Please be patient as the page loads
|
EF2_HUMAN
|
||||||
| SwissProt Accessions | P13639 | Gene names | EEF2, EF2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 2 (EF-2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
EF2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P13639 | Gene names | EEF2, EF2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 2 (EF-2). | |||||
|
EF2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999009 (rank : 2) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P58252, Q544E4 | Gene names | Eef2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 2 (EF-2). | |||||
|
U5S1_MOUSE
|
||||||
| θ value | 1.77876e-181 (rank : 3) | NC score | 0.950023 (rank : 4) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O08810 | Gene names | Eftud2, Snrp116 | |||
|
Domain Architecture |

|
|||||
| Description | 116 kDa U5 small nuclear ribonucleoprotein component (U5 snRNP- specific protein, 116 kDa) (U5-116 kDa) (Elongation factor Tu GTP- binding domain protein 2). | |||||
|
U5S1_HUMAN
|
||||||
| θ value | 3.96267e-181 (rank : 4) | NC score | 0.950196 (rank : 3) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15029, Q9BUR0 | Gene names | EFTUD2, KIAA0031, SNRP116 | |||
|
Domain Architecture |

|
|||||
| Description | 116 kDa U5 small nuclear ribonucleoprotein component (U5 snRNP- specific protein, 116 kDa) (U5-116 kDa) (Elongation factor Tu GTP- binding domain protein 2) (hSNU114). | |||||
|
EFG2_MOUSE
|
||||||
| θ value | 9.8567e-31 (rank : 5) | NC score | 0.712637 (rank : 5) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R2Q4 | Gene names | Gfm2, Efg2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 2, mitochondrial precursor (mEF-G 2) (Elongation factor G2). | |||||
|
EFG2_HUMAN
|
||||||
| θ value | 1.4233e-29 (rank : 6) | NC score | 0.681178 (rank : 6) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q969S9, Q8WYI0 | Gene names | GFM2, EFG2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 2, mitochondrial precursor (mEF-G 2) (Elongation factor G2). | |||||
|
EFG1_HUMAN
|
||||||
| θ value | 8.36355e-22 (rank : 7) | NC score | 0.637180 (rank : 8) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96RP9, Q96T39 | Gene names | GFM1, EFG, EFG1, GFM | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 1, mitochondrial precursor (mEF-G 1) (Elongation factor G1). | |||||
|
EFG1_MOUSE
|
||||||
| θ value | 2.69047e-20 (rank : 8) | NC score | 0.646344 (rank : 7) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K0D5, Q921D6, Q924I0 | Gene names | Gfm1, Efg, Efg1, Gfm | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 1, mitochondrial precursor (mEF-G 1) (Elongation factor G1). | |||||
|
EF1A1_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 9) | NC score | 0.307951 (rank : 11) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P68104, P04719, P04720 | Gene names | EEF1A1, EEF1A, EF1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor 1-alpha 1 (EF-1-alpha-1) (Elongation factor 1 A-1) (eEF1A-1) (Elongation factor Tu) (EF-Tu). | |||||
|
EF1A1_MOUSE
|
||||||
| θ value | 3.41135e-07 (rank : 10) | NC score | 0.307939 (rank : 12) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P10126, Q61511, Q6ZWN2, Q8BMB8, Q8BVS8, Q99KU5 | Gene names | Eef1a1, Eef1a | |||
|
Domain Architecture |
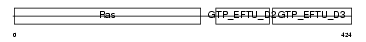
|
|||||
| Description | Elongation factor 1-alpha 1 (EF-1-alpha-1) (Elongation factor 1 A-1) (eEF1A-1) (Elongation factor Tu) (EF-Tu). | |||||
|
EF1A2_HUMAN
|
||||||
| θ value | 4.92598e-06 (rank : 11) | NC score | 0.299131 (rank : 14) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q05639, P54266 | Gene names | EEF1A2, EEF1AL, STN | |||
|
Domain Architecture |
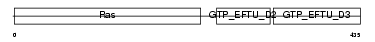
|
|||||
| Description | Elongation factor 1-alpha 2 (EF-1-alpha-2) (Elongation factor 1 A-2) (eEF1A-2) (Statin S1). | |||||
|
EF1A2_MOUSE
|
||||||
| θ value | 4.92598e-06 (rank : 12) | NC score | 0.299137 (rank : 13) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P62631, P27706 | Gene names | Eef1a2, Eef1al, Stn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor 1-alpha 2 (EF-1-alpha-2) (Elongation factor 1 A-2) (eEF1A-2) (Statin S1). | |||||
|
IF2M_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 13) | NC score | 0.373541 (rank : 9) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P46199 | Gene names | MTIF2 | |||
|
Domain Architecture |
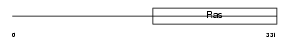
|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
IF2M_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 14) | NC score | 0.367214 (rank : 10) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91YJ5 | Gene names | Mtif2 | |||
|
Domain Architecture |
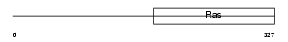
|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
IF2P_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 15) | NC score | 0.172041 (rank : 23) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
EFTU_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 16) | NC score | 0.276753 (rank : 15) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P49411, O15276 | Gene names | TUFM | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor Tu, mitochondrial precursor (EF-Tu) (P43). | |||||
|
EFTU_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 17) | NC score | 0.271047 (rank : 16) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BFR5, Q6P919 | Gene names | Tufm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor Tu, mitochondrial precursor. | |||||
|
HBS1L_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 18) | NC score | 0.249246 (rank : 17) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y450, Q5T7G3, Q8NDW9, Q9UPW3 | Gene names | HBS1L, HBS1, KIAA1038 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HBS1-like protein (ERFS). | |||||
|
HBS1L_MOUSE
|
||||||
| θ value | 0.125558 (rank : 19) | NC score | 0.239846 (rank : 18) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q69ZS7, Q91Z32, Q9CVT2, Q9CZ95, Q9WTY5 | Gene names | Hbs1l, Hbs1, Kiaa1038 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HBS1-like protein. | |||||
|
IF2H_MOUSE
|
||||||
| θ value | 0.125558 (rank : 20) | NC score | 0.107695 (rank : 24) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z0N2 | Gene names | Eif2s3y | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3, Y-linked (Eukaryotic translation initiation factor 2 subunit gamma, Y-linked) (eIF-2-gamma Y). | |||||
|
RT27_HUMAN
|
||||||
| θ value | 0.21417 (rank : 21) | NC score | 0.091780 (rank : 25) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92552 | Gene names | MRPS27, KIAA0264 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial 28S ribosomal protein S27 (S27mt) (MRP-S27). | |||||
|
SELB_HUMAN
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.187587 (rank : 22) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P57772 | Gene names | EEFSEC, SELB | |||
|
Domain Architecture |
|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific). | |||||
|
SELB_MOUSE
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.224250 (rank : 19) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9JHW4 | Gene names | Eefsec, Selb | |||
|
Domain Architecture |

|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific) (mSelB). | |||||
|
DCMC_MOUSE
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.046172 (rank : 32) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99J39, Q7TNL6 | Gene names | Mlycd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Malonyl-CoA decarboxylase, mitochondrial precursor (EC 4.1.1.9) (MCD). | |||||
|
FAK2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.002414 (rank : 43) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14289, Q13475, Q14290, Q16709 | Gene names | PTK2B, FAK2, PYK2, RAFTK | |||
|
Domain Architecture |

|
|||||
| Description | Protein tyrosine kinase 2 beta (EC 2.7.10.2) (Focal adhesion kinase 2) (FADK 2) (Proline-rich tyrosine kinase 2) (Cell adhesion kinase beta) (CAK beta) (Calcium-dependent tyrosine kinase) (CADTK) (Related adhesion focal tyrosine kinase) (RAFTK). | |||||
|
1A29_HUMAN
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.007474 (rank : 37) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P30512, O02955, O19687, O98011, P30451, P79562 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |
|
|||||
| Description | HLA class I histocompatibility antigen, A-29 alpha chain precursor (MHC class I antigen A*29) (Aw-19). | |||||
|
1A31_HUMAN
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.007467 (rank : 38) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P16189, O62924, O98009, O98137, Q8MHM1, Q9TPQ3, Q9TQ24, Q9UQU6, Q9UQU7 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |
|
|||||
| Description | HLA class I histocompatibility antigen, A-31 alpha chain precursor (MHC class I antigen A*31). | |||||
|
1A33_HUMAN
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.007458 (rank : 39) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P16190, O02939, O02954 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |
|
|||||
| Description | HLA class I histocompatibility antigen, A-33 alpha chain precursor (MHC class I antigen A*33) (Aw-33) (Aw-19). | |||||
|
ANLN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.025233 (rank : 33) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
ICAL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.023859 (rank : 34) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20810, O95360, Q96D08, Q9H1Z5 | Gene names | CAST | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor) (Sperm BS-17 component). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.014320 (rank : 35) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
GTPB2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.084512 (rank : 26) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BX10, Q5T7E8, Q8ND84, Q8TAH7, Q8WUA5, Q9HCS9, Q9NRU4, Q9NX60 | Gene names | GTPBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein 2. | |||||
|
GTPB2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 33) | NC score | 0.084454 (rank : 27) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q3UJK4, Q9EST7, Q9JIX7 | Gene names | Gtpbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein 2 (GTP-binding-like protein 2). | |||||
|
1A32_HUMAN
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.006069 (rank : 41) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P10314, O77937, Q29838, Q8MHN9, Q9MYE9, Q9MYG5, Q9TQF5, Q9TQF9 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |
|
|||||
| Description | HLA class I histocompatibility antigen, A-32 alpha chain precursor (MHC class I antigen A*32). | |||||
|
1A74_HUMAN
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.006095 (rank : 40) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P30459, O19647, O43906, O46874, Q95IZ5, Q9UIN1 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |
|
|||||
| Description | HLA class I histocompatibility antigen, A-74 alpha chain precursor (MHC class I antigen A*74) (Aw-74) (Aw-19). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.010270 (rank : 36) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
S27A2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.005043 (rank : 42) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35488, O70550, O88560 | Gene names | Slc27a2, Acsvl1, Facvl1, Fatp2, Vlacs, Vlcs | |||
|
Domain Architecture |

|
|||||
| Description | Very-long-chain acyl-CoA synthetase (EC 6.2.1.-) (VLCS) (Very-long- chain-fatty-acid-CoA ligase) (VLACS) (THCA-CoA ligase) (Fatty-acid- coenzyme A ligase, very long-chain 1) (Long-chain-fatty-acid--CoA ligase) (EC 6.2.1.3) (Fatty acid transport protein 2) (FATP-2) (Solute carrier family 27 member 2). | |||||
|
GSPT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 38) | NC score | 0.199430 (rank : 20) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15170 | Gene names | GSPT1 | |||
|
Domain Architecture |

|
|||||
| Description | G1 to S phase transition protein 1 homolog (GTP-binding protein GST1- HS). | |||||
|
GSPT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 39) | NC score | 0.199018 (rank : 21) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8R050, O88179 | Gene names | Gspt1 | |||
|
Domain Architecture |

|
|||||
| Description | G1 to S phase transition protein 1 homolog. | |||||
|
GTPB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.079178 (rank : 28) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O00178, Q6IC67 | Gene names | GTPBP1 | |||
|
Domain Architecture |
|
|||||
| Description | GTP-binding protein 1 (G-protein 1) (GP-1) (GP1). | |||||
|
GTPB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.078804 (rank : 29) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O08582, Q3UD96, Q3UGW6, Q545R1, Q80ZY5 | Gene names | Gtpbp1, Gtpbp | |||
|
Domain Architecture |
|
|||||
| Description | GTP-binding protein 1 (G-protein 1) (GP-1) (GP1). | |||||
|
IF2G_HUMAN
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.070602 (rank : 30) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P41091 | Gene names | EIF2S3, EIF2G | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3 (Eukaryotic translation initiation factor 2 subunit gamma) (eIF-2-gamma). | |||||
|
IF2G_MOUSE
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.070600 (rank : 31) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Z0N1 | Gene names | Eif2s3x | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3, X-linked (Eukaryotic translation initiation factor 2 subunit gamma, X-linked) (eIF-2-gamma X). | |||||
|
EF2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P13639 | Gene names | EEF2, EF2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 2 (EF-2). | |||||
|
EF2_MOUSE
|
||||||
| NC score | 0.999009 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P58252, Q544E4 | Gene names | Eef2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 2 (EF-2). | |||||
|
U5S1_HUMAN
|
||||||
| NC score | 0.950196 (rank : 3) | θ value | 3.96267e-181 (rank : 4) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15029, Q9BUR0 | Gene names | EFTUD2, KIAA0031, SNRP116 | |||
|
Domain Architecture |

|
|||||
| Description | 116 kDa U5 small nuclear ribonucleoprotein component (U5 snRNP- specific protein, 116 kDa) (U5-116 kDa) (Elongation factor Tu GTP- binding domain protein 2) (hSNU114). | |||||
|
U5S1_MOUSE
|
||||||
| NC score | 0.950023 (rank : 4) | θ value | 1.77876e-181 (rank : 3) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O08810 | Gene names | Eftud2, Snrp116 | |||
|
Domain Architecture |

|
|||||
| Description | 116 kDa U5 small nuclear ribonucleoprotein component (U5 snRNP- specific protein, 116 kDa) (U5-116 kDa) (Elongation factor Tu GTP- binding domain protein 2). | |||||
|
EFG2_MOUSE
|
||||||
| NC score | 0.712637 (rank : 5) | θ value | 9.8567e-31 (rank : 5) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R2Q4 | Gene names | Gfm2, Efg2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 2, mitochondrial precursor (mEF-G 2) (Elongation factor G2). | |||||
|
EFG2_HUMAN
|
||||||
| NC score | 0.681178 (rank : 6) | θ value | 1.4233e-29 (rank : 6) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q969S9, Q8WYI0 | Gene names | GFM2, EFG2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 2, mitochondrial precursor (mEF-G 2) (Elongation factor G2). | |||||
|
EFG1_MOUSE
|
||||||
| NC score | 0.646344 (rank : 7) | θ value | 2.69047e-20 (rank : 8) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K0D5, Q921D6, Q924I0 | Gene names | Gfm1, Efg, Efg1, Gfm | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 1, mitochondrial precursor (mEF-G 1) (Elongation factor G1). | |||||
|
EFG1_HUMAN
|
||||||
| NC score | 0.637180 (rank : 8) | θ value | 8.36355e-22 (rank : 7) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96RP9, Q96T39 | Gene names | GFM1, EFG, EFG1, GFM | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor G 1, mitochondrial precursor (mEF-G 1) (Elongation factor G1). | |||||
|
IF2M_HUMAN
|
||||||
| NC score | 0.373541 (rank : 9) | θ value | 2.44474e-05 (rank : 13) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P46199 | Gene names | MTIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
IF2M_MOUSE
|
||||||
| NC score | 0.367214 (rank : 10) | θ value | 0.000158464 (rank : 14) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91YJ5 | Gene names | Mtif2 | |||
|
Domain Architecture |

|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
EF1A1_HUMAN
|
||||||
| NC score | 0.307951 (rank : 11) | θ value | 3.41135e-07 (rank : 9) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P68104, P04719, P04720 | Gene names | EEF1A1, EEF1A, EF1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor 1-alpha 1 (EF-1-alpha-1) (Elongation factor 1 A-1) (eEF1A-1) (Elongation factor Tu) (EF-Tu). | |||||
|
EF1A1_MOUSE
|
||||||
| NC score | 0.307939 (rank : 12) | θ value | 3.41135e-07 (rank : 10) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P10126, Q61511, Q6ZWN2, Q8BMB8, Q8BVS8, Q99KU5 | Gene names | Eef1a1, Eef1a | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 1-alpha 1 (EF-1-alpha-1) (Elongation factor 1 A-1) (eEF1A-1) (Elongation factor Tu) (EF-Tu). | |||||
|
EF1A2_MOUSE
|
||||||
| NC score | 0.299137 (rank : 13) | θ value | 4.92598e-06 (rank : 12) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P62631, P27706 | Gene names | Eef1a2, Eef1al, Stn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor 1-alpha 2 (EF-1-alpha-2) (Elongation factor 1 A-2) (eEF1A-2) (Statin S1). | |||||
|
EF1A2_HUMAN
|
||||||
| NC score | 0.299131 (rank : 14) | θ value | 4.92598e-06 (rank : 11) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q05639, P54266 | Gene names | EEF1A2, EEF1AL, STN | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 1-alpha 2 (EF-1-alpha-2) (Elongation factor 1 A-2) (eEF1A-2) (Statin S1). | |||||
|
EFTU_HUMAN
|
||||||
| NC score | 0.276753 (rank : 15) | θ value | 0.0113563 (rank : 16) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P49411, O15276 | Gene names | TUFM | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor Tu, mitochondrial precursor (EF-Tu) (P43). | |||||
|
EFTU_MOUSE
|
||||||
| NC score | 0.271047 (rank : 16) | θ value | 0.0252991 (rank : 17) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BFR5, Q6P919 | Gene names | Tufm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Elongation factor Tu, mitochondrial precursor. | |||||
|
HBS1L_HUMAN
|
||||||
| NC score | 0.249246 (rank : 17) | θ value | 0.0961366 (rank : 18) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y450, Q5T7G3, Q8NDW9, Q9UPW3 | Gene names | HBS1L, HBS1, KIAA1038 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HBS1-like protein (ERFS). | |||||
|
HBS1L_MOUSE
|
||||||
| NC score | 0.239846 (rank : 18) | θ value | 0.125558 (rank : 19) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q69ZS7, Q91Z32, Q9CVT2, Q9CZ95, Q9WTY5 | Gene names | Hbs1l, Hbs1, Kiaa1038 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HBS1-like protein. | |||||
|
SELB_MOUSE
|
||||||
| NC score | 0.224250 (rank : 19) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9JHW4 | Gene names | Eefsec, Selb | |||
|
Domain Architecture |

|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific) (mSelB). | |||||
|
GSPT1_HUMAN
|
||||||
| NC score | 0.199430 (rank : 20) | θ value | θ > 10 (rank : 38) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15170 | Gene names | GSPT1 | |||
|
Domain Architecture |

|
|||||
| Description | G1 to S phase transition protein 1 homolog (GTP-binding protein GST1- HS). | |||||
|
GSPT1_MOUSE
|
||||||
| NC score | 0.199018 (rank : 21) | θ value | θ > 10 (rank : 39) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8R050, O88179 | Gene names | Gspt1 | |||
|
Domain Architecture |

|
|||||
| Description | G1 to S phase transition protein 1 homolog. | |||||
|
SELB_HUMAN
|
||||||
| NC score | 0.187587 (rank : 22) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P57772 | Gene names | EEFSEC, SELB | |||
|
Domain Architecture |

|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.172041 (rank : 23) | θ value | 0.00102713 (rank : 15) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
IF2H_MOUSE
|
||||||
| NC score | 0.107695 (rank : 24) | θ value | 0.125558 (rank : 20) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z0N2 | Gene names | Eif2s3y | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3, Y-linked (Eukaryotic translation initiation factor 2 subunit gamma, Y-linked) (eIF-2-gamma Y). | |||||
|
RT27_HUMAN
|
||||||
| NC score | 0.091780 (rank : 25) | θ value | 0.21417 (rank : 21) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92552 | Gene names | MRPS27, KIAA0264 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial 28S ribosomal protein S27 (S27mt) (MRP-S27). | |||||
|
GTPB2_HUMAN
|
||||||
| NC score | 0.084512 (rank : 26) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BX10, Q5T7E8, Q8ND84, Q8TAH7, Q8WUA5, Q9HCS9, Q9NRU4, Q9NX60 | Gene names | GTPBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein 2. | |||||
|
GTPB2_MOUSE
|
||||||
| NC score | 0.084454 (rank : 27) | θ value | 6.88961 (rank : 33) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q3UJK4, Q9EST7, Q9JIX7 | Gene names | Gtpbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTP-binding protein 2 (GTP-binding-like protein 2). | |||||
|
GTPB1_HUMAN
|
||||||
| NC score | 0.079178 (rank : 28) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O00178, Q6IC67 | Gene names | GTPBP1 | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein 1 (G-protein 1) (GP-1) (GP1). | |||||
|
GTPB1_MOUSE
|
||||||
| NC score | 0.078804 (rank : 29) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O08582, Q3UD96, Q3UGW6, Q545R1, Q80ZY5 | Gene names | Gtpbp1, Gtpbp | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein 1 (G-protein 1) (GP-1) (GP1). | |||||
|
IF2G_HUMAN
|
||||||
| NC score | 0.070602 (rank : 30) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P41091 | Gene names | EIF2S3, EIF2G | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3 (Eukaryotic translation initiation factor 2 subunit gamma) (eIF-2-gamma). | |||||
|
IF2G_MOUSE
|
||||||
| NC score | 0.070600 (rank : 31) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Z0N1 | Gene names | Eif2s3x | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 3, X-linked (Eukaryotic translation initiation factor 2 subunit gamma, X-linked) (eIF-2-gamma X). | |||||
|
DCMC_MOUSE
|
||||||
| NC score | 0.046172 (rank : 32) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99J39, Q7TNL6 | Gene names | Mlycd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Malonyl-CoA decarboxylase, mitochondrial precursor (EC 4.1.1.9) (MCD). | |||||
|
ANLN_HUMAN
|
||||||
| NC score | 0.025233 (rank : 33) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
ICAL_HUMAN
|
||||||
| NC score | 0.023859 (rank : 34) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20810, O95360, Q96D08, Q9H1Z5 | Gene names | CAST | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor) (Sperm BS-17 component). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.014320 (rank : 35) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.010270 (rank : 36) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
1A29_HUMAN
|
||||||
| NC score | 0.007474 (rank : 37) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P30512, O02955, O19687, O98011, P30451, P79562 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |

|
|||||
| Description | HLA class I histocompatibility antigen, A-29 alpha chain precursor (MHC class I antigen A*29) (Aw-19). | |||||
|
1A31_HUMAN
|
||||||
| NC score | 0.007467 (rank : 38) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P16189, O62924, O98009, O98137, Q8MHM1, Q9TPQ3, Q9TQ24, Q9UQU6, Q9UQU7 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |

|
|||||
| Description | HLA class I histocompatibility antigen, A-31 alpha chain precursor (MHC class I antigen A*31). | |||||
|
1A33_HUMAN
|
||||||
| NC score | 0.007458 (rank : 39) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P16190, O02939, O02954 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |

|
|||||
| Description | HLA class I histocompatibility antigen, A-33 alpha chain precursor (MHC class I antigen A*33) (Aw-33) (Aw-19). | |||||
|
1A74_HUMAN
|
||||||
| NC score | 0.006095 (rank : 40) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P30459, O19647, O43906, O46874, Q95IZ5, Q9UIN1 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |

|
|||||
| Description | HLA class I histocompatibility antigen, A-74 alpha chain precursor (MHC class I antigen A*74) (Aw-74) (Aw-19). | |||||
|
1A32_HUMAN
|
||||||
| NC score | 0.006069 (rank : 41) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P10314, O77937, Q29838, Q8MHN9, Q9MYE9, Q9MYG5, Q9TQF5, Q9TQF9 | Gene names | HLA-A, HLAA | |||
|
Domain Architecture |

|
|||||
| Description | HLA class I histocompatibility antigen, A-32 alpha chain precursor (MHC class I antigen A*32). | |||||
|
S27A2_MOUSE
|
||||||
| NC score | 0.005043 (rank : 42) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35488, O70550, O88560 | Gene names | Slc27a2, Acsvl1, Facvl1, Fatp2, Vlacs, Vlcs | |||
|
Domain Architecture |

|
|||||
| Description | Very-long-chain acyl-CoA synthetase (EC 6.2.1.-) (VLCS) (Very-long- chain-fatty-acid-CoA ligase) (VLACS) (THCA-CoA ligase) (Fatty-acid- coenzyme A ligase, very long-chain 1) (Long-chain-fatty-acid--CoA ligase) (EC 6.2.1.3) (Fatty acid transport protein 2) (FATP-2) (Solute carrier family 27 member 2). | |||||
|
FAK2_HUMAN
|
||||||
| NC score | 0.002414 (rank : 43) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14289, Q13475, Q14290, Q16709 | Gene names | PTK2B, FAK2, PYK2, RAFTK | |||
|
Domain Architecture |

|
|||||
| Description | Protein tyrosine kinase 2 beta (EC 2.7.10.2) (Focal adhesion kinase 2) (FADK 2) (Proline-rich tyrosine kinase 2) (Cell adhesion kinase beta) (CAK beta) (Calcium-dependent tyrosine kinase) (CADTK) (Related adhesion focal tyrosine kinase) (RAFTK). | |||||