Please be patient as the page loads
|
ACRBP_MOUSE
|
||||||
| SwissProt Accessions | Q3V140, Q62253, Q62254, Q8C621, Q91VQ1 | Gene names | Acrbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acrosin-binding protein precursor (Proacrosin-binding protein sp32). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ACRBP_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.959778 (rank : 2) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NEB7, Q9BY87 | Gene names | ACRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acrosin-binding protein precursor (Proacrosin-binding protein sp32) (Cancer testis antigen OY-TES-1). | |||||
|
ACRBP_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q3V140, Q62253, Q62254, Q8C621, Q91VQ1 | Gene names | Acrbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acrosin-binding protein precursor (Proacrosin-binding protein sp32). | |||||
|
GOGA6_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 3) | NC score | 0.066688 (rank : 4) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NYA3, Q9NYA7 | Gene names | GOLGA6, GLP | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 6 (Golgin linked to PML) (Golgin-like protein). | |||||
|
EVPL_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 4) | NC score | 0.058217 (rank : 7) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92817 | Gene names | EVPL | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (210 kDa paraneoplastic pemphigus antigen) (p210) (210 kDa cornified envelope precursor protein). | |||||
|
ROCK1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 5) | NC score | 0.025179 (rank : 57) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
ACINU_HUMAN
|
||||||
| θ value | 0.125558 (rank : 6) | NC score | 0.064134 (rank : 5) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
RFIP4_HUMAN
|
||||||
| θ value | 0.163984 (rank : 7) | NC score | 0.056229 (rank : 8) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q86YS3, Q8N829, Q8NDT7, Q969D8 | Gene names | RAB11FIP4, KIAA1821 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 4 (Rab11-FIP4). | |||||
|
ZNF96_HUMAN
|
||||||
| θ value | 0.21417 (rank : 8) | NC score | 0.004939 (rank : 72) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 869 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43309, O43724 | Gene names | ZNF96, KIAA0426, ZNF305, ZSCAN12 | |||
|
Domain Architecture |
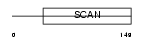
|
|||||
| Description | Zinc finger protein 96 (Zinc finger protein 305) (Zinc finger and SCAN domain-containing protein 12). | |||||
|
CKAP2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.062644 (rank : 6) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3V1H1, Q66LN2, Q8BSF0 | Gene names | Ckap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2. | |||||
|
ADDB_MOUSE
|
||||||
| θ value | 0.47712 (rank : 10) | NC score | 0.032070 (rank : 45) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
CYLN2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 11) | NC score | 0.047053 (rank : 12) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Z0H8, Q7TSI9, Q8CHU1, Q9EP81 | Gene names | Cyln2, Kiaa0291 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115). | |||||
|
TTF1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 12) | NC score | 0.049127 (rank : 10) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
UBP8_HUMAN
|
||||||
| θ value | 0.813845 (rank : 13) | NC score | 0.028830 (rank : 52) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 1.06291 (rank : 14) | NC score | 0.036058 (rank : 38) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
FLIP1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 15) | NC score | 0.044648 (rank : 22) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
GOGA1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.045295 (rank : 16) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
MYRIP_HUMAN
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.039881 (rank : 30) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NFW9, Q8IUF5, Q9Y3V4 | Gene names | MYRIP, SLAC2C | |||
|
Domain Architecture |
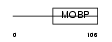
|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
RM32_MOUSE
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.055100 (rank : 9) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DCI9, Q91XR6 | Gene names | Mrpl32, Heg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L32, mitochondrial precursor (L32mt) (MRP-L32) (Heart expressed gene 1 protein). | |||||
|
SWP70_MOUSE
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.047706 (rank : 11) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
CK5P2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.044818 (rank : 20) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1151 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8K389, Q6PCN1, Q6ZPL0 | Gene names | Cdk5rap2, Kiaa1633 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48). | |||||
|
FBX34_MOUSE
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.031535 (rank : 46) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XI1, Q9D290 | Gene names | Fbxo34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 34. | |||||
|
GRAP1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.043455 (rank : 25) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1234 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VD04, O35693, Q3T9C3, Q69ZP9 | Gene names | Gripap1, DXImx47e, Kiaa1167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP1-associated protein 1 (GRASP-1) (HCMV-interacting protein). | |||||
|
INVO_MOUSE
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.090032 (rank : 3) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P48997 | Gene names | Ivl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Involucrin. | |||||
|
LRRC6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.029693 (rank : 49) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86X45, Q13648, Q4G183 | Gene names | LRRC6, LRTP, TSLRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 6 (Leucine-rich testis-specific protein) (Testis-specific leucine-rich repeat protein). | |||||
|
MDN1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.044263 (rank : 23) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
MYO5C_HUMAN
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.030613 (rank : 48) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
RAD50_MOUSE
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.045239 (rank : 17) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
RPTN_MOUSE
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.037663 (rank : 33) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97347 | Gene names | Rptn | |||
|
Domain Architecture |

|
|||||
| Description | Repetin. | |||||
|
SRC8_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.032672 (rank : 44) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14247 | Gene names | CTTN, EMS1 | |||
|
Domain Architecture |
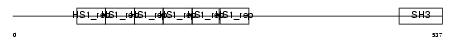
|
|||||
| Description | Src substrate cortactin (Amplaxin) (Oncogene EMS1). | |||||
|
SRC8_MOUSE
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.033153 (rank : 41) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60598 | Gene names | Cttn, Ems1 | |||
|
Domain Architecture |
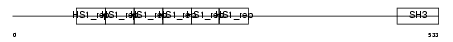
|
|||||
| Description | Src substrate cortactin. | |||||
|
STX3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.032968 (rank : 43) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13277, O43750, O43751, Q15360 | Gene names | STX3, STX3A | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-3. | |||||
|
STX3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.033148 (rank : 42) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q64704 | Gene names | Stx3, Stx3a | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-3. | |||||
|
SYNE2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.037491 (rank : 34) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
GCP6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.042682 (rank : 26) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96RT7, Q5JZ80, Q6PJ40, Q86YE9, Q9BY91, Q9UGX3, Q9UGX4 | Gene names | TUBGCP6, GCP6, KIAA1669 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-tubulin complex component 6 (GCP-6). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.041735 (rank : 28) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
SEN34_MOUSE
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.034723 (rank : 39) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BMZ5, Q58EU2, Q99LV6, Q9CYW1 | Gene names | Tsen34, Leng5, Sen34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen34 (EC 3.1.27.9) (tRNA-intron endonuclease Sen34) (Leukocyte receptor cluster member 5 homolog). | |||||
|
41_MOUSE
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.017550 (rank : 67) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48193, Q68FF1, Q6NVF5 | Gene names | Epb41, Epb4.1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein 4.1 (Band 4.1) (P4.1) (4.1R). | |||||
|
FOS_HUMAN
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.023175 (rank : 60) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P01100, P18849 | Gene names | FOS, G0S7 | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene protein c-fos (Cellular oncogene fos) (G0/G1 switch regulatory protein 7). | |||||
|
MY18A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.036220 (rank : 37) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.046502 (rank : 14) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
SDCG8_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.044744 (rank : 21) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q80UF4 | Gene names | Sdccag8, Cccap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (mCCCAP). | |||||
|
ZN434_HUMAN
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.002829 (rank : 73) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NX65, Q7Z6G2 | Gene names | ZNF434 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 434. | |||||
|
CCHCR_MOUSE
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.044924 (rank : 19) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 932 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8K2I2 | Gene names | Cchcr1, Hcr | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil alpha-helical rod protein 1 (Alpha helical coiled-coil rod protein). | |||||
|
CGRE1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.030913 (rank : 47) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R1U2, Q9D1K1 | Gene names | Cgref1, Cgr11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
CYLN2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.045009 (rank : 18) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
LAMB1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.020020 (rank : 66) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
LIMA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.042285 (rank : 27) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UHB6, Q2TAN7, Q9BVF2, Q9H8J1, Q9HBN5, Q9NX96, Q9NXC3, Q9NXU6, Q9P0H8, Q9UHB5 | Gene names | LIMA1, EPLIN, SREBP3 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm). | |||||
|
MY18A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.036635 (rank : 36) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
41_HUMAN
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.015874 (rank : 68) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11171, P11176, Q14245, Q5TB35, Q5VXN8, Q8IXV9, Q9Y578, Q9Y579 | Gene names | EPB41, E41P | |||
|
Domain Architecture |

|
|||||
| Description | Protein 4.1 (Band 4.1) (P4.1) (EPB4.1) (4.1R). | |||||
|
HMOX1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.020711 (rank : 65) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09601 | Gene names | HMOX1, HO, HO1 | |||
|
Domain Architecture |
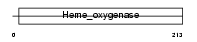
|
|||||
| Description | Heme oxygenase 1 (EC 1.14.99.3) (HO-1). | |||||
|
K1210_HUMAN
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.027023 (rank : 54) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
LIMA1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.026385 (rank : 55) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ERG0, Q9ERG1 | Gene names | Lima1, D15Ertd366e, Eplin | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm) (mEPLIN). | |||||
|
PLVAP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.046761 (rank : 13) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9BX97, Q86VP0, Q8N8Y0, Q8ND68, Q8TER8, Q9BZD5 | Gene names | PLVAP, FELS, PV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plasmalemma vesicle-associated protein (Plasmalemma vesicle protein 1) (PV-1) (Fenestrated endothelial-linked structure protein). | |||||
|
SAS6_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.038798 (rank : 31) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80UK7, Q9CYT4 | Gene names | Sass6, Sas6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spindle assembly abnormal protein 6 homolog. | |||||
|
TRRAP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.028413 (rank : 53) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y4A5, O75218, Q9Y631, Q9Y6H4 | Gene names | TRRAP, PAF400 | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (350/400 kDa PCAF-associated factor) (PAF350/400) (STAF40) (Tra1 homolog). | |||||
|
TRRAP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.028959 (rank : 51) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80YV3, Q8C0Z5, Q8K104 | Gene names | Trrap | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (Tra1 homolog). | |||||
|
CCD21_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.045301 (rank : 15) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BMK0, Q8BUF1, Q8K0E6 | Gene names | Ccdc21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
CENPC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.020798 (rank : 64) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q03188, Q9P0M5 | Gene names | CENPC1, CENPC, ICEN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C) (Interphase centromere complex protein 7). | |||||
|
GLE1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.037210 (rank : 35) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8R322, Q3TT10, Q3TU23, Q3UD65, Q8BT16, Q9D4A6 | Gene names | Gle1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin GLE1 (GLE1-like protein). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.024732 (rank : 59) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.041429 (rank : 29) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
OXR1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.020855 (rank : 63) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
PLCB2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.022794 (rank : 62) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q00722 | Gene names | PLCB2 | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta 2 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (Phospholipase C- beta-2) (PLC-beta-2). | |||||
|
RGNEF_MOUSE
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.026350 (rank : 56) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
RPTN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.033449 (rank : 40) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
SYAM_MOUSE
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.014889 (rank : 69) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14CH7, Q68FH3, Q69ZM9 | Gene names | Aarsl, Aars2, Kiaa1270 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable alanyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.7) (Alanine--tRNA ligase) (AlaRS) (Alanyl-tRNA synthetase like). | |||||
|
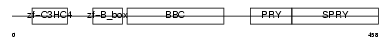
TRI39_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.012670 (rank : 70) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HCM9, Q76BL3, Q8IYT9, Q96IB6 | Gene names | TRIM39, RNF23, TFP | |||
|
Domain Architecture |
|
|||||
| Description | Tripartite motif-containing protein 39 (RING finger protein 23) (Testis-abundant finger protein). | |||||
|
CFAI_HUMAN
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.002591 (rank : 74) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05156, O60442 | Gene names | CFI, IF | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor I precursor (EC 3.4.21.45) (C3B/C4B inactivator) [Contains: Complement factor I heavy chain; Complement factor I light chain]. | |||||
|
CL004_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.022829 (rank : 61) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQ89, Q6MZH5 | Gene names | C12orf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C12orf4. | |||||
|
CL004_MOUSE
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.024914 (rank : 58) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91YN0, Q3UVK4 | Gene names | D6Wsu163e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C12orf4 homolog. | |||||
|
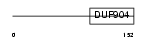
DDIT3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.029623 (rank : 50) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |
|
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
LRRK1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.001016 (rank : 75) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1267 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3UHC2, Q3U476, Q66JQ4, Q6GQR9, Q6NZF5, Q6ZPI4, Q8BKP3, Q8BU93, Q8BUY0, Q8BVV2, Q8R085 | Gene names | Lrrk1, Kiaa1790 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 1 (EC 2.7.11.1). | |||||
|
PCNT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.037994 (rank : 32) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
RAD50_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.043877 (rank : 24) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
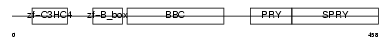
TRI39_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.012246 (rank : 71) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 691 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9ESN2, Q8BPR5, Q8K0F7 | Gene names | Trim39, Rnf23, Tfp | |||
|
Domain Architecture |
|
|||||
| Description | Tripartite motif-containing protein 39 (RING finger protein 23) (Testis-abundant finger protein). | |||||
|
ACRBP_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q3V140, Q62253, Q62254, Q8C621, Q91VQ1 | Gene names | Acrbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acrosin-binding protein precursor (Proacrosin-binding protein sp32). | |||||
|
ACRBP_HUMAN
|
||||||
| NC score | 0.959778 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NEB7, Q9BY87 | Gene names | ACRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acrosin-binding protein precursor (Proacrosin-binding protein sp32) (Cancer testis antigen OY-TES-1). | |||||
|
INVO_MOUSE
|
||||||
| NC score | 0.090032 (rank : 3) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P48997 | Gene names | Ivl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Involucrin. | |||||
|
GOGA6_HUMAN
|
||||||
| NC score | 0.066688 (rank : 4) | θ value | 0.0431538 (rank : 3) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NYA3, Q9NYA7 | Gene names | GOLGA6, GLP | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 6 (Golgin linked to PML) (Golgin-like protein). | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.064134 (rank : 5) | θ value | 0.125558 (rank : 6) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
CKAP2_MOUSE
|
||||||
| NC score | 0.062644 (rank : 6) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3V1H1, Q66LN2, Q8BSF0 | Gene names | Ckap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2. | |||||
|
EVPL_HUMAN
|
||||||
| NC score | 0.058217 (rank : 7) | θ value | 0.0736092 (rank : 4) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92817 | Gene names | EVPL | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (210 kDa paraneoplastic pemphigus antigen) (p210) (210 kDa cornified envelope precursor protein). | |||||
|
RFIP4_HUMAN
|
||||||
| NC score | 0.056229 (rank : 8) | θ value | 0.163984 (rank : 7) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q86YS3, Q8N829, Q8NDT7, Q969D8 | Gene names | RAB11FIP4, KIAA1821 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 4 (Rab11-FIP4). | |||||
|
RM32_MOUSE
|
||||||
| NC score | 0.055100 (rank : 9) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DCI9, Q91XR6 | Gene names | Mrpl32, Heg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L32, mitochondrial precursor (L32mt) (MRP-L32) (Heart expressed gene 1 protein). | |||||
|
TTF1_MOUSE
|
||||||
| NC score | 0.049127 (rank : 10) | θ value | 0.813845 (rank : 12) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
SWP70_MOUSE
|
||||||
| NC score | 0.047706 (rank : 11) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
CYLN2_MOUSE
|
||||||
| NC score | 0.047053 (rank : 12) | θ value | 0.813845 (rank : 11) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Z0H8, Q7TSI9, Q8CHU1, Q9EP81 | Gene names | Cyln2, Kiaa0291 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115). | |||||
|
PLVAP_HUMAN
|
||||||
| NC score | 0.046761 (rank : 13) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9BX97, Q86VP0, Q8N8Y0, Q8ND68, Q8TER8, Q9BZD5 | Gene names | PLVAP, FELS, PV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plasmalemma vesicle-associated protein (Plasmalemma vesicle protein 1) (PV-1) (Fenestrated endothelial-linked structure protein). | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.046502 (rank : 14) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
CCD21_MOUSE
|
||||||
| NC score | 0.045301 (rank : 15) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BMK0, Q8BUF1, Q8K0E6 | Gene names | Ccdc21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
GOGA1_HUMAN
|
||||||
| NC score | 0.045295 (rank : 16) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
RAD50_MOUSE
|
||||||
| NC score | 0.045239 (rank : 17) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
CYLN2_HUMAN
|
||||||
| NC score | 0.045009 (rank : 18) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
CCHCR_MOUSE
|
||||||
| NC score | 0.044924 (rank : 19) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 932 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8K2I2 | Gene names | Cchcr1, Hcr | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil alpha-helical rod protein 1 (Alpha helical coiled-coil rod protein). | |||||
|
CK5P2_MOUSE
|
||||||
| NC score | 0.044818 (rank : 20) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1151 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8K389, Q6PCN1, Q6ZPL0 | Gene names | Cdk5rap2, Kiaa1633 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48). | |||||
|
SDCG8_MOUSE
|
||||||
| NC score | 0.044744 (rank : 21) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q80UF4 | Gene names | Sdccag8, Cccap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (mCCCAP). | |||||
|
FLIP1_HUMAN
|
||||||
| NC score | 0.044648 (rank : 22) | θ value | 1.06291 (rank : 15) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
MDN1_HUMAN
|
||||||
| NC score | 0.044263 (rank : 23) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
RAD50_HUMAN
|
||||||
| NC score | 0.043877 (rank : 24) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
GRAP1_MOUSE
|
||||||
| NC score | 0.043455 (rank : 25) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1234 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VD04, O35693, Q3T9C3, Q69ZP9 | Gene names | Gripap1, DXImx47e, Kiaa1167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP1-associated protein 1 (GRASP-1) (HCMV-interacting protein). | |||||
|
GCP6_HUMAN
|
||||||
| NC score | 0.042682 (rank : 26) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96RT7, Q5JZ80, Q6PJ40, Q86YE9, Q9BY91, Q9UGX3, Q9UGX4 | Gene names | TUBGCP6, GCP6, KIAA1669 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-tubulin complex component 6 (GCP-6). | |||||
|
LIMA1_HUMAN
|
||||||
| NC score | 0.042285 (rank : 27) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UHB6, Q2TAN7, Q9BVF2, Q9H8J1, Q9HBN5, Q9NX96, Q9NXC3, Q9NXU6, Q9P0H8, Q9UHB5 | Gene names | LIMA1, EPLIN, SREBP3 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.041735 (rank : 28) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.041429 (rank : 29) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
MYRIP_HUMAN
|
||||||
| NC score | 0.039881 (rank : 30) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NFW9, Q8IUF5, Q9Y3V4 | Gene names | MYRIP, SLAC2C | |||
|
Domain Architecture |

|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
SAS6_MOUSE
|
||||||
| NC score | 0.038798 (rank : 31) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80UK7, Q9CYT4 | Gene names | Sass6, Sas6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spindle assembly abnormal protein 6 homolog. | |||||
|
PCNT_HUMAN
|
||||||
| NC score | 0.037994 (rank : 32) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
RPTN_MOUSE
|
||||||
| NC score | 0.037663 (rank : 33) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97347 | Gene names | Rptn | |||
|
Domain Architecture |

|
|||||
| Description | Repetin. | |||||
|
SYNE2_HUMAN
|
||||||
| NC score | 0.037491 (rank : 34) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
GLE1_MOUSE
|
||||||
| NC score | 0.037210 (rank : 35) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8R322, Q3TT10, Q3TU23, Q3UD65, Q8BT16, Q9D4A6 | Gene names | Gle1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin GLE1 (GLE1-like protein). | |||||
|
MY18A_HUMAN
|
||||||
| NC score | 0.036635 (rank : 36) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MY18A_MOUSE
|
||||||
| NC score | 0.036220 (rank : 37) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.036058 (rank : 38) | θ value | 1.06291 (rank : 14) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
SEN34_MOUSE
|
||||||
| NC score | 0.034723 (rank : 39) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BMZ5, Q58EU2, Q99LV6, Q9CYW1 | Gene names | Tsen34, Leng5, Sen34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-splicing endonuclease subunit Sen34 (EC 3.1.27.9) (tRNA-intron endonuclease Sen34) (Leukocyte receptor cluster member 5 homolog). | |||||
|
RPTN_HUMAN
|
||||||
| NC score | 0.033449 (rank : 40) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
SRC8_MOUSE
|
||||||
| NC score | 0.033153 (rank : 41) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60598 | Gene names | Cttn, Ems1 | |||
|
Domain Architecture |
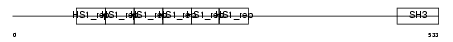
|
|||||
| Description | Src substrate cortactin. | |||||
|
STX3_MOUSE
|
||||||
| NC score | 0.033148 (rank : 42) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q64704 | Gene names | Stx3, Stx3a | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-3. | |||||
|
STX3_HUMAN
|
||||||
| NC score | 0.032968 (rank : 43) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13277, O43750, O43751, Q15360 | Gene names | STX3, STX3A | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-3. | |||||
|
SRC8_HUMAN
|
||||||
| NC score | 0.032672 (rank : 44) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14247 | Gene names | CTTN, EMS1 | |||
|
Domain Architecture |
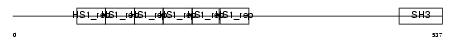
|
|||||
| Description | Src substrate cortactin (Amplaxin) (Oncogene EMS1). | |||||
|
ADDB_MOUSE
|
||||||
| NC score | 0.032070 (rank : 45) | θ value | 0.47712 (rank : 10) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
FBX34_MOUSE
|
||||||
| NC score | 0.031535 (rank : 46) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XI1, Q9D290 | Gene names | Fbxo34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 34. | |||||
|
CGRE1_MOUSE
|
||||||
| NC score | 0.030913 (rank : 47) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R1U2, Q9D1K1 | Gene names | Cgref1, Cgr11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
MYO5C_HUMAN
|
||||||
| NC score | 0.030613 (rank : 48) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
LRRC6_HUMAN
|
||||||
| NC score | 0.029693 (rank : 49) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86X45, Q13648, Q4G183 | Gene names | LRRC6, LRTP, TSLRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 6 (Leucine-rich testis-specific protein) (Testis-specific leucine-rich repeat protein). | |||||
|
DDIT3_HUMAN
|
||||||
| NC score | 0.029623 (rank : 50) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |

|
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
TRRAP_MOUSE
|
||||||
| NC score | 0.028959 (rank : 51) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80YV3, Q8C0Z5, Q8K104 | Gene names | Trrap | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (Tra1 homolog). | |||||
|
UBP8_HUMAN
|
||||||
| NC score | 0.028830 (rank : 52) | θ value | 0.813845 (rank : 13) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
TRRAP_HUMAN
|
||||||
| NC score | 0.028413 (rank : 53) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y4A5, O75218, Q9Y631, Q9Y6H4 | Gene names | TRRAP, PAF400 | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (350/400 kDa PCAF-associated factor) (PAF350/400) (STAF40) (Tra1 homolog). | |||||
|
K1210_HUMAN
|
||||||
| NC score | 0.027023 (rank : 54) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
LIMA1_MOUSE
|
||||||
| NC score | 0.026385 (rank : 55) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ERG0, Q9ERG1 | Gene names | Lima1, D15Ertd366e, Eplin | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm) (mEPLIN). | |||||
|
RGNEF_MOUSE
|
||||||
| NC score | 0.026350 (rank : 56) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
ROCK1_MOUSE
|
||||||
| NC score | 0.025179 (rank : 57) | θ value | 0.0961366 (rank : 5) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
CL004_MOUSE
|
||||||
| NC score | 0.024914 (rank : 58) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91YN0, Q3UVK4 | Gene names | D6Wsu163e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C12orf4 homolog. | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.024732 (rank : 59) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
FOS_HUMAN
|
||||||
| NC score | 0.023175 (rank : 60) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P01100, P18849 | Gene names | FOS, G0S7 | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene protein c-fos (Cellular oncogene fos) (G0/G1 switch regulatory protein 7). | |||||
|
CL004_HUMAN
|
||||||
| NC score | 0.022829 (rank : 61) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQ89, Q6MZH5 | Gene names | C12orf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C12orf4. | |||||
|
PLCB2_HUMAN
|
||||||
| NC score | 0.022794 (rank : 62) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q00722 | Gene names | PLCB2 | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta 2 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (Phospholipase C- beta-2) (PLC-beta-2). | |||||
|
OXR1_MOUSE
|
||||||
| NC score | 0.020855 (rank : 63) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
CENPC_HUMAN
|
||||||
| NC score | 0.020798 (rank : 64) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q03188, Q9P0M5 | Gene names | CENPC1, CENPC, ICEN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C) (Interphase centromere complex protein 7). | |||||
|
HMOX1_HUMAN
|
||||||
| NC score | 0.020711 (rank : 65) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09601 | Gene names | HMOX1, HO, HO1 | |||
|
Domain Architecture |
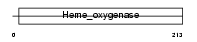
|
|||||
| Description | Heme oxygenase 1 (EC 1.14.99.3) (HO-1). | |||||
|
LAMB1_HUMAN
|
||||||
| NC score | 0.020020 (rank : 66) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
41_MOUSE
|
||||||
| NC score | 0.017550 (rank : 67) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48193, Q68FF1, Q6NVF5 | Gene names | Epb41, Epb4.1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein 4.1 (Band 4.1) (P4.1) (4.1R). | |||||
|
41_HUMAN
|
||||||
| NC score | 0.015874 (rank : 68) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11171, P11176, Q14245, Q5TB35, Q5VXN8, Q8IXV9, Q9Y578, Q9Y579 | Gene names | EPB41, E41P | |||
|
Domain Architecture |

|
|||||
| Description | Protein 4.1 (Band 4.1) (P4.1) (EPB4.1) (4.1R). | |||||
|
SYAM_MOUSE
|
||||||
| NC score | 0.014889 (rank : 69) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14CH7, Q68FH3, Q69ZM9 | Gene names | Aarsl, Aars2, Kiaa1270 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable alanyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.7) (Alanine--tRNA ligase) (AlaRS) (Alanyl-tRNA synthetase like). | |||||
|
TRI39_HUMAN
|
||||||
| NC score | 0.012670 (rank : 70) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HCM9, Q76BL3, Q8IYT9, Q96IB6 | Gene names | TRIM39, RNF23, TFP | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 39 (RING finger protein 23) (Testis-abundant finger protein). | |||||
|
TRI39_MOUSE
|
||||||
| NC score | 0.012246 (rank : 71) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 691 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9ESN2, Q8BPR5, Q8K0F7 | Gene names | Trim39, Rnf23, Tfp | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 39 (RING finger protein 23) (Testis-abundant finger protein). | |||||
|
ZNF96_HUMAN
|
||||||
| NC score | 0.004939 (rank : 72) | θ value | 0.21417 (rank : 8) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 869 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43309, O43724 | Gene names | ZNF96, KIAA0426, ZNF305, ZSCAN12 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 96 (Zinc finger protein 305) (Zinc finger and SCAN domain-containing protein 12). | |||||
|
ZN434_HUMAN
|
||||||
| NC score | 0.002829 (rank : 73) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NX65, Q7Z6G2 | Gene names | ZNF434 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 434. | |||||
|
CFAI_HUMAN
|
||||||
| NC score | 0.002591 (rank : 74) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05156, O60442 | Gene names | CFI, IF | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor I precursor (EC 3.4.21.45) (C3B/C4B inactivator) [Contains: Complement factor I heavy chain; Complement factor I light chain]. | |||||
|
LRRK1_MOUSE
|
||||||
| NC score | 0.001016 (rank : 75) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1267 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3UHC2, Q3U476, Q66JQ4, Q6GQR9, Q6NZF5, Q6ZPI4, Q8BKP3, Q8BU93, Q8BUY0, Q8BVV2, Q8R085 | Gene names | Lrrk1, Kiaa1790 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 1 (EC 2.7.11.1). | |||||