Please be patient as the page loads
|
CRAL_HUMAN
|
||||||
| SwissProt Accessions | P12271 | Gene names | RLBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cellular retinaldehyde-binding protein (CRALBP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CRAL_HUMAN
|
||||||
| θ value | 3.4876e-161 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P12271 | Gene names | RLBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cellular retinaldehyde-binding protein (CRALBP). | |||||
|
CRAL_MOUSE
|
||||||
| θ value | 8.0589e-150 (rank : 2) | NC score | 0.997450 (rank : 2) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Z275 | Gene names | Rlbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Cellular retinaldehyde-binding protein (CRALBP). | |||||
|
CT121_MOUSE
|
||||||
| θ value | 1.12253e-42 (rank : 3) | NC score | 0.909859 (rank : 3) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D3D0, Q3U0R3, Q7TS94 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | Protein C20orf121 homolog. | |||||
|
CT121_HUMAN
|
||||||
| θ value | 1.46607e-42 (rank : 4) | NC score | 0.909603 (rank : 4) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BTX7, Q5QPC1, Q9H1G2, Q9NQG8 | Gene names | C20orf121 | |||
|
Domain Architecture |
|
|||||
| Description | Protein C20orf121. | |||||
|
TTPA_MOUSE
|
||||||
| θ value | 4.42448e-31 (rank : 5) | NC score | 0.883428 (rank : 6) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BWP5, Q9CW51, Q9JL07 | Gene names | Ttpa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-tocopherol transfer protein (Alpha-TTP). | |||||
|
TTPA_HUMAN
|
||||||
| θ value | 5.77852e-31 (rank : 6) | NC score | 0.884521 (rank : 5) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49638, Q71V64 | Gene names | TTPA, TPP1 | |||
|
Domain Architecture |
|
|||||
| Description | Alpha-tocopherol transfer protein (Alpha-TTP). | |||||
|
PTN9_HUMAN
|
||||||
| θ value | 7.58209e-15 (rank : 7) | NC score | 0.246141 (rank : 13) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P43378 | Gene names | PTPN9 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 9 (EC 3.1.3.48) (Protein-tyrosine phosphatase MEG2) (PTPase-MEG2). | |||||
|
PTN9_MOUSE
|
||||||
| θ value | 1.68911e-14 (rank : 8) | NC score | 0.243972 (rank : 14) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35239, Q7TSK0 | Gene names | Ptpn9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 9 (EC 3.1.3.48) (Protein-tyrosine phosphatase MEG2) (PTPase-MEG2). | |||||
|
S14L3_HUMAN
|
||||||
| θ value | 7.34386e-10 (rank : 9) | NC score | 0.546388 (rank : 7) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UDX4 | Gene names | SEC14L3, TAP2 | |||
|
Domain Architecture |
|
|||||
| Description | SEC14-like protein 3 (Tocopherol-associated protein 2). | |||||
|
S14L2_MOUSE
|
||||||
| θ value | 6.21693e-09 (rank : 10) | NC score | 0.519657 (rank : 8) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99J08 | Gene names | Sec14l2 | |||
|
Domain Architecture |
|
|||||
| Description | SEC14-like protein 2 (Alpha-tocopherol-associated protein) (TAP). | |||||
|
S14L2_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 11) | NC score | 0.513068 (rank : 11) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O76054, Q9ULN4 | Gene names | SEC14L2, KIAA1186 | |||
|
Domain Architecture |
|
|||||
| Description | SEC14-like protein 2 (Alpha-tocopherol-associated protein) (TAP) (hTAP) (Supernatant protein factor) (SPF) (Squalene transfer protein). | |||||
|
S14L4_MOUSE
|
||||||
| θ value | 2.36244e-08 (rank : 12) | NC score | 0.513590 (rank : 10) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8R0F9 | Gene names | Sec14l4 | |||
|
Domain Architecture |
|
|||||
| Description | SEC14-like protein 4. | |||||
|
S14L4_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 13) | NC score | 0.515811 (rank : 9) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UDX3 | Gene names | SEC14L4, TAP3 | |||
|
Domain Architecture |
|
|||||
| Description | SEC14-like protein 4 (Tocopherol-associated protein 3). | |||||
|
S14L1_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 14) | NC score | 0.468971 (rank : 12) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92503, Q99780 | Gene names | SEC14L1, SEC14L | |||
|
Domain Architecture |

|
|||||
| Description | SEC14-like protein 1. | |||||
|
GOGA5_HUMAN
|
||||||
| θ value | 0.279714 (rank : 15) | NC score | 0.018756 (rank : 27) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1168 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TBA6, O95287, Q03962, Q9UQQ7 | Gene names | GOLGA5, RETII, RFG5 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (RET-fused gene 5 protein) (Ret-II protein). | |||||
|
RHG01_HUMAN
|
||||||
| θ value | 0.813845 (rank : 16) | NC score | 0.028054 (rank : 23) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q07960 | Gene names | ARHGAP1, CDC42GAP, RHOGAP1 | |||
|
Domain Architecture |
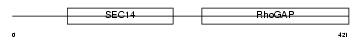
|
|||||
| Description | Rho-GTPase-activating protein 1 (GTPase-activating protein rhoOGAP) (Rho-related small GTPase protein activator) (CDC42 GTPase-activating protein) (p50-RhoGAP). | |||||
|
SMC3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.026362 (rank : 25) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1186 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UQE7, O60464 | Gene names | CSPG6, BAM, BMH, SMC3, SMC3L1 | |||
|
Domain Architecture |
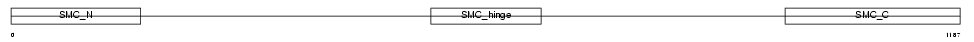
|
|||||
| Description | Structural maintenance of chromosome 3 (Chondroitin sulfate proteoglycan 6) (Chromosome-associated polypeptide) (hCAP) (Bamacan) (Basement membrane-associated chondroitin proteoglycan). | |||||
|
SMC3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.026412 (rank : 24) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1187 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9CW03, O35667, Q9QUS3 | Gene names | Cspg6, Bam, Bmh, Mmip1, Smc3, Smc3l1, Smcd | |||
|
Domain Architecture |
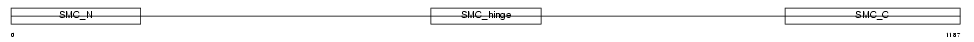
|
|||||
| Description | Structural maintenance of chromosome 3 (Chondroitin sulfate proteoglycan 6) (Chromosome segregation protein SmcD) (Bamacan) (Basement membrane-associated chondroitin proteoglycan) (Mad member- interacting protein 1). | |||||
|
EVPL_MOUSE
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.018960 (rank : 26) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
BNIP2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.052992 (rank : 20) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q12982 | Gene names | BNIP2, NIP2 | |||
|
Domain Architecture |
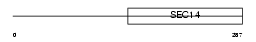
|
|||||
| Description | BCL2/adenovirus E1B 19 kDa protein-interacting protein 2. | |||||
|
ATCAY_HUMAN
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.068516 (rank : 15) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86WG3, Q8NAQ2, Q8TAQ3, Q96HC6, Q96JF5 | Gene names | ATCAY, KIAA1872 | |||
|
Domain Architecture |

|
|||||
| Description | Caytaxin (Ataxia Cayman type protein) (BNIP-H). | |||||
|
GOGA5_MOUSE
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.013863 (rank : 29) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
ATCAY_MOUSE
|
||||||
| θ value | 3.0926 (rank : 23) | NC score | 0.054038 (rank : 19) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BHE3, Q3TR94 | Gene names | Atcay | |||
|
Domain Architecture |
|
|||||
| Description | Caytaxin. | |||||
|
PXR_MOUSE
|
||||||
| θ value | 3.0926 (rank : 24) | NC score | 0.006956 (rank : 31) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O54915 | Gene names | Nr1i2, Pxr | |||
|
Domain Architecture |

|
|||||
| Description | Orphan nuclear receptor PXR (Pregnane X receptor). | |||||
|
BNIP2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.047518 (rank : 22) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54940 | Gene names | Bnip2, Nip2l | |||
|
Domain Architecture |
|
|||||
| Description | BCL2/adenovirus E1B 19 kDa protein-interacting protein 2. | |||||
|
TEKT1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.014760 (rank : 28) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 399 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DAJ2 | Gene names | Tekt1 | |||
|
Domain Architecture |

|
|||||
| Description | Tektin-1. | |||||
|
CE110_HUMAN
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.012030 (rank : 30) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43303, O43335, Q68DV9, Q8NE13 | Gene names | CEP110, CP110, KIAA0419 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 110 kDa (Cep110 protein). | |||||
|
CP135_MOUSE
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.005727 (rank : 32) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
GCP60_HUMAN
|
||||||
| θ value | θ > 10 (rank : 29) | NC score | 0.063379 (rank : 16) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H3P7, Q5VTJ0, Q6P9F1, Q8IZC5, Q8N4D6, Q9H6U3 | Gene names | ACBD3, GCP60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi resident protein GCP60 (Acyl-CoA-binding domain-containing protein 3) (Golgi phosphoprotein 1) (GOLPH1) (Golgi complex-associated protein 1) (GOCAP1) (PBR- and PKA-associated protein 7) (Peripheral benzodiazepine receptor-associated protein PAP7). | |||||
|
GCP60_MOUSE
|
||||||
| θ value | θ > 10 (rank : 30) | NC score | 0.059682 (rank : 17) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BMP6, O35371 | Gene names | Acbd3, Gcp60, Pap7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi resident protein GCP60 (Acyl-CoA-binding domain-containing protein 3) (Golgi phosphoprotein 1) (GOLPH1) (Golgi complex-associated protein 1) (GOCAP1) (PBR- and PKA-associated protein 7) (Peripheral benzodiazepine receptor-associated protein PAP7). | |||||
|
MSPD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 31) | NC score | 0.051904 (rank : 21) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CWP6, Q8BYF8, Q8BZB6, Q8C0G1, Q8R0T7 | Gene names | Mospd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Motile sperm domain-containing protein 2. | |||||
|
TMED8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 32) | NC score | 0.054726 (rank : 18) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PL24, Q9P1V9 | Gene names | TMED8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein TMED8. | |||||
|
CRAL_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 3.4876e-161 (rank : 1) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P12271 | Gene names | RLBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cellular retinaldehyde-binding protein (CRALBP). | |||||
|
CRAL_MOUSE
|
||||||
| NC score | 0.997450 (rank : 2) | θ value | 8.0589e-150 (rank : 2) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Z275 | Gene names | Rlbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Cellular retinaldehyde-binding protein (CRALBP). | |||||
|
CT121_MOUSE
|
||||||
| NC score | 0.909859 (rank : 3) | θ value | 1.12253e-42 (rank : 3) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D3D0, Q3U0R3, Q7TS94 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Protein C20orf121 homolog. | |||||
|
CT121_HUMAN
|
||||||
| NC score | 0.909603 (rank : 4) | θ value | 1.46607e-42 (rank : 4) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BTX7, Q5QPC1, Q9H1G2, Q9NQG8 | Gene names | C20orf121 | |||
|
Domain Architecture |

|
|||||
| Description | Protein C20orf121. | |||||
|
TTPA_HUMAN
|
||||||
| NC score | 0.884521 (rank : 5) | θ value | 5.77852e-31 (rank : 6) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49638, Q71V64 | Gene names | TTPA, TPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-tocopherol transfer protein (Alpha-TTP). | |||||
|
TTPA_MOUSE
|
||||||
| NC score | 0.883428 (rank : 6) | θ value | 4.42448e-31 (rank : 5) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BWP5, Q9CW51, Q9JL07 | Gene names | Ttpa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-tocopherol transfer protein (Alpha-TTP). | |||||
|
S14L3_HUMAN
|
||||||
| NC score | 0.546388 (rank : 7) | θ value | 7.34386e-10 (rank : 9) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UDX4 | Gene names | SEC14L3, TAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SEC14-like protein 3 (Tocopherol-associated protein 2). | |||||
|
S14L2_MOUSE
|
||||||
| NC score | 0.519657 (rank : 8) | θ value | 6.21693e-09 (rank : 10) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99J08 | Gene names | Sec14l2 | |||
|
Domain Architecture |

|
|||||
| Description | SEC14-like protein 2 (Alpha-tocopherol-associated protein) (TAP). | |||||
|
S14L4_HUMAN
|
||||||
| NC score | 0.515811 (rank : 9) | θ value | 4.0297e-08 (rank : 13) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UDX3 | Gene names | SEC14L4, TAP3 | |||
|
Domain Architecture |

|
|||||
| Description | SEC14-like protein 4 (Tocopherol-associated protein 3). | |||||
|
S14L4_MOUSE
|
||||||
| NC score | 0.513590 (rank : 10) | θ value | 2.36244e-08 (rank : 12) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8R0F9 | Gene names | Sec14l4 | |||
|
Domain Architecture |

|
|||||
| Description | SEC14-like protein 4. | |||||
|
S14L2_HUMAN
|
||||||
| NC score | 0.513068 (rank : 11) | θ value | 1.06045e-08 (rank : 11) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O76054, Q9ULN4 | Gene names | SEC14L2, KIAA1186 | |||
|
Domain Architecture |

|
|||||
| Description | SEC14-like protein 2 (Alpha-tocopherol-associated protein) (TAP) (hTAP) (Supernatant protein factor) (SPF) (Squalene transfer protein). | |||||
|
S14L1_HUMAN
|
||||||
| NC score | 0.468971 (rank : 12) | θ value | 2.21117e-06 (rank : 14) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92503, Q99780 | Gene names | SEC14L1, SEC14L | |||
|
Domain Architecture |

|
|||||
| Description | SEC14-like protein 1. | |||||
|
PTN9_HUMAN
|
||||||
| NC score | 0.246141 (rank : 13) | θ value | 7.58209e-15 (rank : 7) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P43378 | Gene names | PTPN9 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 9 (EC 3.1.3.48) (Protein-tyrosine phosphatase MEG2) (PTPase-MEG2). | |||||
|
PTN9_MOUSE
|
||||||
| NC score | 0.243972 (rank : 14) | θ value | 1.68911e-14 (rank : 8) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35239, Q7TSK0 | Gene names | Ptpn9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 9 (EC 3.1.3.48) (Protein-tyrosine phosphatase MEG2) (PTPase-MEG2). | |||||
|
ATCAY_HUMAN
|
||||||
| NC score | 0.068516 (rank : 15) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86WG3, Q8NAQ2, Q8TAQ3, Q96HC6, Q96JF5 | Gene names | ATCAY, KIAA1872 | |||
|
Domain Architecture |

|
|||||
| Description | Caytaxin (Ataxia Cayman type protein) (BNIP-H). | |||||
|
GCP60_HUMAN
|
||||||
| NC score | 0.063379 (rank : 16) | θ value | θ > 10 (rank : 29) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H3P7, Q5VTJ0, Q6P9F1, Q8IZC5, Q8N4D6, Q9H6U3 | Gene names | ACBD3, GCP60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi resident protein GCP60 (Acyl-CoA-binding domain-containing protein 3) (Golgi phosphoprotein 1) (GOLPH1) (Golgi complex-associated protein 1) (GOCAP1) (PBR- and PKA-associated protein 7) (Peripheral benzodiazepine receptor-associated protein PAP7). | |||||
|
GCP60_MOUSE
|
||||||
| NC score | 0.059682 (rank : 17) | θ value | θ > 10 (rank : 30) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BMP6, O35371 | Gene names | Acbd3, Gcp60, Pap7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi resident protein GCP60 (Acyl-CoA-binding domain-containing protein 3) (Golgi phosphoprotein 1) (GOLPH1) (Golgi complex-associated protein 1) (GOCAP1) (PBR- and PKA-associated protein 7) (Peripheral benzodiazepine receptor-associated protein PAP7). | |||||
|
TMED8_HUMAN
|
||||||
| NC score | 0.054726 (rank : 18) | θ value | θ > 10 (rank : 32) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PL24, Q9P1V9 | Gene names | TMED8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein TMED8. | |||||
|
ATCAY_MOUSE
|
||||||
| NC score | 0.054038 (rank : 19) | θ value | 3.0926 (rank : 23) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BHE3, Q3TR94 | Gene names | Atcay | |||
|
Domain Architecture |

|
|||||
| Description | Caytaxin. | |||||
|
BNIP2_HUMAN
|
||||||
| NC score | 0.052992 (rank : 20) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q12982 | Gene names | BNIP2, NIP2 | |||
|
Domain Architecture |

|
|||||
| Description | BCL2/adenovirus E1B 19 kDa protein-interacting protein 2. | |||||
|
MSPD2_MOUSE
|
||||||
| NC score | 0.051904 (rank : 21) | θ value | θ > 10 (rank : 31) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CWP6, Q8BYF8, Q8BZB6, Q8C0G1, Q8R0T7 | Gene names | Mospd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Motile sperm domain-containing protein 2. | |||||
|
BNIP2_MOUSE
|
||||||
| NC score | 0.047518 (rank : 22) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54940 | Gene names | Bnip2, Nip2l | |||
|
Domain Architecture |

|
|||||
| Description | BCL2/adenovirus E1B 19 kDa protein-interacting protein 2. | |||||
|
RHG01_HUMAN
|
||||||
| NC score | 0.028054 (rank : 23) | θ value | 0.813845 (rank : 16) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q07960 | Gene names | ARHGAP1, CDC42GAP, RHOGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 1 (GTPase-activating protein rhoOGAP) (Rho-related small GTPase protein activator) (CDC42 GTPase-activating protein) (p50-RhoGAP). | |||||
|
SMC3_MOUSE
|
||||||
| NC score | 0.026412 (rank : 24) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1187 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9CW03, O35667, Q9QUS3 | Gene names | Cspg6, Bam, Bmh, Mmip1, Smc3, Smc3l1, Smcd | |||
|
Domain Architecture |
|
|||||
| Description | Structural maintenance of chromosome 3 (Chondroitin sulfate proteoglycan 6) (Chromosome segregation protein SmcD) (Bamacan) (Basement membrane-associated chondroitin proteoglycan) (Mad member- interacting protein 1). | |||||
|
SMC3_HUMAN
|
||||||
| NC score | 0.026362 (rank : 25) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1186 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UQE7, O60464 | Gene names | CSPG6, BAM, BMH, SMC3, SMC3L1 | |||
|
Domain Architecture |
|
|||||
| Description | Structural maintenance of chromosome 3 (Chondroitin sulfate proteoglycan 6) (Chromosome-associated polypeptide) (hCAP) (Bamacan) (Basement membrane-associated chondroitin proteoglycan). | |||||
|
EVPL_MOUSE
|
||||||
| NC score | 0.018960 (rank : 26) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
GOGA5_HUMAN
|
||||||
| NC score | 0.018756 (rank : 27) | θ value | 0.279714 (rank : 15) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1168 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TBA6, O95287, Q03962, Q9UQQ7 | Gene names | GOLGA5, RETII, RFG5 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (RET-fused gene 5 protein) (Ret-II protein). | |||||
|
TEKT1_MOUSE
|
||||||
| NC score | 0.014760 (rank : 28) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 399 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DAJ2 | Gene names | Tekt1 | |||
|
Domain Architecture |

|
|||||
| Description | Tektin-1. | |||||
|
GOGA5_MOUSE
|
||||||
| NC score | 0.013863 (rank : 29) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
CE110_HUMAN
|
||||||
| NC score | 0.012030 (rank : 30) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43303, O43335, Q68DV9, Q8NE13 | Gene names | CEP110, CP110, KIAA0419 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 110 kDa (Cep110 protein). | |||||
|
PXR_MOUSE
|
||||||
| NC score | 0.006956 (rank : 31) | θ value | 3.0926 (rank : 24) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O54915 | Gene names | Nr1i2, Pxr | |||
|
Domain Architecture |

|
|||||
| Description | Orphan nuclear receptor PXR (Pregnane X receptor). | |||||
|
CP135_MOUSE
|
||||||
| NC score | 0.005727 (rank : 32) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||