Please be patient as the page loads
|
SURF2_MOUSE
|
||||||
| SwissProt Accessions | P09926 | Gene names | Surf2, Surf-2 | |||
|
Domain Architecture |
|
|||||
| Description | Surfeit locus protein 2 (Surf-2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SURF2_MOUSE
|
||||||
| θ value | 8.63019e-144 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P09926 | Gene names | Surf2, Surf-2 | |||
|
Domain Architecture |

|
|||||
| Description | Surfeit locus protein 2 (Surf-2). | |||||
|
SURF2_HUMAN
|
||||||
| θ value | 1.88829e-90 (rank : 2) | NC score | 0.978261 (rank : 2) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15527, Q96CD1 | Gene names | SURF2 | |||
|
Domain Architecture |
|
|||||
| Description | Surfeit locus protein 2 (Surf-2). | |||||
|
PTTG_HUMAN
|
||||||
| θ value | 0.163984 (rank : 3) | NC score | 0.126999 (rank : 3) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P53801, Q9NS09 | Gene names | PTTG1IP, C21orf1, C21orf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pituitary tumor-transforming gene 1 protein-interacting protein precursor (Pituitary tumor-transforming gene protein-binding factor) (PTTG-binding factor) (PBF). | |||||
|
CDC37_HUMAN
|
||||||
| θ value | 0.365318 (rank : 4) | NC score | 0.050220 (rank : 5) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |
|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
PRGC2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 5) | NC score | 0.049457 (rank : 6) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
HS90A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 6) | NC score | 0.042318 (rank : 9) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07901 | Gene names | Hsp90aa1, Hsp86, Hsp86-1, Hspca | |||
|
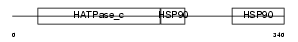
Domain Architecture |
|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (Tumor-specific transplantation 86 kDa antigen) (TSTA). | |||||
|
JADE1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 7) | NC score | 0.029186 (rank : 11) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZPI0, Q6IE84, Q8C7S6, Q8CFM2, Q8R362 | Gene names | Phf17, Jade1, Kiaa1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
TAF3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 8) | NC score | 0.042837 (rank : 8) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
PTTG_MOUSE
|
||||||
| θ value | 1.81305 (rank : 9) | NC score | 0.099365 (rank : 4) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R143, Q3TVT1, Q8BJ96, Q8N7P0 | Gene names | Pttg1ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pituitary tumor-transforming gene 1 protein-interacting protein precursor (Pituitary tumor-transforming gene protein-binding factor) (PTTG-binding factor) (PBF). | |||||
|
SEC62_MOUSE
|
||||||
| θ value | 2.36792 (rank : 10) | NC score | 0.044566 (rank : 7) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BU14, Q6NX81 | Gene names | Tloc1, Sec62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Translocation protein SEC62 (Translocation protein 1) (TP-1). | |||||
|
HS90A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 11) | NC score | 0.036534 (rank : 10) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07900, Q2PP14, Q9BVQ5 | Gene names | HSP90AA1, HSP90A, HSPC1, HSPCA | |||
|
Domain Architecture |
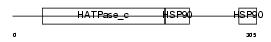
|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (NY-REN-38 antigen). | |||||
|
ZN384_HUMAN
|
||||||
| θ value | 4.03905 (rank : 12) | NC score | 0.002169 (rank : 22) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TF68, O15407, Q8N938 | Gene names | ZNF384, CAGH1, CIZ, NMP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 384 (Nuclear matrix transcription factor 4) (Nuclear matrix protein 4) (CAG repeat protein 1) (CAS-interacting zinc finger protein). | |||||
|
SFR12_MOUSE
|
||||||
| θ value | 5.27518 (rank : 13) | NC score | 0.019034 (rank : 13) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
RYR1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 14) | NC score | 0.014307 (rank : 17) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P21817, Q16314, Q16368, Q9NPK1, Q9P1U4 | Gene names | RYR1, RYDR | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 1 (Skeletal muscle-type ryanodine receptor) (RyR1) (RYR-1) (Skeletal muscle calcium release channel). | |||||
|
TNIK_HUMAN
|
||||||
| θ value | 6.88961 (rank : 15) | NC score | 0.001768 (rank : 23) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
TNNT3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 16) | NC score | 0.015275 (rank : 15) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QZ47, O35575, O35576, O35577, O35578, O35579, O35580, O35581, O35582, O35583, O35584, O35585, P97456, Q99L89, Q9QZ46 | Gene names | Tnnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Troponin T, fast skeletal muscle (TnTf) (Fast skeletal muscle troponin T) (fTnT). | |||||
|
AKAP9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 17) | NC score | 0.005403 (rank : 20) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
CEP35_HUMAN
|
||||||
| θ value | 8.99809 (rank : 18) | NC score | 0.014418 (rank : 16) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
DGKD_HUMAN
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.009718 (rank : 19) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q16760, Q14158, Q6PK55, Q8NG53 | Gene names | DGKD, KIAA0145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase delta (EC 2.7.1.107) (Diglyceride kinase delta) (DGK-delta) (DAG kinase delta) (130 kDa diacylglycerol kinase). | |||||
|
IPP2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 20) | NC score | 0.020188 (rank : 12) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DCL8, Q3UD25, Q8C1A6 | Gene names | Ppp1r2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase inhibitor 2 (IPP-2). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 21) | NC score | 0.013438 (rank : 18) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
SFR17_HUMAN
|
||||||
| θ value | 8.99809 (rank : 22) | NC score | 0.018900 (rank : 14) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
ZN486_HUMAN
|
||||||
| θ value | 8.99809 (rank : 23) | NC score | 0.002355 (rank : 21) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96H40 | Gene names | ZNF486, KRBO2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 486. | |||||
|
SURF2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 8.63019e-144 (rank : 1) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P09926 | Gene names | Surf2, Surf-2 | |||
|
Domain Architecture |

|
|||||
| Description | Surfeit locus protein 2 (Surf-2). | |||||
|
SURF2_HUMAN
|
||||||
| NC score | 0.978261 (rank : 2) | θ value | 1.88829e-90 (rank : 2) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15527, Q96CD1 | Gene names | SURF2 | |||
|
Domain Architecture |

|
|||||
| Description | Surfeit locus protein 2 (Surf-2). | |||||
|
PTTG_HUMAN
|
||||||
| NC score | 0.126999 (rank : 3) | θ value | 0.163984 (rank : 3) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P53801, Q9NS09 | Gene names | PTTG1IP, C21orf1, C21orf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pituitary tumor-transforming gene 1 protein-interacting protein precursor (Pituitary tumor-transforming gene protein-binding factor) (PTTG-binding factor) (PBF). | |||||
|
PTTG_MOUSE
|
||||||
| NC score | 0.099365 (rank : 4) | θ value | 1.81305 (rank : 9) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R143, Q3TVT1, Q8BJ96, Q8N7P0 | Gene names | Pttg1ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pituitary tumor-transforming gene 1 protein-interacting protein precursor (Pituitary tumor-transforming gene protein-binding factor) (PTTG-binding factor) (PBF). | |||||
|
CDC37_HUMAN
|
||||||
| NC score | 0.050220 (rank : 5) | θ value | 0.365318 (rank : 4) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |

|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
PRGC2_MOUSE
|
||||||
| NC score | 0.049457 (rank : 6) | θ value | 0.62314 (rank : 5) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
SEC62_MOUSE
|
||||||
| NC score | 0.044566 (rank : 7) | θ value | 2.36792 (rank : 10) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BU14, Q6NX81 | Gene names | Tloc1, Sec62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Translocation protein SEC62 (Translocation protein 1) (TP-1). | |||||
|
TAF3_MOUSE
|
||||||
| NC score | 0.042837 (rank : 8) | θ value | 0.813845 (rank : 8) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
HS90A_MOUSE
|
||||||
| NC score | 0.042318 (rank : 9) | θ value | 0.813845 (rank : 6) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07901 | Gene names | Hsp90aa1, Hsp86, Hsp86-1, Hspca | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (Tumor-specific transplantation 86 kDa antigen) (TSTA). | |||||
|
HS90A_HUMAN
|
||||||
| NC score | 0.036534 (rank : 10) | θ value | 4.03905 (rank : 11) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07900, Q2PP14, Q9BVQ5 | Gene names | HSP90AA1, HSP90A, HSPC1, HSPCA | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (NY-REN-38 antigen). | |||||
|
JADE1_MOUSE
|
||||||
| NC score | 0.029186 (rank : 11) | θ value | 0.813845 (rank : 7) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZPI0, Q6IE84, Q8C7S6, Q8CFM2, Q8R362 | Gene names | Phf17, Jade1, Kiaa1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
IPP2_MOUSE
|
||||||
| NC score | 0.020188 (rank : 12) | θ value | 8.99809 (rank : 20) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DCL8, Q3UD25, Q8C1A6 | Gene names | Ppp1r2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase inhibitor 2 (IPP-2). | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.019034 (rank : 13) | θ value | 5.27518 (rank : 13) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.018900 (rank : 14) | θ value | 8.99809 (rank : 22) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
TNNT3_MOUSE
|
||||||
| NC score | 0.015275 (rank : 15) | θ value | 6.88961 (rank : 16) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QZ47, O35575, O35576, O35577, O35578, O35579, O35580, O35581, O35582, O35583, O35584, O35585, P97456, Q99L89, Q9QZ46 | Gene names | Tnnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Troponin T, fast skeletal muscle (TnTf) (Fast skeletal muscle troponin T) (fTnT). | |||||
|
CEP35_HUMAN
|
||||||
| NC score | 0.014418 (rank : 16) | θ value | 8.99809 (rank : 18) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
RYR1_HUMAN
|
||||||
| NC score | 0.014307 (rank : 17) | θ value | 6.88961 (rank : 14) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P21817, Q16314, Q16368, Q9NPK1, Q9P1U4 | Gene names | RYR1, RYDR | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 1 (Skeletal muscle-type ryanodine receptor) (RyR1) (RYR-1) (Skeletal muscle calcium release channel). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.013438 (rank : 18) | θ value | 8.99809 (rank : 21) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
DGKD_HUMAN
|
||||||
| NC score | 0.009718 (rank : 19) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q16760, Q14158, Q6PK55, Q8NG53 | Gene names | DGKD, KIAA0145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase delta (EC 2.7.1.107) (Diglyceride kinase delta) (DGK-delta) (DAG kinase delta) (130 kDa diacylglycerol kinase). | |||||
|
AKAP9_HUMAN
|
||||||
| NC score | 0.005403 (rank : 20) | θ value | 8.99809 (rank : 17) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
ZN486_HUMAN
|
||||||
| NC score | 0.002355 (rank : 21) | θ value | 8.99809 (rank : 23) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96H40 | Gene names | ZNF486, KRBO2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 486. | |||||
|
ZN384_HUMAN
|
||||||
| NC score | 0.002169 (rank : 22) | θ value | 4.03905 (rank : 12) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TF68, O15407, Q8N938 | Gene names | ZNF384, CAGH1, CIZ, NMP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 384 (Nuclear matrix transcription factor 4) (Nuclear matrix protein 4) (CAG repeat protein 1) (CAS-interacting zinc finger protein). | |||||
|
TNIK_HUMAN
|
||||||
| NC score | 0.001768 (rank : 23) | θ value | 6.88961 (rank : 15) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||