Please be patient as the page loads
|
HXK2_MOUSE
|
||||||
| SwissProt Accessions | O08528 | Gene names | Hk2 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-2 (EC 2.7.1.1) (Hexokinase type II) (HK II). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HXK1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.998018 (rank : 3) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P19367, O43443, O43444, O75574, Q5VTC3, Q96HC8, Q9NNZ4, Q9NNZ5 | Gene names | HK1 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-1 (EC 2.7.1.1) (Hexokinase type I) (HK I) (Brain form hexokinase). | |||||
|
HXK1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.997587 (rank : 4) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P17710, Q61659, Q64476, Q64479 | Gene names | Hk1 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-1 (EC 2.7.1.1) (Hexokinase type I) (HK I) (Hexokinase, tumor isozyme). | |||||
|
HXK2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999489 (rank : 2) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P52789, Q8WU87, Q9UN82 | Gene names | HK2 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-2 (EC 2.7.1.1) (Hexokinase type II) (HK II) (Muscle form hexokinase). | |||||
|
HXK2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08528 | Gene names | Hk2 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-2 (EC 2.7.1.1) (Hexokinase type II) (HK II). | |||||
|
HXK3_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.993053 (rank : 5) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P52790 | Gene names | HK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-3 (EC 2.7.1.1) (Hexokinase type III) (HK III). | |||||
|
HXK3_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.992919 (rank : 6) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3TRM8, Q3TAX6, Q3UDP1, Q3UEA4 | Gene names | Hk3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hexokinase-3 (EC 2.7.1.1) (Hexokinase type III) (HK III). | |||||
|
HXK4_HUMAN
|
||||||
| θ value | 4.42207e-148 (rank : 7) | NC score | 0.977742 (rank : 7) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35557, Q05810 | Gene names | GCK | |||
|
Domain Architecture |
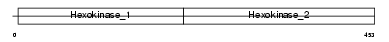
|
|||||
| Description | Glucokinase (EC 2.7.1.2) (Hexokinase-4) (Hexokinase type IV) (HK IV) (HK4) (Hexokinase-D). | |||||
|
HXK4_MOUSE
|
||||||
| θ value | 8.33966e-147 (rank : 8) | NC score | 0.977584 (rank : 8) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P52792, P52791 | Gene names | Gck, Gk | |||
|
Domain Architecture |
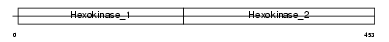
|
|||||
| Description | Glucokinase (EC 2.7.1.2) (Hexokinase-4) (Hexokinase type IV) (HK IV) (HK4) (Hexokinase-D). | |||||
|
BICD1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 9) | NC score | 0.014008 (rank : 12) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96G01, O43892, O43893 | Gene names | BICD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
BICD1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 10) | NC score | 0.010220 (rank : 13) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 837 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BR07, O55206, Q8BQ18, Q8BRR8, Q8C4D3, Q8CAK6, Q8R2J4, Q8R2J5, Q8R2J6, Q91YP7 | Gene names | Bicd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
TIF1B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 11) | NC score | 0.014970 (rank : 10) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13263, O00677, Q7Z632, Q93040, Q96IM1 | Gene names | TRIM28, KAP1, RNF96, TIF1B | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (Nuclear corepressor KAP-1) (KRAB- associated protein 1) (KAP-1) (KRAB-interacting protein 1) (KRIP-1) (RING finger protein 96). | |||||
|
C8AP2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 12) | NC score | 0.018413 (rank : 9) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
TIF1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 13) | NC score | 0.014060 (rank : 11) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
EMSY_HUMAN
|
||||||
| θ value | 4.03905 (rank : 14) | NC score | 0.010215 (rank : 14) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |
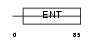
|
|||||
| Description | Protein EMSY. | |||||
|
TRIPB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 15) | NC score | 0.002843 (rank : 17) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
VLDLR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 16) | NC score | 0.006807 (rank : 16) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98155, Q5VVF6 | Gene names | VLDLR | |||
|
Domain Architecture |

|
|||||
| Description | Very low-density lipoprotein receptor precursor (VLDL receptor) (VLDL- R). | |||||
|
VLDLR_MOUSE
|
||||||
| θ value | 4.03905 (rank : 17) | NC score | 0.006900 (rank : 15) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98156, Q64022 | Gene names | Vldlr | |||
|
Domain Architecture |

|
|||||
| Description | Very low-density lipoprotein receptor precursor (VLDL receptor) (VLDL- R). | |||||
|
CYLN2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 18) | NC score | 0.001976 (rank : 18) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z0H8, Q7TSI9, Q8CHU1, Q9EP81 | Gene names | Cyln2, Kiaa0291 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115). | |||||
|
NIN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.001590 (rank : 19) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
HXK2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08528 | Gene names | Hk2 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-2 (EC 2.7.1.1) (Hexokinase type II) (HK II). | |||||
|
HXK2_HUMAN
|
||||||
| NC score | 0.999489 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P52789, Q8WU87, Q9UN82 | Gene names | HK2 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-2 (EC 2.7.1.1) (Hexokinase type II) (HK II) (Muscle form hexokinase). | |||||
|
HXK1_HUMAN
|
||||||
| NC score | 0.998018 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P19367, O43443, O43444, O75574, Q5VTC3, Q96HC8, Q9NNZ4, Q9NNZ5 | Gene names | HK1 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-1 (EC 2.7.1.1) (Hexokinase type I) (HK I) (Brain form hexokinase). | |||||
|
HXK1_MOUSE
|
||||||
| NC score | 0.997587 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P17710, Q61659, Q64476, Q64479 | Gene names | Hk1 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-1 (EC 2.7.1.1) (Hexokinase type I) (HK I) (Hexokinase, tumor isozyme). | |||||
|
HXK3_HUMAN
|
||||||
| NC score | 0.993053 (rank : 5) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P52790 | Gene names | HK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hexokinase-3 (EC 2.7.1.1) (Hexokinase type III) (HK III). | |||||
|
HXK3_MOUSE
|
||||||
| NC score | 0.992919 (rank : 6) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3TRM8, Q3TAX6, Q3UDP1, Q3UEA4 | Gene names | Hk3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hexokinase-3 (EC 2.7.1.1) (Hexokinase type III) (HK III). | |||||
|
HXK4_HUMAN
|
||||||
| NC score | 0.977742 (rank : 7) | θ value | 4.42207e-148 (rank : 7) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35557, Q05810 | Gene names | GCK | |||
|
Domain Architecture |

|
|||||
| Description | Glucokinase (EC 2.7.1.2) (Hexokinase-4) (Hexokinase type IV) (HK IV) (HK4) (Hexokinase-D). | |||||
|
HXK4_MOUSE
|
||||||
| NC score | 0.977584 (rank : 8) | θ value | 8.33966e-147 (rank : 8) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P52792, P52791 | Gene names | Gck, Gk | |||
|
Domain Architecture |

|
|||||
| Description | Glucokinase (EC 2.7.1.2) (Hexokinase-4) (Hexokinase type IV) (HK IV) (HK4) (Hexokinase-D). | |||||
|
C8AP2_MOUSE
|
||||||
| NC score | 0.018413 (rank : 9) | θ value | 3.0926 (rank : 12) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
TIF1B_HUMAN
|
||||||
| NC score | 0.014970 (rank : 10) | θ value | 2.36792 (rank : 11) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13263, O00677, Q7Z632, Q93040, Q96IM1 | Gene names | TRIM28, KAP1, RNF96, TIF1B | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (Nuclear corepressor KAP-1) (KRAB- associated protein 1) (KAP-1) (KRAB-interacting protein 1) (KRIP-1) (RING finger protein 96). | |||||
|
TIF1B_MOUSE
|
||||||
| NC score | 0.014060 (rank : 11) | θ value | 3.0926 (rank : 13) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
BICD1_HUMAN
|
||||||
| NC score | 0.014008 (rank : 12) | θ value | 0.62314 (rank : 9) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96G01, O43892, O43893 | Gene names | BICD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
BICD1_MOUSE
|
||||||
| NC score | 0.010220 (rank : 13) | θ value | 1.81305 (rank : 10) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 837 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BR07, O55206, Q8BQ18, Q8BRR8, Q8C4D3, Q8CAK6, Q8R2J4, Q8R2J5, Q8R2J6, Q91YP7 | Gene names | Bicd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
EMSY_HUMAN
|
||||||
| NC score | 0.010215 (rank : 14) | θ value | 4.03905 (rank : 14) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |

|
|||||
| Description | Protein EMSY. | |||||
|
VLDLR_MOUSE
|
||||||
| NC score | 0.006900 (rank : 15) | θ value | 4.03905 (rank : 17) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98156, Q64022 | Gene names | Vldlr | |||
|
Domain Architecture |

|
|||||
| Description | Very low-density lipoprotein receptor precursor (VLDL receptor) (VLDL- R). | |||||
|
VLDLR_HUMAN
|
||||||
| NC score | 0.006807 (rank : 16) | θ value | 4.03905 (rank : 16) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98155, Q5VVF6 | Gene names | VLDLR | |||
|
Domain Architecture |

|
|||||
| Description | Very low-density lipoprotein receptor precursor (VLDL receptor) (VLDL- R). | |||||
|
TRIPB_HUMAN
|
||||||
| NC score | 0.002843 (rank : 17) | θ value | 4.03905 (rank : 15) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
CYLN2_MOUSE
|
||||||
| NC score | 0.001976 (rank : 18) | θ value | 8.99809 (rank : 18) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z0H8, Q7TSI9, Q8CHU1, Q9EP81 | Gene names | Cyln2, Kiaa0291 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115). | |||||
|
NIN_MOUSE
|
||||||
| NC score | 0.001590 (rank : 19) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||