Please be patient as the page loads
|
HNRLL_HUMAN
|
||||||
| SwissProt Accessions | Q8WVV9, Q659B9, Q8IVH5, Q8IVH6, Q96HR5 | Gene names | HNRPLL, SRRF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L-like (Stromal RNA-regulating factor) (BLOCK24 protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HNRLL_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WVV9, Q659B9, Q8IVH5, Q8IVH6, Q96HR5 | Gene names | HNRPLL, SRRF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L-like (Stromal RNA-regulating factor) (BLOCK24 protein). | |||||
|
HNRLL_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.991782 (rank : 2) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q921F4, Q8BIP6, Q99J40 | Gene names | Hnrpll | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L-like. | |||||
|
HNRPL_MOUSE
|
||||||
| θ value | 1.85788e-146 (rank : 3) | NC score | 0.974754 (rank : 4) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8R081, O54789, Q8K0S7 | Gene names | Hnrpl | |||
|
Domain Architecture |
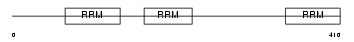
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L (hnRNP L). | |||||
|
HNRPL_HUMAN
|
||||||
| θ value | 1.57279e-145 (rank : 4) | NC score | 0.974846 (rank : 3) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P14866, Q9H3P3 | Gene names | HNRPL | |||
|
Domain Architecture |
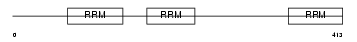
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L (hnRNP L). | |||||
|
PTBP2_MOUSE
|
||||||
| θ value | 3.61106e-41 (rank : 5) | NC score | 0.783929 (rank : 8) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q91Z31, Q7TNW7, Q9CUW2, Q9QYC2, Q9R0V9 | Gene names | Ptbp2, Brptb, Nptb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypyrimidine tract-binding protein 2 (Brain-enriched polypyrimidine tract-binding protein) (Brain-enriched PTB) (RRM-type RNA-binding protein brPTB) (Neural polypyrimidine tract-binding protein). | |||||
|
PTBP2_HUMAN
|
||||||
| θ value | 4.71619e-41 (rank : 6) | NC score | 0.783992 (rank : 7) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UKA9, Q8N0Z1, Q8N160, Q8NFB0, Q8NFB1, Q969N9, Q96Q76 | Gene names | PTBP2, NPTB, PTB, PTBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypyrimidine tract-binding protein 2 (Neurally-enriched homolog of PTB) (Neural polypyrimidine tract-binding protein) (PTB-like protein). | |||||
|
PTBP1_HUMAN
|
||||||
| θ value | 1.16164e-39 (rank : 7) | NC score | 0.769411 (rank : 10) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P26599 | Gene names | PTBP1, PTB | |||
|
Domain Architecture |

|
|||||
| Description | Polypyrimidine tract-binding protein 1 (PTB) (Heterogeneous nuclear ribonucleoprotein I) (hnRNP I) (57 kDa RNA-binding protein PPTB-1). | |||||
|
ROD1_MOUSE
|
||||||
| θ value | 1.42001e-37 (rank : 8) | NC score | 0.807576 (rank : 5) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BHD7, Q923C3 | Gene names | Rod1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of differentiation 1 (Rod1). | |||||
|
ROD1_HUMAN
|
||||||
| θ value | 2.96089e-35 (rank : 9) | NC score | 0.802813 (rank : 6) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95758, Q86YB3, Q86YH9 | Gene names | ROD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of differentiation 1 (Rod1). | |||||
|
PTBP1_MOUSE
|
||||||
| θ value | 7.54701e-31 (rank : 10) | NC score | 0.781014 (rank : 9) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P17225 | Gene names | Ptbp1, Ptb | |||
|
Domain Architecture |

|
|||||
| Description | Polypyrimidine tract-binding protein 1 (PTB) (Heterogeneous nuclear ribonucleoprotein I) (hnRNP I). | |||||
|
ZCRB1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 11) | NC score | 0.106490 (rank : 14) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8TBF4, Q6PJX0 | Gene names | ZCRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC-type and RNA-binding motif-containing protein 1 (U11/U12 snRNP 31 kDa protein) (U11/U12-31K). | |||||
|
NCOR1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 12) | NC score | 0.024212 (rank : 31) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
RBM23_HUMAN
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.048288 (rank : 25) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86U06, Q8ND16, Q8TB88, Q8WY40, Q9BUJ1, Q9NVV7 | Gene names | RBM23, RNPC4 | |||
|
Domain Architecture |
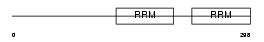
|
|||||
| Description | Probable RNA-binding protein 23 (RNA-binding motif protein 23) (RNA- binding region-containing protein 4) (Splicing factor SF2). | |||||
|
ZCRB1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.081654 (rank : 19) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CZ96, Q3TDL3 | Gene names | Zcrb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC-type and RNA-binding motif-containing protein 1 (MADP-1). | |||||
|
HNRPQ_HUMAN
|
||||||
| θ value | 1.38821 (rank : 15) | NC score | 0.050150 (rank : 24) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O60506, Q53H05, Q5TCG2, Q8IW78, Q8N599, Q96LC1, Q96LC2, Q9Y583 | Gene names | SYNCRIP, HNRPQ, NSAP1 | |||
|
Domain Architecture |
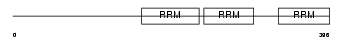
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein Q (hnRNP Q) (hnRNP-Q) (Synaptotagmin-binding, cytoplasmic RNA-interacting protein) (Glycine- and tyrosine-rich RNA-binding protein) (GRY-RBP) (NS1-associated protein 1). | |||||
|
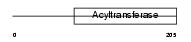
PLCA_MOUSE
|
||||||
| θ value | 1.38821 (rank : 16) | NC score | 0.036188 (rank : 27) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35083, O35446 | Gene names | Agpat1 | |||
|
Domain Architecture |
|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase alpha (EC 2.3.1.51) (1- AGP acyltransferase 1) (1-AGPAT 1) (Lysophosphatidic acid acyltransferase-alpha) (LPAAT-alpha) (1-acylglycerol-3-phosphate O- acyltransferase 1). | |||||
|
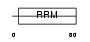
RBM7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 17) | NC score | 0.094165 (rank : 16) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y580 | Gene names | RBM7 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding protein 7 (RNA-binding motif protein 7). | |||||
|
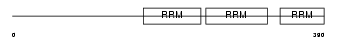
HNRPR_HUMAN
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.033948 (rank : 30) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43390 | Gene names | HNRPR | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein R (hnRNP R). | |||||
|
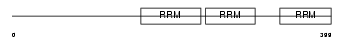
HNRPQ_MOUSE
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.045927 (rank : 26) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TMK9, O88991, Q80YV6, Q8BGP1, Q8C5K6, Q8CGC2, Q91ZR0, Q99KU9, Q9QYF4 | Gene names | Syncrip, Hnrpq, Nsap1, Nsap1l | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein Q (hnRNP Q) (hnRNP-Q) (Synaptotagmin-binding, cytoplasmic RNA-interacting protein) (Glycine- and tyrosine-rich RNA-binding protein) (GRY-RBP) (NS1-associated protein 1) (pp68). | |||||
|
JIP4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.018479 (rank : 33) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 585 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60271, O60905, Q3KQU8, Q3MKM7, Q86WC7, Q86WC8, Q8IZX7, Q96II0, Q9H811 | Gene names | SPAG9, HSS, KIAA0516, MAPK8IP4, SYD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (Human lung cancer protein 6) (HLC-6) (Proliferation-inducing protein 6) (Sperm-specific protein) (Sperm surface protein) (Protein highly expressed in testis) (PHET) (Sunday driver 1). | |||||
|
RBM7_MOUSE
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.083472 (rank : 17) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9CQT2, Q3UB91, Q7TQE3 | Gene names | Rbm7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 7 (RNA-binding motif protein 7). | |||||
|
TRA2B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 22) | NC score | 0.033972 (rank : 28) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P62995, O15449, Q15815, Q64283 | Gene names | SFRS10, TRA2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich-splicing factor 10 (Transformer-2-beta) (HTRA2- beta) (Transformer 2 protein homolog). | |||||
|
TRA2B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 23) | NC score | 0.033972 (rank : 29) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P62996, O15449, Q15815, Q64283 | Gene names | Sfrs10, Silg41, Tra2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich-splicing factor 10 (Transformer-2-beta) (HTRA2- beta) (Transformer 2 protein homolog) (Silica-induced gene 41 protein) (SIG-41). | |||||
|
JIP4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.014278 (rank : 34) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q58A65, Q3UH77, Q3UHF0, Q58VQ4, Q5NC70, Q5NC78, Q6A057, Q6PAS3, Q8CJC2 | Gene names | Spag9, Jip4, Jsap2, Kiaa0516, Mapk8ip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (JNK/SAPK-associated protein 2) (JSAP2). | |||||
|
COG5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.022928 (rank : 32) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UP83, O14555, O95008 | Gene names | COG5, GOLTC1, GTC90 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Conserved oligomeric Golgi complex component 5 (13S Golgi transport complex 90 kDa subunit) (GTC-90) (Golgi transport complex 1). | |||||
|
MATR3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 26) | NC score | 0.161094 (rank : 12) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P43243, Q9UHW0, Q9UQ27 | Gene names | MATR3, KIAA0723 | |||
|
Domain Architecture |

|
|||||
| Description | Matrin-3. | |||||
|
MATR3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 27) | NC score | 0.169442 (rank : 11) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K310 | Gene names | Matr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Matrin-3. | |||||
|
NONO_HUMAN
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.061369 (rank : 22) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15233, O00201, P30807, Q12786, Q9BQC5 | Gene names | NONO, NRB54 | |||
|
Domain Architecture |
|
|||||
| Description | Non-POU domain-containing octamer-binding protein (NonO protein) (54 kDa nuclear RNA- and DNA-binding protein) (p54(nrb)) (p54nrb) (55 kDa nuclear protein) (NMT55) (DNA-binding p52/p100 complex, 52 kDa subunit). | |||||
|
NONO_MOUSE
|
||||||
| θ value | θ > 10 (rank : 29) | NC score | 0.062146 (rank : 21) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99K48, Q63887, Q9CYQ4, Q9DBP2 | Gene names | Nono | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Non-POU domain-containing octamer-binding protein (NonO protein). | |||||
|
RU2B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 30) | NC score | 0.106953 (rank : 13) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P08579, Q9UJD4 | Gene names | SNRPB2 | |||
|
Domain Architecture |
|
|||||
| Description | U2 small nuclear ribonucleoprotein B''. | |||||
|
RU2B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 31) | NC score | 0.105926 (rank : 15) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9CQI7, Q9CW35, Q9CZ66 | Gene names | Snrpb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U2 small nuclear ribonucleoprotein B''. | |||||
|
SNRPA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 32) | NC score | 0.051083 (rank : 23) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62189, Q52LA2 | Gene names | Snrpa, Rnu1a-1 | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein A (U1 snRNP protein A) (U1A protein) (U1-A). | |||||
|
ZN638_HUMAN
|
||||||
| θ value | θ > 10 (rank : 33) | NC score | 0.080165 (rank : 20) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
ZN638_MOUSE
|
||||||
| θ value | θ > 10 (rank : 34) | NC score | 0.083458 (rank : 18) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
HNRLL_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WVV9, Q659B9, Q8IVH5, Q8IVH6, Q96HR5 | Gene names | HNRPLL, SRRF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L-like (Stromal RNA-regulating factor) (BLOCK24 protein). | |||||
|
HNRLL_MOUSE
|
||||||
| NC score | 0.991782 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q921F4, Q8BIP6, Q99J40 | Gene names | Hnrpll | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L-like. | |||||
|
HNRPL_HUMAN
|
||||||
| NC score | 0.974846 (rank : 3) | θ value | 1.57279e-145 (rank : 4) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P14866, Q9H3P3 | Gene names | HNRPL | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L (hnRNP L). | |||||
|
HNRPL_MOUSE
|
||||||
| NC score | 0.974754 (rank : 4) | θ value | 1.85788e-146 (rank : 3) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8R081, O54789, Q8K0S7 | Gene names | Hnrpl | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L (hnRNP L). | |||||
|
ROD1_MOUSE
|
||||||
| NC score | 0.807576 (rank : 5) | θ value | 1.42001e-37 (rank : 8) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BHD7, Q923C3 | Gene names | Rod1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of differentiation 1 (Rod1). | |||||
|
ROD1_HUMAN
|
||||||
| NC score | 0.802813 (rank : 6) | θ value | 2.96089e-35 (rank : 9) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95758, Q86YB3, Q86YH9 | Gene names | ROD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of differentiation 1 (Rod1). | |||||
|
PTBP2_HUMAN
|
||||||
| NC score | 0.783992 (rank : 7) | θ value | 4.71619e-41 (rank : 6) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UKA9, Q8N0Z1, Q8N160, Q8NFB0, Q8NFB1, Q969N9, Q96Q76 | Gene names | PTBP2, NPTB, PTB, PTBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypyrimidine tract-binding protein 2 (Neurally-enriched homolog of PTB) (Neural polypyrimidine tract-binding protein) (PTB-like protein). | |||||
|
PTBP2_MOUSE
|
||||||
| NC score | 0.783929 (rank : 8) | θ value | 3.61106e-41 (rank : 5) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q91Z31, Q7TNW7, Q9CUW2, Q9QYC2, Q9R0V9 | Gene names | Ptbp2, Brptb, Nptb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypyrimidine tract-binding protein 2 (Brain-enriched polypyrimidine tract-binding protein) (Brain-enriched PTB) (RRM-type RNA-binding protein brPTB) (Neural polypyrimidine tract-binding protein). | |||||
|
PTBP1_MOUSE
|
||||||
| NC score | 0.781014 (rank : 9) | θ value | 7.54701e-31 (rank : 10) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P17225 | Gene names | Ptbp1, Ptb | |||
|
Domain Architecture |

|
|||||
| Description | Polypyrimidine tract-binding protein 1 (PTB) (Heterogeneous nuclear ribonucleoprotein I) (hnRNP I). | |||||
|
PTBP1_HUMAN
|
||||||
| NC score | 0.769411 (rank : 10) | θ value | 1.16164e-39 (rank : 7) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P26599 | Gene names | PTBP1, PTB | |||
|
Domain Architecture |

|
|||||
| Description | Polypyrimidine tract-binding protein 1 (PTB) (Heterogeneous nuclear ribonucleoprotein I) (hnRNP I) (57 kDa RNA-binding protein PPTB-1). | |||||
|
MATR3_MOUSE
|
||||||
| NC score | 0.169442 (rank : 11) | θ value | θ > 10 (rank : 27) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K310 | Gene names | Matr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Matrin-3. | |||||
|
MATR3_HUMAN
|
||||||
| NC score | 0.161094 (rank : 12) | θ value | θ > 10 (rank : 26) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P43243, Q9UHW0, Q9UQ27 | Gene names | MATR3, KIAA0723 | |||
|
Domain Architecture |

|
|||||
| Description | Matrin-3. | |||||
|
RU2B_HUMAN
|
||||||
| NC score | 0.106953 (rank : 13) | θ value | θ > 10 (rank : 30) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P08579, Q9UJD4 | Gene names | SNRPB2 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein B''. | |||||
|
ZCRB1_HUMAN
|
||||||
| NC score | 0.106490 (rank : 14) | θ value | 0.21417 (rank : 11) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8TBF4, Q6PJX0 | Gene names | ZCRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC-type and RNA-binding motif-containing protein 1 (U11/U12 snRNP 31 kDa protein) (U11/U12-31K). | |||||
|
RU2B_MOUSE
|
||||||
| NC score | 0.105926 (rank : 15) | θ value | θ > 10 (rank : 31) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9CQI7, Q9CW35, Q9CZ66 | Gene names | Snrpb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U2 small nuclear ribonucleoprotein B''. | |||||
|
RBM7_HUMAN
|
||||||
| NC score | 0.094165 (rank : 16) | θ value | 1.38821 (rank : 17) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y580 | Gene names | RBM7 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 7 (RNA-binding motif protein 7). | |||||
|
RBM7_MOUSE
|
||||||
| NC score | 0.083472 (rank : 17) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9CQT2, Q3UB91, Q7TQE3 | Gene names | Rbm7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 7 (RNA-binding motif protein 7). | |||||
|
ZN638_MOUSE
|
||||||
| NC score | 0.083458 (rank : 18) | θ value | θ > 10 (rank : 34) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
ZCRB1_MOUSE
|
||||||
| NC score | 0.081654 (rank : 19) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CZ96, Q3TDL3 | Gene names | Zcrb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC-type and RNA-binding motif-containing protein 1 (MADP-1). | |||||
|
ZN638_HUMAN
|
||||||
| NC score | 0.080165 (rank : 20) | θ value | θ > 10 (rank : 33) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
NONO_MOUSE
|
||||||
| NC score | 0.062146 (rank : 21) | θ value | θ > 10 (rank : 29) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99K48, Q63887, Q9CYQ4, Q9DBP2 | Gene names | Nono | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Non-POU domain-containing octamer-binding protein (NonO protein). | |||||
|
NONO_HUMAN
|
||||||
| NC score | 0.061369 (rank : 22) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15233, O00201, P30807, Q12786, Q9BQC5 | Gene names | NONO, NRB54 | |||
|
Domain Architecture |

|
|||||
| Description | Non-POU domain-containing octamer-binding protein (NonO protein) (54 kDa nuclear RNA- and DNA-binding protein) (p54(nrb)) (p54nrb) (55 kDa nuclear protein) (NMT55) (DNA-binding p52/p100 complex, 52 kDa subunit). | |||||
|
SNRPA_MOUSE
|
||||||
| NC score | 0.051083 (rank : 23) | θ value | θ > 10 (rank : 32) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62189, Q52LA2 | Gene names | Snrpa, Rnu1a-1 | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein A (U1 snRNP protein A) (U1A protein) (U1-A). | |||||
|
HNRPQ_HUMAN
|
||||||
| NC score | 0.050150 (rank : 24) | θ value | 1.38821 (rank : 15) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O60506, Q53H05, Q5TCG2, Q8IW78, Q8N599, Q96LC1, Q96LC2, Q9Y583 | Gene names | SYNCRIP, HNRPQ, NSAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein Q (hnRNP Q) (hnRNP-Q) (Synaptotagmin-binding, cytoplasmic RNA-interacting protein) (Glycine- and tyrosine-rich RNA-binding protein) (GRY-RBP) (NS1-associated protein 1). | |||||
|
RBM23_HUMAN
|
||||||
| NC score | 0.048288 (rank : 25) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86U06, Q8ND16, Q8TB88, Q8WY40, Q9BUJ1, Q9NVV7 | Gene names | RBM23, RNPC4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable RNA-binding protein 23 (RNA-binding motif protein 23) (RNA- binding region-containing protein 4) (Splicing factor SF2). | |||||
|
HNRPQ_MOUSE
|
||||||
| NC score | 0.045927 (rank : 26) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TMK9, O88991, Q80YV6, Q8BGP1, Q8C5K6, Q8CGC2, Q91ZR0, Q99KU9, Q9QYF4 | Gene names | Syncrip, Hnrpq, Nsap1, Nsap1l | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein Q (hnRNP Q) (hnRNP-Q) (Synaptotagmin-binding, cytoplasmic RNA-interacting protein) (Glycine- and tyrosine-rich RNA-binding protein) (GRY-RBP) (NS1-associated protein 1) (pp68). | |||||
|
PLCA_MOUSE
|
||||||
| NC score | 0.036188 (rank : 27) | θ value | 1.38821 (rank : 16) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35083, O35446 | Gene names | Agpat1 | |||
|
Domain Architecture |

|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase alpha (EC 2.3.1.51) (1- AGP acyltransferase 1) (1-AGPAT 1) (Lysophosphatidic acid acyltransferase-alpha) (LPAAT-alpha) (1-acylglycerol-3-phosphate O- acyltransferase 1). | |||||
|
TRA2B_HUMAN
|
||||||
| NC score | 0.033972 (rank : 28) | θ value | 3.0926 (rank : 22) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P62995, O15449, Q15815, Q64283 | Gene names | SFRS10, TRA2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich-splicing factor 10 (Transformer-2-beta) (HTRA2- beta) (Transformer 2 protein homolog). | |||||
|
TRA2B_MOUSE
|
||||||
| NC score | 0.033972 (rank : 29) | θ value | 3.0926 (rank : 23) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P62996, O15449, Q15815, Q64283 | Gene names | Sfrs10, Silg41, Tra2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich-splicing factor 10 (Transformer-2-beta) (HTRA2- beta) (Transformer 2 protein homolog) (Silica-induced gene 41 protein) (SIG-41). | |||||
|
HNRPR_HUMAN
|
||||||
| NC score | 0.033948 (rank : 30) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43390 | Gene names | HNRPR | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein R (hnRNP R). | |||||
|
NCOR1_HUMAN
|
||||||
| NC score | 0.024212 (rank : 31) | θ value | 0.62314 (rank : 12) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
COG5_HUMAN
|
||||||
| NC score | 0.022928 (rank : 32) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UP83, O14555, O95008 | Gene names | COG5, GOLTC1, GTC90 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Conserved oligomeric Golgi complex component 5 (13S Golgi transport complex 90 kDa subunit) (GTC-90) (Golgi transport complex 1). | |||||
|
JIP4_HUMAN
|
||||||
| NC score | 0.018479 (rank : 33) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 585 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60271, O60905, Q3KQU8, Q3MKM7, Q86WC7, Q86WC8, Q8IZX7, Q96II0, Q9H811 | Gene names | SPAG9, HSS, KIAA0516, MAPK8IP4, SYD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (Human lung cancer protein 6) (HLC-6) (Proliferation-inducing protein 6) (Sperm-specific protein) (Sperm surface protein) (Protein highly expressed in testis) (PHET) (Sunday driver 1). | |||||
|
JIP4_MOUSE
|
||||||
| NC score | 0.014278 (rank : 34) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 25 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q58A65, Q3UH77, Q3UHF0, Q58VQ4, Q5NC70, Q5NC78, Q6A057, Q6PAS3, Q8CJC2 | Gene names | Spag9, Jip4, Jsap2, Kiaa0516, Mapk8ip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (JNK/SAPK-associated protein 2) (JSAP2). | |||||