growth factor 2-interacting factor) (FIF) (Protein XAGL) (Antiapoptosis clone 11 protein) (AAC- 11). | |||||
|
TRAF3_HUMAN
|
||||||
| NC score | 0.049001 (rank : 11) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13114, Q12990, Q13076, Q13947, Q6AZX1, Q9UNL1 | Gene names | TRAF3, CAP1, CRAF1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 3 (CD40 receptor-associated factor 1) (CRAF1) (CD40-binding protein) (CD40BP) (LMP1-associated protein) (LAP1) (CAP-1). | |||||
|
TRAF3_MOUSE
|
||||||
| NC score | 0.048647 (rank : 12) | θ value | 3.0926 (rank : 18) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 476 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60803, Q62380 | Gene names | Traf3, Craf1, Trafamn | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 3 (CD40 receptor-associated factor 1) (CRAF1) (TRAFAMN). | |||||
|
LIPA2_MOUSE
|
||||||
| NC score | 0.047802 (rank : 13) | θ value | 0.365318 (rank : 7) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1282 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BSS9, Q8BN73 | Gene names | Ppfia2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
DOS_HUMAN
|
||||||
| NC score | 0.040101 (rank : 14) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N350, O43385 | Gene names | DOS, C19orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Dos. | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.038982 (rank : 15) | θ value | 2.36792 (rank : 15) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.037834 (rank : 16) | θ value | 2.36792 (rank : 14) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
BUB1B_MOUSE
|
||||||
| NC score | 0.037443 (rank : 17) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1S0 | Gene names | Bub1b, Mad3l | |||
|
Domain Architecture |

|
|||||
| Description | Mitotic checkpoint serine/threonine-protein kinase BUB1 beta (EC 2.7.11.1) (MAD3/BUB1-related protein kinase) (Mitotic checkpoint kinase MAD3L). | |||||
|
LIPB1_HUMAN
|
||||||
| NC score | 0.032735 (rank : 18) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86W92, O75336, Q86X70, Q9NY03, Q9ULJ0 | Gene names | PPFIBP1, KIAA1230 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 1) (PTPRF-interacting protein-binding protein 1) (hSGT2). | |||||
|
NCOR1_HUMAN
|
||||||
| NC score | 0.031455 (rank : 19) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
NCOR1_MOUSE
|
||||||
| NC score | 0.030812 (rank : 20) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
GSH0_HUMAN
|
||||||
| NC score | 0.029580 (rank : 21) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48507, Q6FHC1, Q9NPX9, Q9NU74 | Gene names | GCLM, GLCLR | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate--cysteine ligase regulatory subunit (EC 6.3.2.2) (Gamma- glutamylcysteine synthetase) (Gamma-ECS) (GCS light chain) (Glutamate--cysteine ligase modifier subunit). | |||||
|
GSH0_MOUSE
|
||||||
| NC score | 0.029546 (rank : 22) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O09172 | Gene names | Gclm, Glclr | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate--cysteine ligase regulatory subunit (EC 6.3.2.2) (Gamma- glutamylcysteine synthetase) (Gamma-ECS) (GCS light chain) (Glutamate--cysteine ligase modifier subunit). | |||||
|
BRE1A_MOUSE
|
||||||
| NC score | 0.022959 (rank : 23) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1025 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5DTM8, Q3UT10, Q3V350, Q7TT11, Q8BKA8, Q8BKN8, Q8BUF7, Q8BVU4 | Gene names | Rnf20, Bre1a, Kiaa4116 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (RING finger protein 20). | |||||
|
BRE1A_HUMAN
|
||||||
| NC score | 0.021893 (rank : 24) | θ value | 5.27518 (rank : 21) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1071 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5VTR2, Q69YL5, Q6P527, Q8N3J4, Q96JD3, Q9H9Y7, Q9HA51, Q9NUR4, Q9NWQ3, Q9NX83 | Gene names | RNF20, BRE1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (hBRE1) (RING finger protein 20). | |||||
|
TRAF1_HUMAN
|
||||||
| NC score | 0.020963 (rank : 25) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13077, Q8NF13 | Gene names | TRAF1, EBI6 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 1 (Epstein-Barr virus-induced protein 6). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.019060 (rank : 26) | θ value | 1.06291 (rank : 11) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
CABIN_HUMAN
|
||||||
| NC score | 0.015668 (rank : 27) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
PDCD7_MOUSE
|
||||||
| NC score | 0.015288 (rank : 28) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WTY1, Q8R5D9 | Gene names | Pdcd7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Programmed cell death protein 7 (ES18). | |||||
|
RBP24_HUMAN
|
||||||
| NC score | 0.014998 (rank : 29) | θ value | 5.27518 (rank : 23) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
SCN9A_HUMAN
|
||||||
| NC score | 0.014312 (rank : 30) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15858, Q6B4R9, Q6B4S0, Q6B4S1, Q70HX1, Q70HX2, Q8WTU1, Q8WWN4 | Gene names | SCN9A, NENA, PN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 9 subunit alpha (Sodium channel protein type IX subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.7) (Neuroendocrine sodium channel) (hNE-Na) (Peripheral sodium channel 1). | |||||
|
ZHX2_MOUSE
|
||||||
| NC score | 0.014284 (rank : 31) | θ value | 4.03905 (rank : 20) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C0C0 | Gene names | Zhx2, Afr1, Raf | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
TLK2_HUMAN
|
||||||
| NC score | 0.012056 (rank : 32) | θ value | 0.279714 (rank : 5) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1044 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86UE8, Q9UKI7, Q9Y4F7 | Gene names | TLK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 2 (EC 2.7.11.1) (Tousled- like kinase 2) (PKU-alpha). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.011584 (rank : 33) | θ value | 5.27518 (rank : 22) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
TLK1_HUMAN
|
||||||
| NC score | 0.008987 (rank : 34) | θ value | 4.03905 (rank : 19) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UKI8, Q14150, Q8N591, Q9NYH2, Q9Y4F6 | Gene names | TLK1, KIAA0137 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 1 (EC 2.7.11.1) (Tousled- like kinase 1) (PKU-beta). | |||||
|
TRI13_MOUSE
|
||||||
| NC score | 0.007138 (rank : 35) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CYB0, Q8CEV0, Q923J0, Q925P1, Q99PQ0 | Gene names | Trim13, Rfp2 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 13 (Ret finger protein 2) (Putative tumor suppressor RFP2). | |||||
|
SYT11_MOUSE
|
||||||
| NC score | 0.004769 (rank : 36) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R0N3 | Gene names | Syt11 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-11 (Synaptotagmin XI) (SytXI). | |||||